INTRODUCTION
Chronic lymphocytic leukemia (CLL) is a heterogeneous disease, running an indolent course in some patients and a clinically aggressive course in others [1, 2]. Therefore, in clinical practice, prediction of outcome and response to treatment is important in an era in which several chemoimmunotherapy combinations and effective mechanism-driven treatments are available [3–5].
Prognostic/predictive factors include advanced stage [6], positivity for CD38, ZAP70 and CD49d [7–9], the unmutated configuration of the variable region of the immunoglobulin heavy chain (IGHV) gene [7] and specific cytogenetic lesions revealed by fluorescent in situ hybridization (FISH) [10]. Recent studies also demonstrated the independent negative prognostic impact of mutations of several genes, including TP53, NOTCH1 and SF3B1 [11–15].
Next generation sequencing (NGS) techniques documented that previously unidentified genes may be mutated in CLL possibly promoting disease progression and drug resistance [16–19]. By NGS it was also demonstrated that TP53 mutated subclones (< 15–20% of the cells) may confer the same negative prognosis as major clones (> 15–20% of the cells) detected by conventional sequencing techniques (i.e. Sanger sequencing) [20–22].
Comprehensive prognostic indexes including clinical and biological parameters were published [23, 24] and a recent systematic review [25] recommended IGHV and FISH analyses as useful tests for all newly diagnosed CLL patients as they may identify those 40–50% of cases with unfavorable prognosis based on the presence of an unmutated IGHV gene and/or 11q22/17p13 deletion [25].
Recently, karyotype aberrations were shown to represent a strong prognostic factor [26–28]. Moreover, the complex karyotype emerged as an independent predictor of inferior time to first treatment (TTFT) and shorter overall survival (OS) in patients investigated at diagnosis [29] or at disease progression [30] and in relapsed/refractory CLL treated with ibrutinib [31].
We therefore investigated whether an extended genetic characterization, including karyotype analysis using novel mitogens and mutational screening by NGS, could predict outcome in high-risk CLL patients. By correlating genetic data with clinical and biological parameters we showed that karyotype represents an independent prognostic factor for TTFT, OS and time to chemorefractoriness (TTCR).
RESULTS
Patients, mutational analysis and cytogenetic data
The clinical and biological characteristics of 101 high-risk CLL patients are presented in Table 1. Cytogenetic analysis showed clonal karyotype aberrations in 88/101 (87.1%) of the cases (Supplementary Table 1), a favorable karyotype in 30 (29.7%) patients, an intermediate karyotype in 36 (35.6%) and an unfavorable karyotype in 34 (33.7%) cases. A complex karyotype was present in 21 (20.8%) patients. Recurring aberrations included: 14q deletions in 12 cases, 6q deletions in 4 and 7q deletions in 3. A balanced translocation was observed in 12 patients, while 6 cases presented unbalanced translocations.
Table 1: Clinical and biological characteristics of the 101 CLL patients
Variable |
|
|---|---|
Age, median yrs (range) |
65.6 (38.4–89.9) |
Sex m/f |
63/38 |
Binet Stage a/b/c |
76/17/8 |
CD38 neg/pos |
41/60 |
ZAP70 neg/pos |
61/27 |
Normal FISH yes/no |
32/69 |
13q14 deletion yes/no |
52/49 (20/81 hierarchical) |
Trisomy 12 yes/no |
23/78 (21/80 hierarchical) |
11q22 deletion yes/no |
20/81 |
17p13 deletion yes/no |
8/93 |
FISH fav/int/unfav |
52/21/28 |
karyotype fav/int/unfav |
30/36/34 |
Complex karyotype yes/no |
21/79 |
IGHV mut/unmut |
9/92 |
Mutated patients by NGS no/yes |
45/56 |
N. of mutations by NGS 0/1/2/3/4 |
45/30/16/7/3 |
TP53 disruption* yes/no |
19/82 |
Chemotherapy yes/no |
63/38 |
Chemorefractory yes/no |
26/37 |
Lines of therapy 1/> 1 |
37/26 |
*17p13 deletion and/or TP53 mutation.
Abbreviations: f, female; fav, favorable; int, intermediate; m, male; mut, mutated; neg, negative; pos, positive; unfav, unfavorable; unmut, unmutated; yrs, years.
95 somatic mutations were found in 56/101 (55.4%) cases; 80 missense mutations, 5 nonsense mutations and 10 frameshit deletions. Mutations were detected with a frequency ranging from 5.0 to 96.7% of the reads. 16 cases (15.8%) showed mutations in the TP53 gene, 11 (10.9%) in the NOTCH1 gene, 11 (10.9%) in the SF3B1 gene, 8 (7.9%) in the ATM gene, 5 (4.9%) in the BIRC3 gene, 5 (4.9%) in the PTEN gene, 4 (4.0%) in the MYD88 gene, 4 (4.0%) in the BRAF gene, 4 (4.0%) in the POT1 gene, and 18 (17.8%) cases in the remaining 11 genes (Supplementary Tables 2 and 3). 26/56 (46.4%) mutated patients presented two or more mutations.
Correlations between mutational status, cytogenetic findings and clinical and biological parameters
The presence of gene mutations did not correlate with age, sex, Binet stage and CD38 positivity, while a significant association was found with unmutated IGHV status (p = 0.040) and a complex karyotype (p = 0.047; Tables 2 and 3). A complex karyotype (Table 3) was also associated with unfavorable FISH (p < 0.001) and TP53 disruption (p = 0.012). A trend for a significant association was observed with the presence of ≥ 2 mutations (p = 0.061) and with TP53 and ATM mutations (p = 0.080 and 0.058, respectively). A positive correlation was found between unfavorable FISH and unfavorable karyotype (p < 0.001). 11q22 deletions by FISH were never found in association with TP53 mutations or with TP53 disruption. TP53 disruption was associated with the presence of ≥ 2 mutations (p = 0.001) and mutated IGHV status (p < 0.001), the latter association being accounted for by the selection criteria adopted in this analysis, which included patients with unmutated IGHV and TP53 lesions. An overview of the distribution of mutations and principal biological findings according to cytogenetic results is presented in Supplementary Figure 1.
Table 2: Correlations between clinical biological parameters and molecular and molecular cytogenetic results
NGS mutations |
11q22 deletion by FISH |
TP53 disruption* |
|||||||
|---|---|---|---|---|---|---|---|---|---|
variable |
yes |
no |
P |
No |
yes |
P |
No |
yes |
P |
Age < 65/ > = 65 years |
30/26 |
26/19 |
0.692 |
43/38 |
13/7 |
0.453 |
46/36 |
10/9 |
0.803 |
Sex m/f |
33/23 |
30/15 |
0.536 |
51/30 |
12/8 |
0.802 |
51/31 |
12/7 |
1.000 |
Binet stage a/b-c |
43/13 |
33/12 |
0.817 |
62/19 |
14/6 |
0.569 |
62/20 |
14/5 |
1.000 |
CD38 neg/pos |
23/33 |
18/27 |
1.000 |
33/48 |
8/12 |
0.580 |
31/51 |
10/9 |
0.302 |
FISH fav-int/unfav |
39/17 |
34/11 |
0.655 |
73/8 |
0/20 |
< 0.001 |
62/20 |
11/8 |
0.156 |
IGHV mut/unmut |
8/48 |
1/44 |
0.040 |
8/73 |
1/19 |
0.684 |
1/81 |
8/11 |
< 0.001 |
TP53 disruption no/yes |
na |
na |
- |
62/19 |
20/0 |
0.012 |
na |
na |
na |
Muts by NGS no/yes |
na |
na |
- |
36/45 |
9/11 |
1.000 |
43/39 |
9/10^ |
0.800 |
n. of muts by NGS no/1/≥2 |
na |
na |
- |
36/23/22 |
9/7/4 |
0.730 |
43/23/16 |
2/7/10 |
0.001 |
TP53 WT/mut |
na |
na |
- |
65/16 |
20/0 |
0.037 |
82/0 |
3/16 |
< 0.001 |
SF3B1 WT/mut |
na |
na |
- |
73/8 |
17/3 |
0.452 |
74/8 |
16/3 |
0.429 |
NOTCH1 WT/mut |
na |
na |
- |
73/8 |
17/3 |
0.452 |
72/10 |
18/1 |
0.685 |
ATM WT/mut |
na |
na |
- |
76/5 |
17/3 |
0.192 |
74/8 |
19/0 |
0.346 |
BIRC3 WT/mut |
na |
na |
- |
77/4 |
19/1 |
1.000 |
79/3 |
17/2 |
0.236 |
Others WT/mut |
na |
na |
- |
60/21 |
15/5 |
1.000 |
61/21 |
14/5 |
1.000 |
*TP53 disruption: 17p13 deletion by FISH and/or TP53 mutation by NGS.
^TP53 mutations not included.
Abbreviations: f, female; fav, favorable; int, intermediate; m, male; mut, mutated; na; not assessed; neg, negative; pos, positive; unfav, unfavorable; unmut, unmutated; yrs, years.
Table 3: Correlations between clinical biologic parameters and cytogenetic findings
Karyotype abnormality |
Complex karyotype |
|||||
|---|---|---|---|---|---|---|
variable |
Fav-int |
Unfav |
P |
No |
Yes |
P |
Age < 65 / > = 65 years |
36/30 |
19/15 |
1.000 |
46/23 |
9/12 |
0.227 |
Sex m/f |
41/25 |
22/12 |
0.831 |
51/28 |
12/9 |
0.614 |
Binet stage a/b-c |
52/14 |
23/11 |
0.234 |
61/18 |
14/7 |
0.396 |
CD38 neg/pos |
26/40 |
14/20 |
1.000 |
33/46 |
7/14 |
0.618 |
FISH fav-int/unfav |
65/1 |
8/26 |
< 0.001 |
64/14 |
8/13 |
< 0.001 |
IGHV mut/unmut |
5/61 |
3/31 |
1.000 |
6/73 |
2/19 |
0.673 |
TP53 disruption no/yes |
57/9 |
25/9 |
0.168 |
69/10 |
13/8 |
0.021 |
Mutations s by NGS no/yes |
33/33 |
12/22 |
0.204 |
40/39 |
5/16 |
0.047 |
n. of mutations by NGS no/1/≥ 2 |
33/16/17 |
12/14/8 |
0.201 |
40/20/19 |
5/10/6 |
0.061 |
TP53 WT/mut |
57/9 |
28/6 |
0.572 |
70/9 |
15/6 |
0.080 |
SF3B1 WT/mut |
59/7 |
31/3 |
1.000 |
72/7 |
18/3 |
0.434 |
NOTCH1 WT/mut |
60/6 |
29/5 |
0.502 |
70/9 |
19/2 |
1.000 |
ATM WT/mut |
63/3 |
29/5 |
0.117 |
75/4 |
17/4 |
0.058 |
BIRC3 WT/mut |
63/3 |
32/2 |
1.000 |
75/4 |
20/1 |
1.000 |
Others WT/mut |
49/17 |
25/9 |
1.000 |
59/20 |
15/6 |
0.480 |
Abbreviations: f, female; fav, favorable; int, intermediate; m, male; mut, mutated; na; not assessed; neg, negative; pos, positive; unfav, unfavorable; unmut, unmutated; yrs, years.
Correlations between mutational status, cytogenetic findings and clinical outcome
The median follow-up for these 101 CLL patients was 41 months. In univariate analysis (Table 4), a worse TTFT was associated with advanced Binet stage (p < 0.001), a complex karyotype (p < 0.001) and an unfavorable karyotype (p = 0.001; Figure 1A). No correlations were observed between TTFT and gene mutations.
Table 4: Univariate analysis for TTFT and OS
TTFT |
OS |
||||
|---|---|---|---|---|---|
Variable |
N pts |
HR (CI 95%) |
P |
HR (CI 95%) |
P |
Binet stage b-c vs a |
25 vs 76 |
2.441 (1.851–3.218) |
< 0.001 |
1.457 (1.021–2.080) |
0.038 |
CD38 pos vs neg |
60 vs 41 |
1.490 (0.884–2.512) |
0.134 |
0.931 (0.466–1.859) |
0.839 |
IGHV unmut vs mut |
92 vs 9 |
2.169 (0.774–6.080) |
0.141 |
1.596 (0.382–6.673) |
0.522 |
11q22 deletion yes vs no |
20 vs 81 |
1.350 (0.711–2.561) |
0.359 |
0.946 (0.389–2.302) |
0.903 |
TP53 disruption yes/no |
19 vs 82 |
1.340 (0.733–2.452) |
0.342 |
2.419 (1.145–5.111) |
0.021 |
Complex karyotype yes vs no |
21 vs 79 |
3.024 (1.705–5.362) |
< 0.001 |
3.364 (1.596–7.091) |
0.001 |
Mutations by NGS yes/no |
56 vs 45 |
0.662 (0.395–1.109) |
0.117 |
0.694 (0.341–1.414) |
0.315 |
Karyotype abnormalities unfav vs fav-int |
34 vs 66 |
1.611 (1.228–2.112) |
0.001 |
1.562 (1.099–2.219) |
0.013 |
TP53 mut vs wt |
16 vs 85 |
1.107 (0.571–2.149) |
0.763 |
1.612 (0.697–3.728) |
0.264 |
SF3B1 mut vs wt |
11 vs 90 |
1.120 (0.753–1.666) |
0.575 |
1.210 (0.666–2.195) |
0.532 |
NOTCH1 mut vs wt |
11 vs 90 |
0.739 (0.519–1.050) |
0.096 |
0.744 (0.476–1.164) |
0.196 |
ATM mut vs wt |
8 vs 93 |
0.716 (0.466–1.100) |
0.127 |
0.810 (0.479–1.370) |
0.432 |
BIRC3 mut vs wt |
5 vs 96 |
1.287 (0.719–2.305) |
0.396 |
1.104 (0.538–2.269) |
0.787 |
OTHERS mut vs wt |
26 vs 75 |
0.958 (0.725–1.267) |
0.766 |
1.126 (0.741–1.710) |
0.578 |
A poorer OS (Table 4), was associated with advanced Binet stage (p = 0.038), TP53 disruption (p = 0.021), complex karyotype (p = 0.001) and an unfavorable karyotype (p = 0.013; Figure 1B).
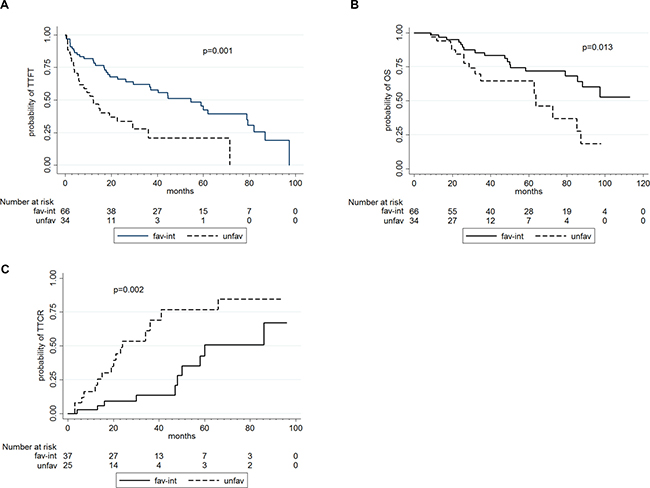
Figure 1: TTFT (in A), OS (in B) and TTCR (in C) according to karyotype abnormalities.
In univariate analysis (Table 5), a shorter TTCR correlated with unmutated IGHV status, (p = 0.036), TP53 disruption (p < 0.001), a complex karyotype (p = 0.004), an unfavorable karyotype (p = 0.002; Figure 1C) and TP53 mutations (p = 0.013).
Table 5: Univariate analysis for TTCR
TTCR |
|||
|---|---|---|---|
Variable |
N pts |
HR (CI 95%) |
P |
Binet stage b-c vs a |
24 vs 39 |
1.159 (0.783–1.715) |
0.463 |
CD38 pos vs neg |
40 vs 23 |
0.840 (0.370–1.904) |
0.676 |
IGHV unmut vs mut |
58 vs 5 |
0.148 (0.037–0.899) |
0.036 |
11q22 deletion yes vs no |
12 vs 51 |
1.409 (0.590–3.364) |
0.441 |
TP53 disruption yes/no |
15 vs 48 |
6.021 (2.366–15.320) |
< 0.001 |
Complex karyotype yes vs no |
18 vs 44 |
3.153 (1.429–6.954) |
0.004 |
Mutations by NGS yes/no |
40 vs 23 |
0.717 (0.311–1.653) |
0.435 |
Karyotype abnormalities unfav vs fav-int |
25 vs 37 |
1.864 (1.250–2.779) |
0.002 |
TP53 mut vs wt |
12 vs 51 |
3.484 (1.297–9.355) |
0.013 |
SF3B1 mut vs wt |
7 vs 56 |
1.159 (0.271–4.952) |
0.842 |
NOTCH1 mut vs wt |
9 vs 54 |
1.222 (0.419–3.561) |
0.713 |
ATM mut vs wt |
6 vs 57 |
1.107 (0.328–3.734) |
0.870 |
BIRC3 mut vs wt |
3 vs 60 |
1.144 (0.263–4.982) |
0.858 |
OTHERS mut vs wt |
18 vs 45 |
0.696 (0.309–1.566) |
0.381 |
Lines of therapy 1 vs > 1 |
37 vs 26 |
1.062 (0.437–2.580) |
0.894 |
By multivariate analysis (Table 6) an advanced Binet stage (p < 0.001) and an unfavorable karyotype (p = 0.001) predicted a shorter TTFT while TP53 disruption (p = 0.019) and an unfavorable karyotype (p = 0.028) were associated with a worse OS. A shorter TTCR (Table 6) was predicted, in multivariate analysis, by TP53 disruption (p = 0.001) and unfavorable cytogenetics (p = 0.025).
Table 6: Multivariate analysis for TTFT, OS and TTCR
Variable |
After bootstrapping |
|||||
|---|---|---|---|---|---|---|
TTFT |
HR |
CI |
p |
CI |
p |
|
Binet stage b-c vs a |
2.457 |
1.848–3.264 |
< 0.001 |
1.788–3.374 |
< 0.001 |
|
Karyotype abnormalities unfav vs fav-int |
1.630 |
1.231–2.159 |
0.001 |
1.241–2.141 |
< 0.001 |
|
OS |
||||||
Binet stage b-c vs a |
1.414 |
0.978–2.049 |
0.065 |
0.891–2.246 |
0.142 |
|
TP53 disruption yes vs no |
3.029 |
1.406–6.527 |
0.005 |
1.200–7.648 |
0.019 |
|
Karyotype abnormalities unfav vs fav-int |
1.512 |
1.046–2.184 |
0.028 |
1.031–2.218 |
0.034 |
|
TTCR |
||||||
IGHV unmut vs mut |
0.583 |
0.113–3.018 |
0.522 |
0.321–8.060 |
0.960 |
|
TP53 disruption yes vs no |
5.049 |
1.820–14.009 |
0.002 |
1.894–13.458 |
0.001 |
|
Karyotype abnormalities unfav vs fav-int |
1.761 |
1.171–2.649 |
0.007 |
1.072–2.894 |
0.025 |
|
The following variables were included in multivariate analysis:
TTFT: Binet stage and karyotype abnormalities.
OS: Binet stage, TP53 disruption and karyotype abnormalities.
TTCR: IGHV, TP53 disruption and karyotype abnormalities.
When in the model of multivariate analysis we introduced the complex karyotype instead of karyotypic abnormalities, the complex karyotype was significantly associated with a worse TTFT (p < 0.001) and an inferior OS (p = 0.024) while a trend for significance was observed for a shorter TTCR (p = 0.077; Supplementary Table 4).
By combining TP53 disruption status and cytogenetic results, 9 patients showed the coexistence of TP53 disruption and unfavorable karyotype: this subset of very high-risk patients presented a median TTFT of 5.5 months, a median OS of 28.7 months and a median TTCR of 15 months (Table 7, Figure 2).
Table 7: Median TTFT, OS and TTCR according to TP53 disruption status and karyotype abnormalities
n. pts |
TTFT* |
OS* |
N pts |
TTCR* |
||||
|---|---|---|---|---|---|---|---|---|
Median months |
se |
Median months |
se |
Median months |
se |
|||
No TP53 disruption & fav-int karyotype |
57 |
54.5 |
11.1 |
NR |
- |
32 |
86.0 |
17.4 |
TP53 disruption & fav-int karyotype |
9 |
97.3 |
1.3 |
NR |
- |
5 |
30.0 |
6.1 |
No TP53 disruption & unfav karyotype |
25 |
22.6 |
9.2 |
72.5 |
14.5 |
16 |
36.0 |
2.7 |
TP53 disruption & unfav karyotype |
9 |
5.5 |
2.2 |
28.7 |
6.2 |
9 |
15.0 |
5.2 |
*P < 0.0001 for the comparison by log-rank test.
Abbreviations: fav: favorable; int: intermediate; NR: not reached; se: standard error; unfav: unfavorable.
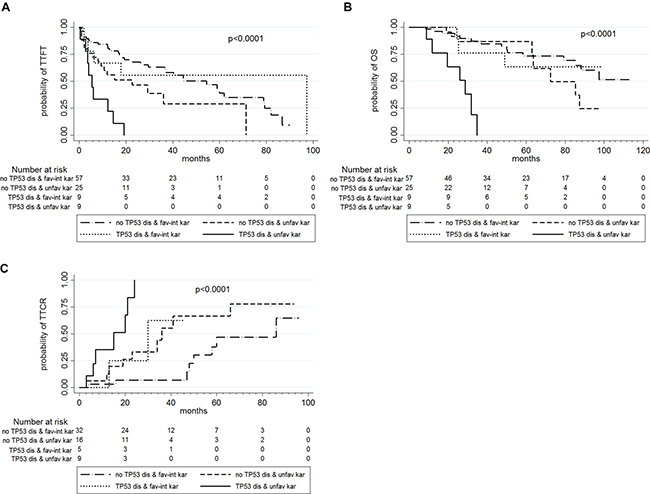
Figure 2: TTFT (in A), OS (in B) and TTCR (in C) according to TP53 disruption status and karyotype abnormalities.
In patients with favorable or intermediate cytogenetic findings (Supplementary Table 5), TP53 mutations correlated with a shorter TTCR (p = 0.037; Supplementary Figure 2) while in patients without TP53 disruption or complex karyotype (Supplementary Table 6), NOTCH1 mutations predicted a shorter TTFT and an inferior OS (p = 0.008 and 0.016, respectively; Supplementary Figures 3 and 4). A worse TTFT was also associated with the presence of ≥ 2 gene mutations (p = 0.038).
DISCUSSION
IGHV status and FISH results represent robust prognostic tests and their inclusion in the standard prognostic work up of newly diagnosed and/or previously untreated CLL was recently recommended [25]. Indeed unmutated IGHV status and 11q22 deletion were recognized as independent predictors of an unfavorable outcome [23, 24] while 17p13 deletion was aggregated with TP53 mutations in a category that received the highest risk score in the international prognostic index (IPI) of CLL [24]. Thus, based on IGHV status, FISH results and TP53 mutational analysis it is possible to identify a category of CLL patients with poor prognosis, representing approximately 50% of all CLL cases [13].
Karyotype analysis was reinforced as an important variable in CLL risk assessment [29, 31, 32, 33] though it is relatively time consuming and needs standardization of culture conditions [34]. Because few data are available on cytogenetic and molecular findings in high-risk CLL, we perform an extensive genetic characterization of 101 CLL with unmutated IGHV and/or 11q22 deletion and/or TP53 disruption. These patients were diagnosed and followed at a single center for a median of 41 months over the last 10 years. Notably, our center has a > 90% capture of incident CLL cases in our region allowing for meaningful analyses of TTFT, TTCR and OS in a real world scenario.
In this study, cytogenetic analysis was performed following DSP30/IL2 stimulation, thus reducing the rate of cytogenetic failure [28, 34]. A complex karyotype was present at diagnosis in 20.8% of patients, while a favorable karyotype was observed in 29.7% of the cases. Similar data were observed in the CLL 11 trial when karyotyping was performed before treatment and chromosomal aberrations were found in 68.8% of 154 patients, with 19.5% of the cases showing a complex karyotype [30]. These figures, as expected, are higher than those observed in unselected series of patients that included also low-risk CLL and cases with stable disease [29, 33].
We performed mutational screening with the Ion Torrent PGM, a very sensitive NGS platform, allowing for multiplexing of samples and gene targets in one experimental setup thus resulting in higher speed of analysis and lower costs [35]. Parallel sequencing of exonic regions in 20 CLL-related genes showed gene mutations in 56/101 (55.4%) cases by using a 5% cut off. Mutations were detected with a frequency ranging from 5.0 to 96.7% of the reads, showing cases with major clones that represent early leukemogenetic events and cases with minor clones that represent late-appearing aberrations [36, 37]. In this series, the frequency of mutations involving TP53, NOTCH1, SF3B1, ATM and BIRC3 genes (15.8, 10.9%, 10.9%, 7.9% and 4.9%, respectively) reflected the high-risk genetic features of our patients. Similar incidences for these mutations were reported in patients enrolled in clinical trials at disease progression [13]. The frequency of mutations involving the other investigated genes was in line with published data using whole exome sequencing [38–41]. Interestingly, we observed that 25.7% of the cases presented more than one mutation. In the CLL11 trial, NGS analysis revealed mutations in 42 out of 85 analyzed genes, with 76.4% and 42.2% of the patients presenting one or ≥ 2 mutations, respectively [30].
Moreover, an unfavorable karyotype, defined by the presence of a complex karyotype and/or 11q22 and/or 17p13 deletion, was independently associated with a worse OS and TTFT, while TP53 disruption was associated with poorer OS. It is worth noting, however, that our patients may have received different lines of treatment including various chemotherapeutic agents, with or without rituximab and BTK inhibitors. These observations need therefore to be validated in larger series of patients and in clinical trials with homogeneous treatments. Interestingly, in the CLL11 trial a complex karyotype emerged as an independent negative prognostic factor for OS after front-line therapy [30] while in relapsed/refractory CLL treated with ibrutinib-based regimens, a complex karyotype was stronger than 17p13 deletion in predicting an inferior outcome [31]. A complex karyotype was also found to be associated with unmutated IGHV genes and aberrations of chromosome 17p and was identified as an independent prognostic factor for shorter TTFT in a series of 1001 cases [33]. Likewise, an unfavorable karyotype predicted a shorter OS and represented the strongest prognostic parameter for disease progression in unselected patients studied at diagnosis [28, 29].
The mechanisms responsible for the unfavorable prognostic significance of the complex karyotype remain to be investigated, though in CLL it was shown that telomere shortening was associated with genetic complexity [42].
We also assessed the impact of karyotyping and gene mutations on TTCR. While confirming the negative role of TP53 disruption [36, 37], we showed, for the first time, that patients with unfavorable cytogenetics had a shorter TTCR. The identification of patients that are likely to develop chemoresistance is very important, given the possibility to use alternative effective BCR-target treatments [43, 44].
Nor the presence of gene mutations, nor single gene mutations correlated with TTFT, OS and TTCR, with the notable exception of TP53 mutations. This is not surprising if we consider that the high-risk genetic profile of our patients may mirror a genetic instability or dysregulation [45] that may overcome the prognostic impact of single gene mutations [12, 13].
Interestingly our data showed that high-risk CLL patients can be further sub-classified into four different groups, based on TP53 disruption status and karyotype. Those patients with both unfavorable cytogenetics and TP53 disruption presented a dismal clinical course (median TTFT, OS and TTCR of 5.5, 28.7 and 15.0 months, respectively). This observation may justify a specific treatment strategy for this subset of patients [46]. By contrast, there are cases with TP53 disruption and favorable or intermediate cytogenetics that may display a less aggressive course of the disease [47]. Recently, it has been shown that also the percentage of positive cells by FISH may help to refine the prognosis of high-risk CLL patients [48]. By contrast, mutational analysis may refine the prognosis of high-risk patients without unfavorable or complex karyotype or TP53 disruption as shown by the negative prognostic impact of NOTCH1 mutations and of the coexistence of two or more mutations [49].
In conclusion, we described the results of a comprehensive analysis of chromosomal aberrations and gene mutations in high-risk CLL, providing the first demonstration that the cytogenetic profile was independently associated with a shorter TTFT, OS and TTCR. The introduction of this technique in future CLL trials seems warranted to identify those high-risk patients that could be considered as ideal candidates for consolidation treatments or novel treatment combinations [50–53]. Larger series of homogenously treated patients and with longer follow-up could confirm this observation.
MATERIALS AND METHODS
Patients
One hundred-one untreated CLL patients with unmutated IGHV gene and/or 17p13/11q22 deletion by FISH and/or mutated TP53 gene (here referred to as high-risk CLL) were included in this analysis. TP53 disruption was defined by the presence of 17p13 deletion by FISH and/or TP53 mutation by NGS. These patients belong to a consecutive series of 200 patients diagnosed and followed between 2007 and 2014 [49]. All patients were diagnosed according to NCI criteria [54]. Only patients with a Matutes immunophenotypic score [55] ≥ 3 (i.e., typical CLL) were included. CD38 and ZAP70 were tested on peripheral blood (PB) cells, as described [56]. When needed, mantle cell lymphoma was excluded by cyclin D1 evaluation. The study was approved by the local ethics committee. Indications for treatment included: increased white blood cell count with < 6 month lymphocyte doubling time, anemia or thrombocytopenia due to bone marrow infiltration or autoimmune phenomena not responding to steroids, and clinically significant disease progression in the Binet staging system. Fludarabine or bendamustine containing regimens, with or without rituximab were used as first-line treatment in fit patients; chlorambucil with or without rituximab was used in elderly and/or unfit patients according to the treatment policy adopted at our center. Since 2015, ibrutinib or idelalisib plus rituximab were offered to relapsed/refractory patients.
Cytogenetic and FISH analyses
Interphase FISH was performed on PB samples obtained at diagnosis using probes for the following regions: 13q14, 12q13, 11q22/ATM, 17p13/TP53 (Vysis/Abbott Co, Downers Grove, IL) as described [57]. Each patient was categorized into a FISH risk group according to the following classification: favorable group (isolated 13q14 deletion or absence of FISH aberrations), intermediate group (trisomy 12); unfavorable group (deletions of 11q22 or of 17p13).
Cytogenetic analysis was performed on the same samples used for FISH analysis using CpG-oligonucleotide DSP30 (2 μmol/l TibMolBiol Berlin, Germany) plus IL2 (100 U/ml Stem Cell Technologies Inc., Milan, Italy) as described [57]. Whenever possible, at least 20 metaphases were karyotyped and each patient was categorized into a cytogenetic risk group according to the following classification: favorable group (isolated 13q14 deletion or normal karyotype), unfavorable group (deletions of 11q22 or 17p13, or complex karyotype, ie, at least three chromosome aberrations); intermediate group (all other karyotypic abnormalities).
IGHV analysis
IGHV genes were amplified from genomic DNA and sequenced according to standard methods with the cut-off of 98% homology to the germline sequence to discriminate between mutated (< 98%) and unmutated (≥ 98%) cases, as reported [58].
Next generation sequencing
NGS analysis was performed on the same samples used for FISH and cytogenetic analyses. In all samples, the percentage of CLL cells was over 90% as assessed by flow cytometry analysis. Agilent HaloPlex Target Enrichment kit (Agilent Technologies, Santa Clara, CA, USA) was used to produce libraries of exonic regions from 20 genes (ATM, BIRC3, BRAF, CDKN2A, PTEN, CDH2, DDX3X, FBXW7, KIT, KLHL6, KRAS, MYD88, NOTCH1, NRAS, PIK3CA, POT1, SF3B1, TP53, XPO1, ZMYM3) starting from genomic DNA from PB samples, according to HaloPlex Target Enrichment System (Agilent Technologies, Santa Clara, CA, USA). Diluted libraries were linked to Ion Sphere Particles, clonally amplified in an emulsion PCR and enriched using Ion OneTouch emulsion PCR System (Life technologies, Foster City, CA, USA). Exon-enriched DNA was precipitated with magnetic beads coated with streptavidin. Enriched, template-positive Ion Sphere Particles were loaded in one Ion chip and sequenced using Ion Torrent PGM (Life technologies, Foster City, CA, USA). Sequencing data were aligned to the human reference genome (GRCh37). Data analysis and variants identification were performed using Torrent Suite 3.4 and Variant Caller plugin 3.4.4 (Life technologies, Foster City, CA, USA). To identify pathogenic variations, mutations that did not affect the protein coding regions (intronic, 3′ and 5′ UTR variations, silent exonic mutations and polymorphisms) were filtered out; insertions and deletions belonging to homopolymeric regions were removed, because sequencing error rate is high in these regions [49].
Statistical analysis
The Fischer’s exact test was applied for categorical variables. TTFT was calculated as the interval between diagnosis and the start of first line treatment. OS was calculated from the date of diagnosis until death due to any cause or until the last patient follow-up. Refractory disease was defined as treatment failure (stable disease, nonresponse, progressive disease, or death from any cause) or disease progression within 6 months from antileukemic therapy [54]. TTCR was measured from date of first-line treatment until date of refractoriness to fludarabine/bendamustine-based regimens, date of alkylator refractoriness in patients who had never been exposed to fludarabine/bendamustine, death, or last follow-up [59]. Survival curves were compared by the log-rank test. Proportional hazards regression analysis was used to identify the significant independent prognostic variables on TTFT. The stability of the Cox model was internally validated using bootstrapping procedures [11]. Statistical analysis was performed using Stata 14.0 (Stata Corp, College Station, TX).
CONFLICTS OF INTEREST
The authors declare that they have no potential conflicts of interests.
GRANT SUPPORT
Supported by FAR (Fondo di Ateneo per la Ricerca) 2012, 2013, 2014, 2016 of the University of Ferrara (GMR, AC), Programma Ricerca Regione Università 2007-2009 University of Ferrara (GMR, AC), PRIN 2008 (AC), Ricerca Finalizzata (AC, project RF-2011-02349712) and AIL (Associazione Italiana contro le Leucemie Linfomi e Mieloma, Ferrara). EV and ES are supported by AIL-Ferrara.
REFERENCES
1. Chiorazzi N, Rai KR, Ferrarini M. Chronic lymphocytic leukemia. N Engl J Med. 2005; 352:804–815.
2. Zenz T, Mertens D, Küppers R, Döhner H, Stilgenbauer S. From pathogenesis to treatment of chronic lymphocytic leukaemia. Nat Rev Cancer. 2010; 10:37–50.
3. Shanafelt TD. Predicting clinical outcome in CLL: how and why. Hematology Am Soc Hematol Educ Program. 2009: 421–429.
4. Cuneo A, Cavazzini F, Ciccone M, Daghia G, Sofritti O, Saccenti E, Negrini M, Rigolin GM. Modern treatment in chronic lymphocytic leukemia: impact on survival and efficacy in high-risk subgroups. Cancer Med. 2014; 3:555–564.
5. Hallek M. Chronic lymphocytic leukemia: 2015 Update on diagnosis, risk stratification, and treatment. Am J Hematol. 2015; 90:446–460.
6. Binet JL, Auquier A, Dighiero G, Chastang C, Piguet H, Goasguen J, Vaugier G, Potron G, Colona P, Oberling F, Thomas M, Tchernia G, Jacquillat C, et al. A new prognostic classification of chronic lymphocytic leukemia derived from a multivariate survival analysis. Cancer. 1981; 48:198–206.
7. Damle RN, Wasil T, Fais F, Ghiotto F, Valetto A, Allen SL, Buchbinder A, Budman D, Dittmar K, Kolitz J, Lichtman SM, Schulman P, Vinciguerra VP, et al. Ig V gene mutation status and CD38 expression as novel prognostic indicators in chronic lymphocytic leukemia. Blood. 1999; 94:1840–1847.
8. Wiestner A, Rosenwald A, Barry TS, Wright G, Davis RE, Henrickson SE, Zhao H, Ibbotson RE, Orchard JA, Davis Z, Stetler-Stevenson M, Raffeld M, Arthur DC, et al. ZAP70 expression identifies a chronic lymphocytic leukemia subtype with unmutated immunoglobulin genes, inferior clinical outcome, and distinct gene expression profile. Blood. 2003; 101:4944–4951.
9. Bulian P, Shanafelt TD, Fegan C, Zucchetto A, Cro L, Nückel H, Baldini L, Kurtova AV, Ferrajoli A, Burger JA, Gaidano G, Del Poeta G, Pepper C, et al. CD49d is the strongest flow cytometry-based predictor of overall survival in chronic lymphocytic leukemia. J Clin Oncol. 2014; 32:897–904.
10. Döhner H, Stilgenbauer S, Benner A, Leupolt E, Kröber A, Bullinger L, Döhner K, Bentz M, Lichter P. Genomic aberrations and survival in chronic lymphocytic leukemia. N Engl J Med. 2000; 343:1910–1916.
11. Rossi D, Rasi S, Spina V, Bruscaggin A, Monti S, Ciardullo C, Deambrogi C, Khiabanian H, Serra R, Bertoni F, Forconi F, Laurenti L, Marasca R, et al. Integrated mutational and cytogenetic analysis identifies new prognostic subgroups in chronic lymphocytic leukemia. Blood. 2013; 121:1403–1412.
12. Jeromin S, Weissmann S, Haferlach C, Dicker F, Bayer K, Grossmann V, Alpermann T, Roller A, Kohlmann A, Haferlach T, Kern W, Schnittger S. SF3B1 mutations correlated to cytogenetics and mutations in NOTCH1, FBXW7, MYD88, XPO1 and TP53 in 1160 untreated CLL patients. Leukemia. 2014; 28:108–117.
13. Baliakas P, Hadzidimitriou A, Sutton LA, Rossi D, Minga E, Villamor N, Larrayoz M, Kminkova J, Agathangelidis A, Davis Z, Tausch E, Stalika E, Kantorova B, et al. Recurrent mutations refine prognosis in chronic lymphocytic leukemia. Leukemia. 2015; 29:329–336.
14. Oscier DG1, Rose-Zerilli MJ, Winkelmann N, Gonzalez de Castro D, Gomez B, Forster J, Parker H, Parker A, Gardiner A, Collins A, Else M, Cross NC, Catovsky D, et al. The clinical significance of NOTCH1 and SF3B1 mutations in the UK LRF CLL4 trial. Blood. 2013; 121:468–75.
15. Stilgenbauer S, Schnaiter A, Paschka P, Zenz T, Rossi M, Döhner K, Bühler A, Böttcher S, Ritgen M, Kneba M, Winkler D, Tausch E, Hoth P, et al. Gene mutations and treatment outcome in chronic lymphocytic leukemia: results from the CLL8 trial. Blood. 2014; 123:3247–3254.
16. Guièze R, Wu CJ. Genomic and epigenomic heterogeneity in chronic lymphocytic leukemia. Blood. 2015; 126:445–453.
17. Rossi D, Bruscaggin A, Spina V, Rasi S, Khiabanian H, Messina M, Fangazio M, Vaisitti T, Monti S, Chiaretti S, Guarini A, Del Giudice I, Cerri M, et al. Mutations of the SF3B1 splicing factor in chronic lymphocytic leukemia: association with progression and fludarabine-refractoriness. Blood. 2011; 118:6904–6908.
18. Landau DA, Tausch E, Taylor-Weiner AN, Stewart C, Reiter JG, Bahlo J, Kluth S, Bozic I, Lawrence M, Böttcher S, Carter SL, Cibulskis K, Mertens D, et al. Mutations driving CLL and their evolution in progression and relapse. Nature. 2015; 526:525–530.
19. Puente XS, Pinyol M, Quesada V, Conde L, Ordóñez GR, Villamor N, Escaramis G, Jares P, Beà S, González-Díaz M, Bassaganyas L, Baumann T, Juan M, et al. Whole-genome sequencing identifies recurrent mutations in chronic lymphocytic leukaemia. Nature. 2011; 475:101–105.
20. Rossi D, Khiabanian H, Spina V, Ciardullo C, Bruscaggin A, Famà R, Rasi S, Monti S, Deambrogi C, De Paoli L, Wang J, Gattei V, Guarini A, et al. Clinical impact of small TP53 mutated subclones in chronic lymphocytic leukemia. Blood. 2014; 123:2139–2147.
21. Malcikova J, Stano-Kozubik K, Tichy B, Kantorova B, Pavlova S, Tom N, Radova L, Smardova J, Pardy F, Doubek M, Brychtova Y, Mraz M, Plevova K, et al. Detailed analysis of therapy-driven clonal evolution of TP53 mutations in chronic lymphocytic leukemia. Leukemia. 2015; 29:877–885.
22. Rasi S, Khiabanian H, Ciardullo C, Terzi-di-Bergamo L, Monti S, Spina V, Bruscaggin A, Cerri M, Deambrogi C, Martuscelli L, Biasi A, Spaccarotella E, De Paoli L, et al. Clinical impact of small subclones harboring NOTCH1, SF3B1 or BIRC3 mutations in chronic lymphocytic leukemia. Haematologica. 2016; 101:e135–138.
23. Pflug N, Bahlo J, Shanafelt TD, Eichhorst BF, Bergmann MA, Elter T, Bauer K, Malchau G, Rabe KG, Stilgenbauer S, Döhner H, Jäger U, Eckart MJ, et al. Development of a comprehensive prognostic index for patients with chronic lymphocytic leukemia. Blood. 2014; 124:49–62.
24. International CLL-IPI working group. An international prognostic index for patients with chronic lymphocytic leukaemia (CLL-IPI): a meta-analysis of individual patient data. Lancet Oncol. 2016; 17:779–790.
25. Parikh SA, Strati P, Tsang M, West CP, Shanafelt TD. Should IGHV status and FISH testing be performed in all CLL patients at diagnosis? A systematic review and meta-analysis. Blood. 2016; 127:1752–1760.
26. Cavazzini F, Hernandez JA, Gozzetti A, Russo Rossi A, De Angeli C, Tiseo R, Bardi A, Tammiso E, Crupi R, Lenoci MP, Forconi F, Lauria F, Marasca R, et al. Chromosome 14q32 translocations involving the immunoglobulin heavy chain locus in chronic lymphocytic leukaemia identify a disease subset with poor prognosis. Br J Haematol. 2008; 142:529–537.
27. Cuneo A, Rigolin GM, Bigoni R, De Angeli C, Veronese A, Cavazzini F, Bardi A, Roberti MG, Tammiso E, Agostini P, Ciccone M, Della Porta M, Tieghi A, et al. Chronic lymphocytic leukemia with 6q- shows distinct hematological features and intermediate prognosis. Leukemia. 2004; 18:476–483.
28. Rigolin GM, Cibien F, Martinelli S, Formigaro L, Rizzotto L, Tammiso E, Saccenti E, Bardi A, Cavazzini F, Ciccone M, Nichele I, Pizzolo G, Zaja F, et al. Chromosome aberrations detected by conventional karyotyping using novel mitogens in chronic lymphocytic leukemia with “normal” FISH: correlations with clinicobiologic parameters. Blood. 2012; 119:2310–2313.
29. Rigolin GM, del Giudice I, Formigaro L, Saccenti E, Martinelli S, Cavallari M, Lista E, Tammiso E, Volta E, Lupini L, Bassi C, Bardi A, Sofritti O, et al. Chromosome aberrations detected by conventional karyotyping using novel mitogens in chronic lymphocytic leukemia: Clinical and biologic correlations. Genes Chromosomes Cancer. 2015; 54:818–826.
30. Herling CD, Klaumünzer M, Rocha CK, Altmüller J, Thiele H, Bahlo J, Kluth S, Crispatzu G, Herling M, Schiller J, Engelke A, Tausch E, Döhner H, et al. Complex karyotypes, KRAS and POT1 mutations impact outcome in CLL after chlorambucil based chemo- or chemoimmunotherapy. Blood. 2016; 128:395–404.
31. Thompson PA, O’Brien SM, Wierda WG, Ferrajoli A, Stingo F, Smith SC, Burger JA, Estrov Z, Jain N, Kantarjian HM, Keating MJ. Complex karyotype is a stronger predictor than del(17p) for an inferior outcome in relapsed or refractory chronic lymphocytic leukemia patients treated with ibrutinib-based regimens. Cancer. 2015; 121:3612–3621.
32. Haferlach C, Dicker F, Schnittger S, Kern W, Haferlach T. Comprehensive genetic characterization of CLL: a study on 506 cases analysed with chromosome banding analysis, interphase FISH, IgHV status and immunophenotyping. Leukemia. 2007; 21:2442–2451.
33. Baliakas P, Iskas M, Gardiner A, Davis Z, Plevova K, Nguyen-Khac F, Malcikova J, Anagnostopoulos A, Glide S, Mould S, Stepanovska K, Brejcha M, Belessi C, et al. Chromosomal translocations and karyotype complexity in chronic lymphocytic leukemia: a systematic reappraisal of classic cytogenetic data. Am J Hematol. 2014; 89:249–255.
34. Dicker F, Schnittger S, Haferlach T, Kern W, Schoch C. Immunostimulatory oligonucleotide-induced metaphase cytogenetics detect chromosomal aberrations in 80% of CLL patients: A study of 132 CLL cases with correlation to FISH, IgHV status, and CD38 expression. Blood. 2006; 108:3152–3160.
35. Sutton LA, Sutton LA, Ljungström V, Mansouri L, Young E, Cortese D, Navrkalova V, Malcikova J, Muggen AF, Trbusek M, Panagiotidis P, Davi F, Belessi C, et al. Targeted next-generation sequencing in chronic lymphocytic leukemia: a high-throughput yet tailored approach will facilitate implementation in a clinical setting. Haematologica. 2015; 100:370–376.
36. Schnaiter A, Paschka P, Rossi M, Zenz T, Bühler A, Winkler D, Cazzola M, Döhner K, Edelmann J, Mertens D, Kless S, Mack S, Busch R, et al. NOTCH1, SF3B1, and TP53 mutations in fludarabine-refractory CLL patients treated with alemtuzumab: results from the CLL2H trial of the GCLLSG. Blood. 2013; 122:1266–1270.
37. Rossi D, Fangazio M, Rasi S, Vaisitti T, Monti S, Cresta S, Chiaretti S, Del Giudice I, Fabbri G, Bruscaggin A, Spina V, Deambrogi C, Marinelli M, et al. Disruption of BIRC3 associates with fludarabine chemorefractoriness in TP53 wild-type chronic lymphocytic leukemia. Blood. 2012; 119:2854–2862.
38. Wang L, Lawrence MS, Wan Y, Stojanov P, Sougnez C, Stevenson K, Werner L, Sivachenko A, DeLuca DS, Zhang L, Zhang W, Vartanov AR, Fernandes SM, et al. SF3B1 and other novel cancer genes in chronic lymphocytic leukemia. N Engl J Med. 2011; 365:2497–2506.
39. Fabbri G1, Rasi S, Rossi D, Trifonov V, Khiabanian H, Ma J, Grunn A, Fangazio M, Capello D, Monti S, Cresta S, Gargiulo E, Forconi F, et al. Analysis of the chronic lymphocytic leukemia coding genome: role of NOTCH1 mutational activation. J Exp Med. 2011; 208:1389–1401.
40. Quesada V, Conde L, Villamor N, Ordóñez GR, Jares P, Bassaganyas L, Ramsay AJ, Beà S, Pinyol M, Martínez-Trillos A, López-Guerra M, Colomer D, Navarro A, et al. Exome sequencing identifies recurrent mutations of the splicing factor SF3B1 gene in chronic lymphocytic leukemia. Nat Genet. 2011; 44:47–52.
41. Landau DA, Carter SL, Stojanov P, McKenna A, Stevenson K, Lawrence MS, Sougnez C, Stewart C, Sivachenko A, Wang L, Wan Y, Zhang W, Shukla SA, et al. Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell. 2013; 152:714–726.
42. Roos G, Kröber A, Grabowski P, Kienle D, Bühler A, Döhner H, Rosenquist R, Stilgenbauer S. Short telomeres are associated with genetic complexity, high-risk genomic aberrations, and short survival in chronic lymphocytic leukemia. Blood. 2008; 111:2246–2252.
43. Burger JA, Tedeschi A, Barr PM, Robak T, Owen C, Ghia P, Bairey O, Hillmen P, Bartlett NL, Li J, Simpson D, Grosicki S, Devereux S, McCarthy H, et al. Ibrutinib as Initial Therapy for Patients with Chronic Lymphocytic Leukemia. N Engl J Med 2015; 373:2425–2437
44. O’Brien SM, Lamanna N, Kipps TJ, Flinn I, Zelenetz AD, Burger JA, Keating M, Mitra S, Holes L, Yu AS, Johnson DM, Miller LL, Kim Y, et al. A phase 2 study of idelalisib plus rituximab in treatment-naïve older patients with chronic lymphocytic leukemia. Blood 2015; 126:2686–2694.
45. Dicker F, Herholz H, Schnittger S, Nakao A, Patten N, Wu L, Kern W, Haferlach T, Haferlach C. The detection of TP53 mutations in chronic lymphocytic leukemia independently predicts rapid disease progression and is highly correlated with a complex aberrant karyotype. Leukemia. 2009; 23:117–124.
46. Dreger P, Schetelig J, Andersen N, Corradini P, van Gelder M, Gribben J, Kimby E, Michallet M, Moreno C, Stilgenbauer S, Montserrat E; European Research Initiative on CLL (ERIC) and the European Society for Blood and Marrow Transplantation (EBMT). Managing high-risk CLL during transition to a new treatment era: stem cell transplantation or novel agents? Blood. 2014; 124:3841–3849.
47. Tam CS, Shanafelt TD, Wierda WG, Abruzzo LV, Van Dyke DL, O’Brien S, Ferrajoli A, Lerner SA, Lynn A, Kay NE, Keating MJ. De novo deletion 17p13.1 chronic lymphocytic leukemia shows significant clinical heterogeneity: the M. D. Anderson and Mayo Clinic experience. Blood. 2009; 114:957–964.
48. Hernández JÁ, Hernández-Sánchez M, Rodríguez-Vicente AE, Grossmann V, Collado R, Heras C, Puiggros A, Martín AÁ, Puig N, Benito R, Robledo C, Delgado J, González T, et al. A low frequency of losses in 11q chromosome is associated with better outcome and lower rate of genomic mutations in patients with chronic lymphocytic leukemia. PLoS One. 2015; 10:e0143073.
49. Rigolin GM, Saccenti E, Bassi C, Lupini L, Quaglia FM, Cavallari M, Martinelli S, Formigaro L, Lista E, ardi MA, Volta E, Tammiso E, Melandri A, et al. Extensive Next Generation Sequencing analysis in Chronic Lymphocytic Leukemia at diagnosis: clinical and biological correlations. J Hematol Oncol. 2016; 9:88.
50. Foà R, Del Giudice I, Cuneo A, Del Poeta G, Ciolli S, Di Raimondo F, Lauria F, Cencini E, Rigolin GM, Cortelezzi A, Nobile F, Callea V, Brugiatelli M, et al. Chlorambucil plus rituximab with or without maintenance rituximab as first-line treatment for elderly chronic lymphocytic leukemia patients. Am J Hematol 2014; 89:480–486.
51. Rossi D, Terzi-di-Bergamo L, De Paoli L, Cerri M, Ghilardi G, Chiarenza A, Bulian P, Visco C, Mauro FR, Morabito F, Cortelezzi A, Zaja F, Forconi F, et al. Molecular prediction of durable remission after first-line fludarabine-cyclophosphamide-rituximab in chronic lymphocytic leukemia. Blood. 2015; 126:1921–1924.
52. Strati P, Lanasa M, Call TG, Leis JF, Brander DM, LaPlant BR, Pettinger AM, Ding W, Parikh SA, Hanson CA, Chanan-Khan AA, Bowen DA, Conte M, et al. Ofatumumab monotherapy as a consolidation strategy in patients with previously untreated chronic lymphocytic leukaemia: a phase 2 trial. Lancet Haematol. 2016; 3:e407–414.
53. Stilgenbauer S, Eichhorst B, Schetelig J, Coutre S, Seymour JF, Munir T, Puvvada SD, Wendtner CM, Roberts AW, Jurczak W, Mulligan SP, Böttcher S, Mobasher M, et al. Venetoclax in relapsed or refractory chronic lymphocytic leukaemia with 17p deletion: a multicentre, open-label, phase 2 study. Lancet Oncol. 2016; 17:768–778.
54. Hallek M, Cheson BD, Catovsky D, Caligaris-Cappio F, Dighiero G, Döhner H, Hillmen P, Keating MJ, Montserrat E, Rai KR, Kipps TJ; International Workshop on Chronic Lymphocytic Leukemia.. Guidelines for the diagnosis and treatment of chronic lymphocytic leukemia: a report from the International Workshop on Chronic Lymphocytic Leukemia updating the National Cancer Institute-Working Group 1996 guidelines. Blood. 2008; 111:5446–5456.
55. Matutes E, Owusu-Ankomah K, Morilla R, Garcia Marco J, Houlihan A, Que TH, Catovsky D. The immunological profile of B-cell disorders and proposal of a scoring system for the diagnosis of CLL. Leukemia. 1994; 8:1640–1645.
56. Rigolin GM, Maffei R, Rizzotto L, Ciccone M, Sofritti O, Daghia G, Cibien F, Cavazzini F, Marasca R, Cuneo A. Circulating endothelial cells in patients with chronic lymphocytic leukemia: clinical-prognostic and biologic significance. Cancer. 2010; 116:1926–1937.
57. Bardi A, Cavazzini F, Rigolin GM, Tammiso E, Volta E, Pezzolo E, Formigaro L, Sofritti O, Daghia G, Ambrosio C, Rizzotto L, Abass AE, D’Auria F, et al. Employment of oligodeoxynucleotide plus interleukin-2 improves cytogenetic analysis in splenic marginal zone lymphoma. J Biomed Biotechnol. 2011; 2011:691493.
58. Rigolin GM, Saccenti E, Rizzotto L, Ferracin M, Martinelli S, Formigaro L, Cibien F, Cavallari M, Lista E, Daghia G, Sofritti O, Ciccone M, Cavazzini F, et al. Genetic subclonal complexity and miR125a-5p down-regulation identify a subset of patients with inferior outcome in low-risk CLL patients. Oncotarget. 2014; 5:140–149. doi: 10.18632/oncotarget.1382.
59. Rossi D, Cerri M, Deambrogi C, Sozzi E, Cresta S, Rasi S, De Paoli L, Spina V, Gattei V, Capello D, Forconi F, Lauria F, Gaidano G. The prognostic value of TP53 mutations in chronic lymphocytic leukemia is independent of Del17p13: implications for overall survival and chemorefractoriness. Clin Cancer Res. 2009; 15:995–1004.

