INTRODUCTION
Female reproductive system cancer, which includes cervical cancer, endometrial cancer, and ovarian cancer, is a major threat to women’s health. In fact, cervical cancer ranks third among all gynecologic cancers in the world [1], 65 new endometrial cancer cases occur annually per 100,000 women between the ages of 65 to 75 [2], and approximately 140,200 new ovarian cancer cases worldwide per year are recorded [3]. Elucidating the etiology of female reproductive system cancer and identifying at-risk populations may allow more effective early detection and perhaps even prevention.
The causes of these cancers remain poorly understood. Infection with oncogenic human papilloma virus (HPV) is a risk factor in tumorigenesis [4], but many HPV carriers do not develop cervical cancer, indicating that there must be other cancer risk factors, such as genetic and environmental factors. One possible genetic factor may be polymorphism in the gene encoding x-ray repair cross complementing 1 (XRCC1) protein. The gene is located on chromosome 19 (19q13.2), and the expressed protein is involved in the base excision repair (BER) pathway [5, 6], which helps correct errors during DNA replication and recombination as well as preserve genome integrity [7]. Functional single-nucleotide polymorphisms (SNPs) in XRCC1 have been linked to development of esophageal squamous cell carcinoma [8], lung cancer [9], pancreatic cancer [10], breast cancer [11], colorectal cancer [12], and gastric cancer [13]. While more than 300 XRCC1 SNPs have been described in the dbSNP database, three functional SNPs have been extensively studied, all of which cause amino acid substitutions in the encoded protein: rs25487 [Arg399Gln], rs1799782 [Arg194Trp] and rs25489 [Arg280His].
Associations between XRCC1 SNPs and risk of female reproducitve system cancer are unclear, because the several molecular epidemiologic studies conducted so far have been inconclusive. This lack of clarity may reflect the relatively small statistical power in individual studies, as well as heterogeneity in genetic backgrounds of study participants. Therefore we performed this meta-analysis to comprehensively assess available evidence on the association between XRCC1 polymorphism and risk of female reproductive system cancer.
RESULTS
Study characteristics
Systematic search of the PubMed, EMBASE and China National Knowledge Infrastructure (CNKI) databases identified 157 potentially relevant studies (Figure 1). Further screening allowed elimination of all but 31 studies, which were read in full. In the end, 26 case-control studies were included in this study (Table 1): data on rs25487 were reported in 24 studies involving 4,265 cases and 5,495 controls; data on rs1799782 were reported in 15 studies involving 2,672 cases and 3,578 controls; and data on rs25489 were reported in 5 studies with 907 cases and 1,416 controls. The various ethnic groups involved in the studies were divided into two categories: Asian or Non-Asian, with the latter including Caucasian, Latino, and mixed.
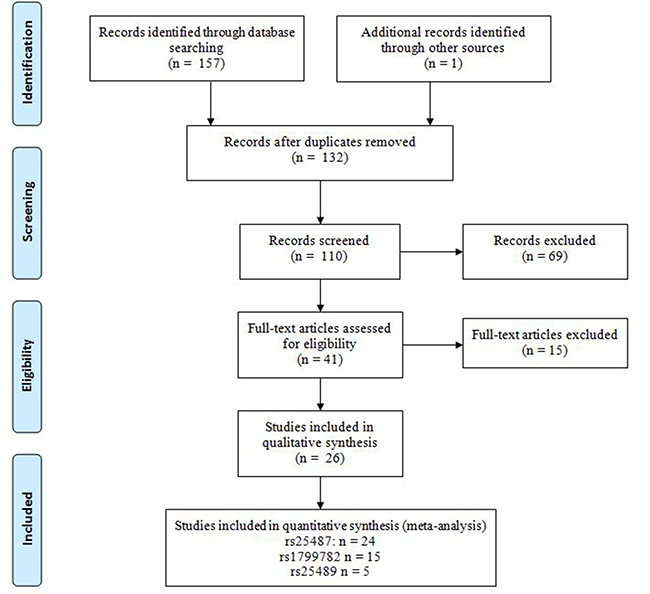
Figure 1: PRISMA flow diagram.
Table 1: Characteristics of studies included in the meta-analysis
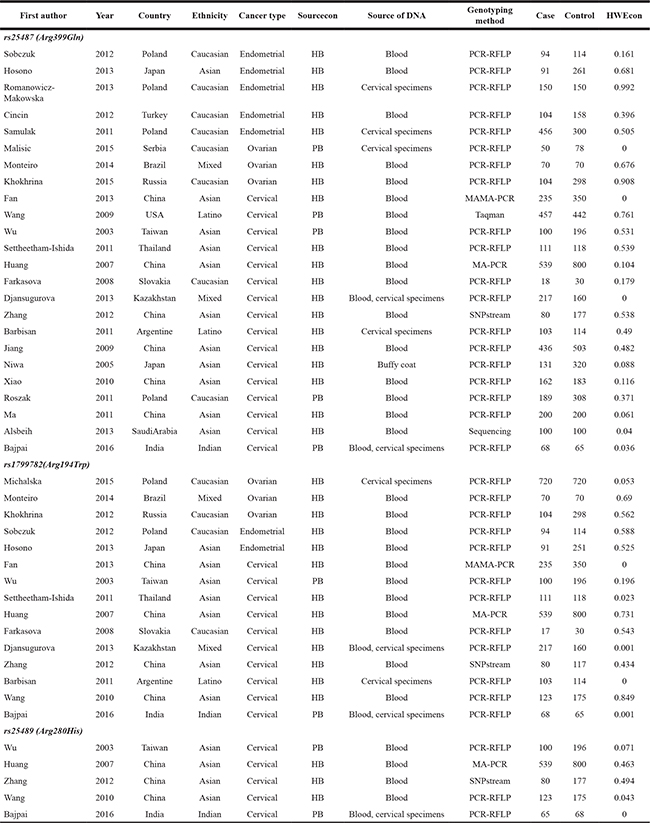
Sourcecon: Source of control. HWEcon: Hardy–Weinberg equilibrium in controls. PB, population-based; HB, hospital-based.
Of the 24 studies related to rs25487, 16 focused on cervical cancer [14–29], 5 on endometrial cancer [30–34], and 3 on ovarian cancer [35–37]. The ethnic group was Asian in 11 studies and Non-Asian in the others. Of the 15 studies related to rs1799782, 10 focused on cervical cancer [16–20, 25, 27, 28], 3 on endometrial cancer [35, 37, 38], and 2 on ovarian cancer [30, 31]. The ethnic group was Asian in 7 studies and Non-Asian in 8 studies. Of the 5 studies related to rs25489, all focused on cervical cancer. The population was Asian in 4 studies and Non-Asian in one study.
Across all studies, the distribution of genotypes in controls was mostly in agreement with Hardy-Weinberg equilibrium (HWE), except in 5 studies on rs25487 [6, 15, 17, 18, 36], 5 studies on rs1799782 [15–18, 26] and 2 studies on rs25489 [15, 39].
Quantitative data synthesis
Across the entire pooled study population, a significant association was not found between rs25487 and risk of female reproductive system cancer (Table 2). In the subgroup of Asian participants, however, we detected a significant association of the A variant at rs25487 with increased risk of female reproductive system cancer (GA vs. GG, OR 1.16, 95%CI 1.00–1.36). This association disappeared when we excluded 5 studies that deviated from HWE, instead appearing in the subgroup of Non-Asian participants (AA vs. GA/GG, OR 1.61, 95%CI 1.41–1.85). Subgroup analysis by cancer type indicated an association between the A allele at rs25487 and increased risk of cervical cancer (AA vs. GA/GG, OR 1.22, 95%CI 1.05–1.41), endometrial cancer (AA vs. GG, OR 2.16, 95%CI 1.00–4.67) and ovarian cancer (AA vs. GA/GG, OR 2.01, 95%CI 1.70–2.38). The significant associations with cervical cancer and ovarian cancer disappeared after removing studies that deviated from HWE.
Table 2: Meta-analysis of the associations between XRCC1 polymorphisms and risk of female reproductive system cancer
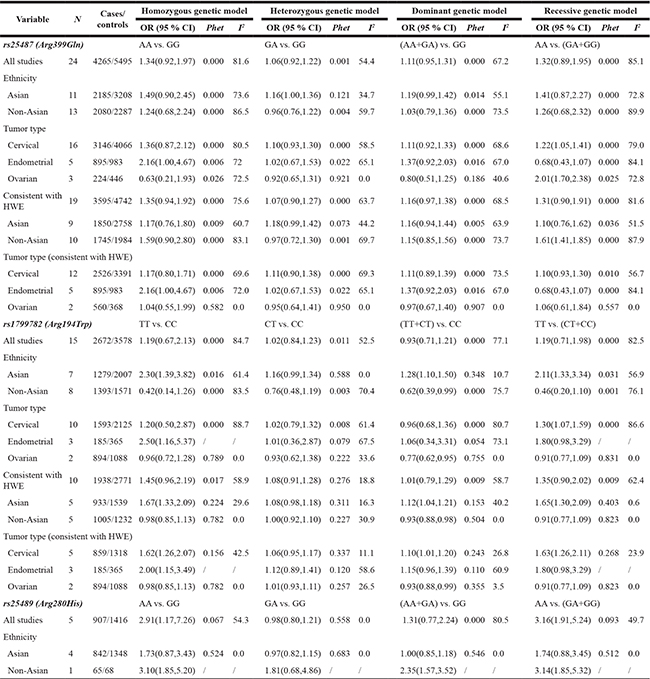
N: Number of studies. Phet: P value for heterogeneity test. HWE: Hardy–Weinberg equilibrium.
Across the entire pooled study population, no association was found between rs1799782 and risk of female reproductive system cancer (Table 2). In subgroup analysis by ethnicity, the T variant was significantly associated with increased risk of female reproductive system cancer in Asians (TT vs. CC, OR 2.30, 95%CI 1.39–3.82; TT/CT vs. CC, OR 1.28, 95%CI 1.10–1.50; TT vs. CT/CC, OR 2.11, 95%CI 1.33–3.34). This association remained significant after excluding 5 studies that deviated from HWE(TT vs. CC, OR 1.67, 95%CI 1.33–2.09; TT/CT vs. CC, OR 1.12, 95%CI 1.04–1.21; TT vs. CT/CC, OR 1.65, 95%CI 1.30–2.09). In subgroup analysis by tumor type, the T allele was associated with increased risk of cervical cancer (TT vs. CT/CC, OR 1.30, 95%CI 1.07–1.59), and this association remained significant after excluding studies that deviated from HWE (TT vs. CC, OR 1.62, 95%CI 1.26–2.07; TT/CT vs. CC, OR 1.10, 95%CI 1.01–1.20; TT vs. CT/CC, OR 1.63, 95%CI 1.26–2.11). The same T variant increased risk of endometrial cancer based on all study participants (TT vs. CC, OR 2.50, 95%CI 1.16–5.37) as well as based on only studies consistent with HWE (TT vs. CC, OR 2.00, 95%CI 1.15–3.49).
Data on rs25489 SNPs were limited to cervical cancer studies. Meta-analysis suggested that the A variant was associated with increased risk of this cancer (AA vs. GG, OR 2.91, 95%CI 1.17–7.26; AA vs. GA/GG, OR 3.16, 95%CI 1.91–5.24).
Heterogeneity and sensitivity analyses
Significant heterogeneity across studies was observed in the meta-analysis of the association between the A variant at rs25487 and risk of female reproductive system cancer (homozygous model, I2 = 81.6, P = 0.000; heterozygous model: I2 = 54.4, P = 0.001; dominant model: I2 = 67.2, P < 0.001; recessive model: I2 = 85.1, P < 0.001). Similarly, significant heterogeneity across studies was observed in the meta-analysis of the association between the T allele at rs1799782 and cancer risk (homozygous model, I2 = 84.7, P < 0.001; heterozygous model, I2 = 52.5, P = 0.011; dominant model, I2 = 77.1, P < 0.001; recessive model: I2 = 82.5, P < 0.001). Among studies used in meta-analyses involving rs25489, we found moderate heterogeneity in the homozygous model (I2 = 54.3, P = 0.067) and dominant model (I2 = 80.5, P < 0.001), but no significant heterogeneity in the heterozygous or recessive models.
Then we performed sensitivity analysis, in which we recalculated the meta-analysis after deleting each study systematically. The results were not substantially different after excluding any single study, indicating the robustness of our original meta-analyses.
DISCUSSION
XRCC1 is the first protein to participate in the BER pathway, acting as a scaffold for other DNA repair proteins, such as DNA ligase IIIa, DNA polymerase β and poly (ADP-ribose) polymerase [40]. The XRCC1 SNPs rs25487, rs1799782, and rs25489 have been linked to susceptibility for several types of cancer, but it is unclear whether this is also true for female reproductive system cancer. The present meta-analysis suggests that as in other cancers, XRCC1 polymorphism may also influence tumorigenesis in the female reproductive system.
The present study may be the first quantitative meta-analysis of XRCC1 polymorphism and risk of female reproductive system cancer. Previous meta-analyses focused only on cervical cancer risk [41–45], and they reported that rs25487 and rs1799782 were associated with increased risk in Asian populations. The present meta-analysis extended this finding by showing that in Asians, the A allele of rs25487 and T allele of rs1799782 are associated with increased risk of female reproductive system cancer. These findings should be verified in larger studies, especially since the association with rs25487 disappeared in Asians and appeared in Non-Asians after we excluded studies deviating from HWE.
We did not observe a significant association between XRCC1 rs25489 and risk of ovarian cancer, even though estrogens and their metabolites damage DNA by forming bulky DNA adducts [46], which are normally repaired by the BER pathway. It is possible that our negative results reflect limited sample size. Further studies are needed to verify our findings.
To ensure results as reliable as possible, we analogized the studies in our meta-analysis to approximate randomized controlled trials. At the same time, our study does have several limitations. Our results were based on OR analyses that did not adjust for age, family history, gender, reproductive history, or other biological factors that might influence risk of female reproductive system cancer. Similarly, we did not take into account potential effects of gene-environment interaction. Therefore further work is needed before definitive conclusions can be drawn about these three XRCC1 SNPs and risk of female reproductive system cancer.
Despite these limitations, our data provide up-to-date evidence from a comprehensive review of the literature that the A allele of rs25487 and T allele of rs1799782 are low-penetration risk factors for female reproductive system cancer in Asians. It may be that these alleles translate to weaker interaction between XRCC1 and other repair proteins, thereby reducing DNA repair capacity [47]. Our findings add to the growing evidence that polymorphism in DNA repair genes can destabilize the genome and increase tumor susceptibility [32].
MATERIALS AND METHODS
Literature search strategy
We performed a comprehensive literature search in the PubMed, EMBASE, Scopus, and CNKI databases up to December 17, 2016. The following search strings were used: “X-ray repair cross complementing protein1” or “XRCC1”; “polymorphisms” or “variants”; “carcinoma” or “cancer” or “malignancy” or “neoplasm” or “tumour” or “tumor”; “cervical” or “endometrial” or “ovarian” or “vaginal” or “vulvar” or “fallopian tube” or “female reproductive system”. The reference lists of relevant articles were also searched manually to identify additional eligible studies. When different studies presented overlapping data, we included only the larger study.
Inclusion and exclusion criteria
Studies included in this meta-analysis had to be case-control or genome-wide association studies for which full text was available and that reported adequate data on genotype frequencies for cases and controls. Studies were excluded if they reported data overlapping with those of a larger study.
Data extraction
Two investigators (N.-N.Y and Y.-F.H) independently extracted the following data from all eligible publications: the first author’s name, year of publication, country of origin, ethnicity, study type (retrospective or prospective), source of control subjects (population-based [PB] or hospital-based [HB]), DNA source (e.g., blood, lymphocytes or buffy coat), genotyping method, total numbers of cases and controls and P value for HWE. Conflicts were resolved by consensus among all authors.
Statistical analysis
Agreement between study results and HWE predictions was tested using the goodness-of-fit χ2 test, with the threshold for HWE defined as P > 0.05. Strength of association between XRCC1 SNPs and cancer risk was assessed using odds ratios (ORs) and associated 95% confidence intervals (95%CI). Four different genetic models were conducted to detect the association: homozygous model (VV vs. WW), heterozygous model (WV vs. WW), dominant genetic model (VV+WV vs. WW) and recessive model (VV vs. WW+WV), with W and V representing the wild and variant alleles of each SNP. Heterogeneity across studies was assessed using a χ2-test-based Q statistic test, and the level of heterogeneity was quantified using the I2 test. When heterogeneity across studies was obvious (P ≥ 0.05 or I2 < 50%) [48], the random effects model was used to meta-analyze data from different studies. Otherwise, the fixed effects model was adopted [49]. All studies were analogized into interim randomized controlled clinical trials in order to control for type I and type II error.
Subgroup analyses were carried out based on ethnicity, tumor type and HWE [50]. Sensitivity analysis was performed in which we recalculated the meta-analysis after deleting each study systematically. Publication bias was investigated using Begg’s and Egger’s test [51], with significant risk of bias defined as P < 0.05. All statistical analyses were performed using STATA 11.0 (Stata Corp, College Station, Texas USA). All P values were two-sided.
Authors’ contributions
Guarantors of integrity of entire study, J.F.J.; study concepts/study design or data acquisition or data analysis/interpretation, all authors; manuscript drafting or manuscript revision for important intellectual content, all authors; manuscript final version approval, all authors; literature research, N.N.Y., Y.F.H.; statistical analysis, N.N.Y., Y.F.H.; and manuscript editing, N.N.Y., J.S., J.F.J.
CONFLICTS OF INTEREST
None.
FUNDING
This research was funded by the Guangxi University of Science and Technology Research Project (KY2015YB073) and the Guangxi Medical Care Appropriate Technology Research and Development (S201627). The funding source had no role in the design or conduct of the study; in the collection, analysis or interpretation of the data; or in the preparation, review, or approval of the manuscript.
REFERENCES
1. Siegel RL, Miller KD, Jemal A. Cancer Statistics, 2017. CA Cancer J Clin. 2017; 67:7–30.
2. Smith RA, Andrews K, Brooks D, DeSantis CE, Fedewa SA, Lortet-Tieulent J, Manassaram-Baptiste D, Brawley OW, Wender RC. Cancer screening in the United States, 2016: A review of current American Cancer Society guidelines and current issues in cancer screening. CA Cancer J Clin. 2016; 66:96–114.
3. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015; 65:87–108.
4. Ahn WS, Bae SM, Chung JE, Lee HK, Kim BK, Lee JM, Namkoong SE, Kim CK, Sin J. Evaluation of adenoassociated virus 2 and human papilloma virus 16 and 18 infection in cervical cancer biopsies. Gynecol Oncol. 2003; 89:105–111.
5. Fan J, Wilson DM 3rd. Protein-protein interactions and posttranslational modifications in mammalian base excision repair. Free Radic Biol Med. 2005; 38:1121–1138.
6. Al Zoubi MS. X-ray repair cross-complementing protein 1 and 3 polymorphisms and susceptibility of breast cancer in a Jordanian population. Saudi Med J. 2015; 36:1163–1167.
7. Rasouly A, Pani B, Nudler E. A Magic Spot in Genome Maintenance. Trends Genet. 2017; 33:58–67.
8. Yun YX, Dai LP, Wang P, Wang KJ, Zhang JY, Xie W. Association of polymorphisms in X-ray repair cross complementing 1 gene and risk of esophageal squamous cell carcinoma in a Chinese population. Biomed Res Int. 2015; 2015:509215.
9. Fan X, Xiu Q. Effect of X-ray repair cross complementing group 1 polymorphisms on the efficacy of platinum-based chemotherapy in patients with nonsmall cell lung cancer. J Cancer Res Ther. 2015; 11:571–574.
10. Hou BH, Jian ZX, Cui P, Li SJ, Tian RQ, Ou JR. Association and Intragenic Single-Nucleotide Polymorphism Interactions of the XRCC1 Polymorphisms for Pancreatic Cancer Susceptibility. Pancreas. 2016; 45:546–551.
11. Zheng L, Yang F, Zhang X, Zhu J, Zhou P, Yu F, Hou L, Zhao G, He Q, Wang B. Investigation on XRCC1 genetic polymorphism and its relationship with breast cancer risk factors in Chinese women. Med Oncol. 2013; 30:738.
12. Nissar S, Sameer AS, Rasool R, Chowdri NA, Rashid F. Polymorphism of the DNA Repair Gene XRCC1 (Arg194Trp) and its role in Colorectal Cancer in Kashmiri Population: a Case Control Study. Asian Pac J Cancer Prev. 2015; 16:6385–6390.
13. Zhao DY, Cheng L, Yu J, Shen H. XRCC1 genetic polymorphism Arg339Gln, Arg194Trp, Arg280His and gastric cancer risk: an evidence based decision. Cancer Biomark. 2014; 14:449–456.
14. Alsbeih G, Al-Harbi N, El-Sebaie M, Al-Badawi I. HPV prevalence and genetic predisposition to cervical cancer in Saudi Arabia. Infect Agent Cancer. 2013; 8:15.
15. Bajpai D, Banerjee A, Pathak S, Thakur B, Jain SK, Singh N. Single nucleotide polymorphisms in the DNA repair genes in HPV-positive cervical cancer. Eur J Cancer Prev. 2016; 25:224–231.
16. Barbisan G, Perez LO, Difranza L, Fernandez CJ, Ciancio NE, Golijow CD. XRCC1 Arg399Gln polymorphism and risk for cervical cancer development in Argentine women. Eur J Gynaecol Oncol. 2011; 32:274–279.
17. Djansugurova LB, Perfilyeva AV, Zhunusova GS, Djantaeva KB, Iksan OA, Khussainova EM. The determination of genetic markers of age-related cancer pathologies in populations from Kazakhstan. Front Genet. 2013; 4:70.
18. Fan XM, Li KX, Niu SH, Fang ZH, Liu H. [Relationship of XRCC1 polymorphism with the risks and clinicopathological factors of cervical cancer]. [Article in Chinese]. Zhonghua Yi Xue Za Zhi. 2013; 93:3454–6.
19. Farkasova T, Gurska S, Witkovsky V, Gabelova A. Significance of amino acid substitution variants of DNA repair genes in radiosusceptibility of cervical cancer patients; a pilot study. Neoplasma. 2008; 55:330–337.
20. Huang J, Ye F, Chen H, Lu W, Xie X. The nonsynonymous single nucleotide polymorphisms of DNA repair gene XRCC1 and susceptibility to the development of cervical carcinoma and high-risk human papillomavirus infection. Int J Gynecol Cancer. 2007; 17:668–675.
21. JiangW, Wang ML, Zhang ZZ, Chen XJ, Zhu H. The relationship between XRCC1 polymorphisms and the risk of cervical cancer in Jiangsu population. Acta Univ Med Nanjing. 2009; 29:1–6.
22. Ma W, Jin P. Single nucleotide polymorphisms of the DNA repair genes XPD and XRCC1 and the susceptibility to cervical squamous cell carcinoma. Prog Obstet Gynecol. 2011; 20:881–885.
23. Niwa Y, Matsuo K, Ito H, Hirose K, Tajima K, Nakanishi T, Nawa A, Kuzuya K, Tamakoshi A, Hamajima N. Association of XRCC1 Arg399Gln and OGG1 Ser326Cys polymorphisms with the risk of cervical cancer in Japanese subjects. Gynecol Oncol. 2005; 99:43–49.
24. Roszak A, Lianeri M, Jagodzinski PP. Involvement of the XRCC1 Arg399Gln gene polymorphism in the development of cervical carcinoma. Int J Biol Markers. 2011; 26:216–220.
25. Settheetham-Ishida W, Yuenyao P, Natphopsuk S, Settheetham D, Ishida T. Genetic risk of DNA repair gene polymorphisms (XRCC1 and XRCC3) for high risk human papillomavirus negative cervical cancer in Northeast Thailand. Asian Pac J Cancer Prev. 2011; 12:963–966.
26. Wang SS, Bratti MC, Rodriguez AC, Herrero R, Burk RD, Porras C, González P, Sherman ME, Wacholder S, Lan ZE, Schiffman M, Chanock SJ, Hildesheim A. Common variants in immune and DNA repair genes and risk for human papillomavirus persistence and progression to cervical cancer. J Infect Dis. 2009; 199:20–30.
27. Wu MT, Chen SY, Wu TN, Hwang HY, Ho CK, Lee LH, Wu SC. No association between polymorphisms of the DNA repair geneXRCC1 and cervical neoplasm risk. Environ Health Prev Med. 2003; 8:100–103.
28. Zhang L, Ruan Z, Hong Q, Gong X, Hu Z, Huang Y, Xu A. Single nucleotide polymorphisms in DNA repair genes and risk of cervical cancer: A case-control study. Oncol Lett. 2012; 3:351–362.
29. Xiao HY, Wu WQ, Xie H. Relationship between the polymorphism of XRCC-Arg399Gln and incidence risk of cervical cancer in the population of Guangdong. Hainan Med J. 2010; 21:35–37.
30. Sobczuk A, Poplawski T, Blasiak J. Polymorphisms of DNA repair genes in endometrial cancer. Pathol Oncol Res. 2012; 18:1015–1020.
31. Hosono S, Matsuo K, Ito H, Oze I, Hirose K, Watanabe M, Nakanishi T, Tajima K, Tanaka H. Polymorphisms in base excision repair genes are associated with endometrial cancer risk among postmenopausal Japanese women. Int J Gynecol Cancer. 2013; 23:1561–1568.
32. Romanowicz-Makowska H, Smolarz B, Houli A, Szyllo K. Single nucleotide polymorphism in DNA base excision repair genes XRCC1 and hOGG1 and the risk of endometrial carcinoma in the Polish population. Pol J Pathol. 2011; 62:89–94.
33. Cincin ZB, Iyibozkurt AC, Kuran SB, Cakmakoglu B. DNA repair gene variants in endometrial carcinoma. Med Oncol. 2012; 29:2949–2954.
34. Samulak D, Romanowicz-Makowska H, Smolarz B, Kulig A, Sporny S. Association between Arg399Gln polymorphism of X-ray repair cross-complementing 1 (XRCC1) gene and sporadic endometrial cancer in the Polish population. Eur J Gynaecol Oncol. 2011; 32:491–495.
35. Michalska MM, Samulak D, Romanowicz H, Bieńkiewicz J, Sobkowski M, Ciesielski K, Smolarz B. Single nucleotide polymorphisms (SNPs) of hOGG1 and XRCC1 DNA repair genes and the risk of ovarian cancer in Polish women. Tumour Biol. 2015; 36:9457–9463.
36. Malisic EJ, Krivokuca AM, Boljevic IZ, Jankovic RN. Impact of RAD51 G135C and XRCC1 Arg399Gln polymorphisms on ovarian carcinoma risk in Serbian women. Cancer Biomark. 2015; 15:685–691.
37. Monteiro MS, Vilas Boas DB, Gigliotti CB, Salvadori DM. Association among XRCC1, XRCC3, and BLHX gene polymorphisms and chromosome instability in lymphocytes from patients with endometriosis and ovarian cancer. Genet Mol Res. 2014; 13:636–648.
38. Khokhrin DV, Khrunin AV, Moiseev AA, Gorbunov VA, Limborskaia SA. Association of polymorphisms in glutathione-S-transferase and DNA repair genes with ovarian cancer risk in the Russian population. Genetika. 2012; 48:901–904.
39. Wang X. The association between XRCC1 polymorphisms and cervical cancer risk. Mod Med Health. 2010; 26:2137–2138.
40. Whitehouse CJ, Taylor RM, Thistlethwaite A, Zhang H, Karimi-Busheri F, Lasko DD, Weinfeld M, Caldecott KW. XRCC1 stimulates human polynucleotide kinase activity at damaged DNA termini and accelerates DNA single-strand break repair. Cell. 2001; 104:107–117.
41. Zeng X, Zhang Y, Yue T, Zhang T, Wang J, Xue Y, An R. Association between XRCC1 polymorphisms and the risk of cervical cancer: a meta-analysis based on 4895 subjects. Oncotarget. 2017; 8:2249–2260. doi: 10.18632/oncotarget.13663.
42. Liu DY, Liang HC, Xiao XM. Association between the XRCC1 Arg399Gln polymorphism and risk of cervical carcinoma: a meta-analysis. Genet Mol Res. 2015; 14:9821–9828.
43. Liu YT, Shi JP, Fu LY, Zhou B, Wang HL, Wu XM. Gene polymorphism of XRCC1 Arg399Gln and cervical carcinoma susceptibility in Asians: a meta-analysis based on 1,759 cases and 2,497 controls. Asian Pac J Cancer Prev. 2013; 14:189–193.
44. Mei J, Duan HX, Wang LL, Yang S, Lu JQ, Shi TY, Zhao Y. XRCC1 polymorphisms and cervical cancer risk: an updated meta-analysis. Tumour Biol. 2014; 35:1221–1231.
45. Li Y, Liu F, Tan SQ, Wang Y, Li SW. X-ray repair cross-complementing group 1 (XRCC1) genetic polymorphisms and cervical cancer risk: a huge systematic review and meta-analysis. PLoS One. 2012; 7:e44441.
46. Zhu BT, Conney AH. Functional role of estrogen metabolism in target cells: review and perspectives. Carcinogenesis. 1998; 19:1–27.
47. Tudek B. Base excision repair modulation as a risk factor for human cancers. Mol Aspects Med. 2007; 28:258–275.
48. Higgins JP, Thompson SG. Quantifying heterogeneity in a meta-analysis. Stat Med. 2002; 21:1539–1558.
49. DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986; 7:177–188.
50. Zhong JH, Zhang ZM, Li LQ. mEH Tyr113His polymorphism and the risk of ovarian cancer development. J Ovarian Res. 2013; 6:40.
51. Begg CB, Mazumdar M. Operating characteristics of a rank correlation test for publication bias. Biometrics. 1994; 50:1088–1101.

