Introduction
Osteosarcoma is an extremely aggressive form of bone cancer which has a peak incidence in the second decade of life [1, 2]. Prior to systemic chemotherapy amputation was the only choice of treatment, and the high rate of pulmonary micrometastasis makes 5 year survival very low [3]. For the last 4 decades, neoadjuvant chemotherapy and limb salvage surgery have greatly improved both the survival and life quality of the patient [4]. However, despite intensive chemotherapy, the survival rate for high-grade osteosarcoma has improved little and almost 40% of osteosarcoma patients die of their disease [5].
Osteosarcoma cells within tumor tissue demonstrate significant heterogeneity with respect to tumorigenic and metastatic potential [6, 7]. The biological basis underlying this cellular heterogeneity has potential therapeutic implications. In contrast to the stochastic model, the cancer stem cell model predicts that heterogeneity within a tumor is derived from a hierarchal organization [8-11]. Cancer stem cells (CSCs) are thus thought to share important properties with normal tissue stem cells such as self-renewal and differentiation. In addition, cancer stem cells have been proposed to be responsible for metastasis and resistance to anticancer drugs [12, 13]. Therefore, the inadequacy of current treatments may result from the inability to effectively target the cancer stem cells within osteosarcoma.
Properties of normal stem cells have been widely used to identify cancer stem cells. There are several methods used to isolate osteosarcoma stem cells, including cell surface markers [14, 15], Hoechst effluxing (SP) [16], label retention (PKH26) [17], the Aldefluor assay [18] and promoter reporter assays [19]. For example, the mesenchymal stem cell markers CD117 and Stro-1 have been successfully applied to isolate osteosarcoma initiating cells associated with metastasis and drug resistance [14]. However, these methods have not identified a way to target these tumorigenic cellular subpopulations.
Human telomerase is a reverse transcriptase enzyme composed of a catalytic component, telomerase reverse-transcriptase (TERT) and a telomerase RNA component (TERC) [20]. In most human normal somatic cells, telomerase activity is undetectable, however, stem/progenitor cells in self-renewing tissues express telomerase. In addition, telomerase activity is detected in most malignant cancers, including osteosarcoma [21]. Given the important role of telomeres and telomerase in stem cells and cancer [22-24], we predicted a high expression of telomerase in cancer stem cells, and aimed to investigate its role in the cancer stem cell phenotype.
To test this, we stably transduced osteosarcoma cell lines with a lentiviral vector in which the human TERT promoter drives expression of green fluorescent protein (GFP). Based on high activity of the fluorescent reporter, we isolated a distinct subpopulation of cells with tumor-initiating properties from the human osteosarcoma cell lines. These cells show high drug-resistant and metastatic properties, and furthermore we demonstrate that telomerase inhibition is effective for eradicating these osteosarcoma stem-like cells.
Results
Osteosarcoma sphere cells are enriched for high telomerase activity
A proportion of osteosarcoma cells are capable of forming spheres when cultured in a serum free, anchorage-independent environment [25]. Spheres show high levels of staining for the osteosarcoma stem cell markers Stro-1 and CD117, and sphere-forming cells have greater capacity to initiate tumors, which supports their increased stem cell content [14]. To test whether stem cells have higher telomerase activity, we used the TRAP-ELISA assay and analysed hTERT mRNA expression levels in the cells in different culture conditions. We found telomerase activity to be much higher in sphere cultured cells than monolayer cultured cells both in osteosarcoma cell lines and primary osteosarcoma cells, with an average fold change of 3.15±0.34 (P<0.01) (Fig. 1A). In accordance with the telomerase activity, expression of hTERT mRNA was also upregulated under sphere culture condition, with an average fold change of 3.47±1.10 (P<0.01) (Fig. 1B).
A subset of osteosarcoma cells can activate an exogenous hTERT Promoter and GFP expression
In order to perform live tracing of the telomerase activity, we constructed a lentiviral hTERTpromoter-GFP reporter vector with a puromycin selection cassette, and established osteosarcoma cell lines stably expressing the hTERT promoter reporter. To assess the validity of the lentivirus, we used both a transcriptionally inactive negative control virus and a transcriptionally active positive control virus (Fig. 1C). The negative control group had no GFP positive cells, which demonstrates the specificity of the cloned hTERT promoter reporter. The positive control vector, without puromycin selection, demonstrated high numbers of GFP positive cells implying that the osteosarcoma cell lines are easily infected by this lentiviral vector. We found that only a subset of osteosarcoma cells can activate the exogenous hTERT promoter and express GFP (Fig. 1C). The GFP positive portion of MG63, MNNG/HOS and 143B was 23.24±3.88, 15.51±2.79 and 22.44±2.74, respectively. We then asked whether hTERT transcription activity correlates with hTERT mRNA and telomerase activity in human osteosarcoma cell lines. We divided the osteosarcoma cells into GFP-pos and GFP-neg groups according to their GFP expression, and found that both the telomerase activity and hTERT mRNA levels were higher in the GFP-pos cells, with the average fold increase of 3.42±0.89 (P<0.01) and 3.29±0.45 (P<0.01), respectively (Fig. 1D-E).
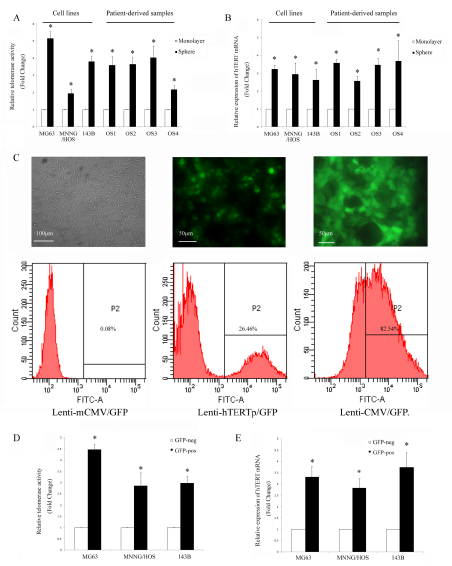
Figure 1: Osteosarcoma cells are heterogeneous according to their telomerase activity. (A) Telomerase activity was detected by TRAP-ELISA, and it was higher in sphere cultured cells. (B) hTERT mRNA level (determined by qPCR) was upregulated in sphere-cultured cells. *P<0.01 compared with the monolayer. (C) Representative images of fluorescence of MG63 cells transduced with different GFP lentiviruses (left: negative control, mCMV-GFP, 100×; middle: hTERT-GFP, 200× and right: positive control, CMV-GFP, all without puromycin selection, 200×). The proportion of GFP positive cells was analyzed using flow cytometry. (D) The osteosarcoma cells were sorted according to their GFP status. Telomerase activity was increased in GFP positive cells and (E) hTERT mRNA level was also higher in GFP positive cells. *P<0.01 compared with the GFP negative cells.
Osteosarcoma cells with high telomerase promoter activity express cancer stem cell markers and related genes
Next, we separated three osteosarcoma cell lines into TELpos and TELneg subpopulations according to their GFP expression status in order to compare their stem cell-like properties. CD117 and Stro-1 are linked to cancer stem cells with metastatic and drug resistant properties in osteosarcoma [14], therefore we investigated if TELpos cells were enriched with cells expressing these markers. The CD117 and Stro-1 double positive rate for MG63, MNNG/HOS and 143B TELpos cells were 5.62±0.75, 6.24±0.09 and 8.27±0.41, respectively. Across the three cell lines tested, we observed an average of 4.51±0.55 fold higher of double positive cells in TELpos cells compared with TELneg cells (P<0.01) (Fig. 2A-B). As a control, the Stro-1 and CD117 status of the non-transduced parental MG63 cells was also assessed (Supplementary Fig.1A). CSCs are reported to possess a transcriptional profile similar to that of embryonic stem cells (ES cells) [19], therefore we investigated if there is differential expression of these genes between TELpos and TELneg cells. We found that Sox2 and Oct4 were greatly upregulated in GFP-positive cells, whereas Nanog was expressed in both GFP-positive and negative cells without significant difference (Fig. 2C) (Detailed densitometry data is shown in Supplementary Table 2).
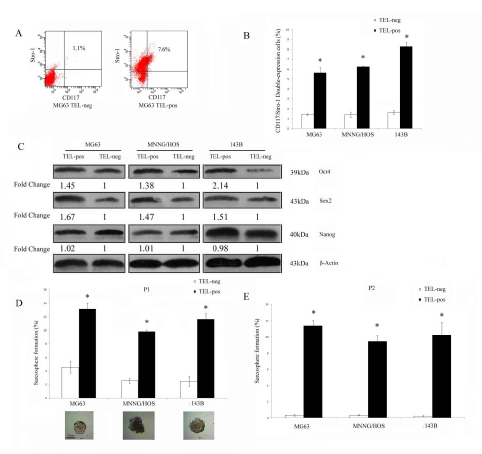
Figure 2: TELpos osteosarcoma cells show increased stem cell-like properties in vitro. (A) A representative FACS analysis of CD117/Stro-1 double positive cells in TELpos and TELneg MG63 cells. (B) TELpos cells had an increased CD117 and Stro-1 double positive cell population. *P<0.01 compared with the TELneg cells. (C) TELpos cells had upregulated expression of Sox2 and Oct4, but not Nanog. Fold changes were calculated from 3 independent experiments normalised using β-actin expression (see also Supplementary Table 2). (D) Sphere culture of osteosarcoma cells. The ability to form sarcospheres was higher in TELpos cells at the first passage (P1). *P<0.01 compared with the TELneg cells. Representative images of sarcosphere are shown, scale bar was 50 μm. (E) The ability of TELneg cells cells to form sarcospheres was almost depleted at the secondary passage (P2). *P<0.01 compared with the TELneg cells.
Osteosarcoma cells possessing high telomerase activity have increased tumorigenic potential both in vitro and in vivo
Sarcosphere culture is an in vitro system used to evaluate the ability of osteosarcoma stem-like cells to form sphere colonies in suspension and whether they self-renew in secondary culture. We compared the ability of TELpos and TELneg cells to form primary and secondary sarcospheres. TELpos cells formed more sarcospheres than TELneg cells, with an average fold increase of 3.8±0.9 (Fig. 2D). Significantly, when dissociated sphere cells were plated for a second generation of sphere culture, self-renewal from TELneg spheres was almost depleted, whereas cells from spheres grown from TELpos cells underwent self-renewal very efficiently (Fig.2E).
The most stringent test of CSC activity is their ability to initiate tumors. We therefore subcutaneously injected serial dilutions of TELpos and TELneg MG63 cells into immunocompromised mice and examined the rate of tumor formation over a period of 6 months. As shown in Table 1, the majority of mice (7/8) injected with 5,000 TELpos cells formed tumors, whereas only one in 8 mice injected with 5×104 TELneg cells showed tumor formation. The extreme limiting dilution assay (ELDA) calculation estimated a 374-fold increase in cancer stem cell frequency in TELpos compared to TELneg cells (Fig. 3A; Table 1). Tumors were further analysed by histological examination, and expression of vimentin indicated their mesenchymal origin (Fig. 3B). Furthermore, we isolated TELpos cells from two different MG63 derived tumors and serially transplanted these into further mice. Tumor formation was observed in 83.3% (5/6) of mice (n = 6) injected with 5,000 cells (Fig. 3C). Serial transplantability of TELpos cells confirmed their in vivo self-renewal activity. We next tested the ability of TELpos cells to initiate osteosarcomas in the bone niche using MNNG/HOS cells. Mice were injected orthotopically into the tibia with TELpos or TELneg cells. 6 out of 8 mice injected with 5,000 TELpos cells formed tumors, whereas no tumours were formed in mice injected with TELneg cells, even when 5×104 cells were injected. ELDA analysis indicated a 232-fold increase in tumour-initiating cell frequencies in TELpos compared to TELneg cells (Fig. 3D; Table 1).
Table 1: Tumor forming ability following subcutaneous and orthotopic injections
Cell line |
MG63 (s.c.) |
P-value |
MNNG/HOS (orthotopic) |
P-value |
||
Cell number |
TELneg |
TELpos |
TELneg |
TELpos |
||
5,000 |
0/8 |
7/8 |
0.001 |
0/8 |
6/8 |
0.003 |
50,000 |
1/8 |
8/8 |
0.001 |
0/8 |
8/8 |
0.000 |
500,000 |
3/8 |
8/8 |
0.013 |
4/8 |
8/8 |
0.038 |
ELDA (95% CI)a |
1/898899 (1/2455639- 1/329046) |
1/2404 (1/5807- 1/996) |
0.000 |
1/835203 (1/2222499- 1/313865) |
1/3607 (1/8572- 1/1518) |
0.000 |
a ELDA: Extreme limiting dilution analysis; CI: confidence interval.
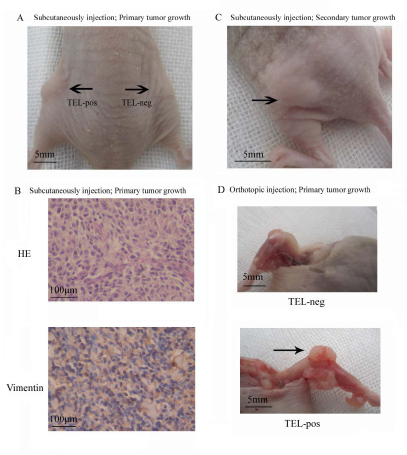
Figure 3: TELpos osteosarcoma cells show increased stem cell-like properties in vivo. (A) MG63 TELpos cells show an increased capacity to form tumors in vivo by subcutaneous injection. The image represents the relative tumorigenic potential of 5×103 TELpos compared with 5×103 TELneg cells. (B) Representative H and E and vimentin staining of MG63 TELpos cells derived tumor (100×). (C) MG63 TELpos cells derived from xenografts form tumor after serial transplantation. (D) MNNG/HOS TELpos cells show an increased capacity to form tumors in vivo by orthotopic injection. The pictures shown the relative tumorigenic potential of 5×103 TELneg compared with 5×103 TELpos cells.
Osteosarcoma cells with high telomerase activity have multipotency
Many cancer stem cell types possess the capability of multipotent differentiation [14, 26]. We demonstrated that cells recovered from TELpos xenograft tumors could be re-sorted into GFP-enriched and non-GFP subpopulations (Fig. 4A). This implies that TELpos cells can differentiate into TELneg cells in vivo, and was confirmed in a TELpos-derived tumor section, where both GFP-positive and GFP-negative cells were observed. To exclude the possibility of contamination from mouse derived-tissue, xenograft tumors were stained with anti-human MHC class I Antibody (Santa Cruz sc-32235), which does not cross-react with mouse tissue. It was found that almost 100% of the cells stained positive, indicating that they were human-derived (Fig. 4B). We also observed that a single TELpos cell could form a sarcosphere colony consisting of TELpos and TELneg cells, suggesting in vitro differentiation of TELpos cells into TELneg cells (Fig.4C).
It is not common to see the differentiation of regular osteosarcoma cells along osteogenic or adipogenic lineage, and therefore this method can be used to test the multipotency of osteosarcoma stem cells. We observed that TELpos cells were able to undergo osteogenic and adipogenic differentiation in vitro, whilst TELneg cells could not be induced along these lineages. These data indicate the multipotency of TELpos cells (Fig. 4D, Supplementary Fig.1B).
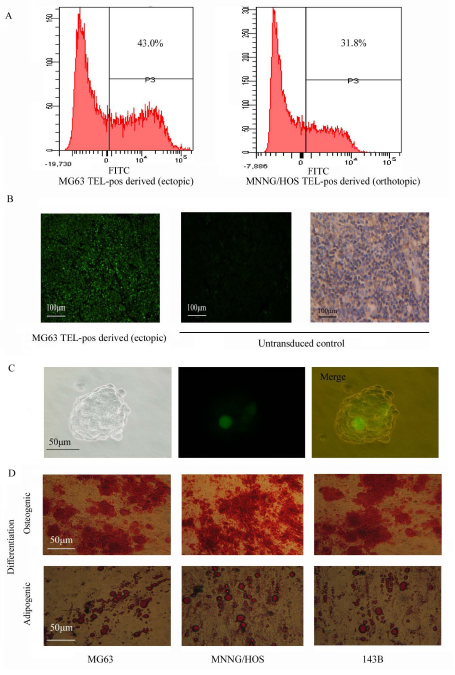
Figure 4: Multipotency of the TELpos cells. (A) Tumor tissues derived from TELpos cells were dissociated into single cells to analyze GFP expression, which showed the production of TELneg cells by TELpos cells. (B) Left: A representative fluorescence image of MG63 TELpos-derived tumor section is shown (100×); middle: non-transduced cells was set as negative control (100×); right: non-transduced cells stained with anti-human MHC ClassⅠantibody (100×). (C) In vitro differentiation of TELpos cells into TELneg, a representative clonally derived sphere of MG63 is shown (400×). (D) In vitro differentiation of TELpos cells, a representative image of TELpos cell differentiation from three osteosarcoma cell lines (200×).
Osteosarcoma cells with high telomerase activity show increased invasion in vitro, metastasis in vivo and drug resistance
We performed a Matrigel Transwell invasion assay to evaluate the invasive properties of different cells in vitro. TELpos cells showed a nearly 3-fold higher invasive potential compared with that of TELneg cells (Fig. 5A). To investigate the metastatic potential of TELpos and TELneg cells, we used the 143B cell line due to its relatively high tendency to metastasise in vivo. We orthotopically injected 1x105 143B cells and the development of pulmonary metastasis was monitored by X-ray detection. Overall, the mice showed a 75% (6/8) appearance rate of lung nodules following TELpos cells injection, whereas none of the TELneg injected mice showed obvious lung nodules (Fig. 5B). We also quantified micrometastatic lesions by serial lung section (Fig.5C), which further confirmed the higher metastatic potential of TELpos cells than TELneg cells .
We next examined whether telomerase active cells could survive current treatments, such as doxorubicin, which is one of the most widely used reagents for osteosarcoma chemotherapy. We found that in all three cell lines TELpos cells showed a higher resistance to doxorubicin than TELneg cells (Fig. 5D).
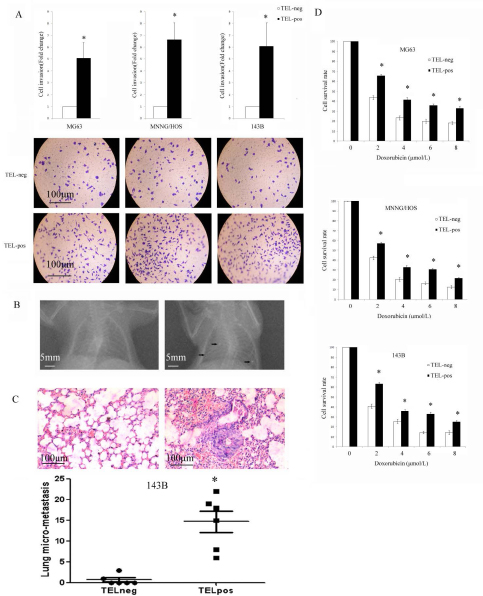
Figure 5: TELpos cells show higher invasion, metastasis potential and drug resistance. (A) The invasive potential of TELpos cells was higher. *P<0.01 compared with the TELneg cells. Representative images of invasion of each cell line are shown (100×). (B) TELpos 143B cells injected in vivo produce obvious detected pulmonary nodules by X-ray examination. (C) The histology examination of 143B cell lung micrometastases. TELpos 143B cells produce a higher number of pulmonary micrometastatic lesions. *P<0.01compared to TELneg cells. (D) Cell sub-populations were treated with increasing concentrations of doxorubicin for 24 h. TELpos cells showed an increased viability in all three cell lines. *P<0.01compared with the TELneg cells.
Treatment with Telomerase inhibitor MST312 preferentially targets GFP positive CSCs
Next, we tested whether the telomerase inhibitor MST312 targets the osteosarcoma cells in vitro. We divided the cells into TELpos and TELneg subgroups and treated with MST312 for 96h. We found a decrease in the cell viability of both subgroups, but a significantly greater inhibitory effect was observed in TELpos cells (Fig. 6A). Furthermore, we found that MST312 inhibited in vitro sphere formation of TELpos cells, with an average inhibition rate of 58.3±5.1% (Fig. 6B). TELpos MG63 cells were then subcutaneously injected into nude mice, and the mice were treated with MST312. After 3 weeks the tumors in control mice were ~ 1cm3, while tumours in the MST312 treated mice were 5-fold smaller (Fig. 6C). We then analysed MG63-TELpos derived tumors treated with MST312 for the GFP positive cell population, and found it to be decreased from 27.3±3.0 to 7.9±2.2 (Fig. 6D).
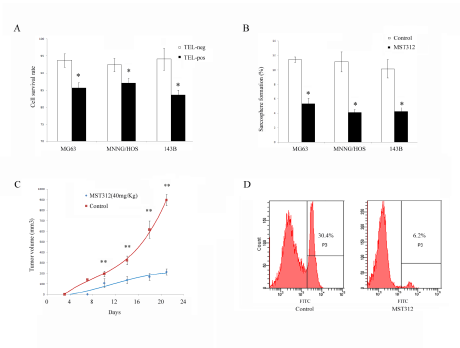
Figure 6: MST312 targets TELpos cells. (A) Cell viability following MST312 treatment of MG63, MNNG/HOS, and 143B cells. Both TELpos and TELneg cells showed a decrease in cell viability but TELpos cells respond significantly more to the treatment (P<0.01). (B) In vitro sphere formation of TELpos cell was inhibited after exposure to MST312 (P<0.01). (C) MST312 inhibited in vivo tumor forming ability of TELpos MG63 cells (P<0.01). (D) TELpos cell population of xenografted tumors were decreased after MST312 treatment.
Discussion
CSCs share many properties with normal stem cells, which are known to express telomerase. Telomerase is also found in 90% of human malignancies and has been reported to predict the outcome of osteosarcoma [27]. We thus proposed to use telomerase activity to isolate osteosarcoma stem cells.
First, we asked whether there was differential activity of telomerase between osteosarcoma stem cells and non-stem cells. Using anchorage-independent, serum-starved culture conditions, a subpopulation of cells capable of self-renewal can be enriched as spherical clones, termed sarcospheres [25]. We found that telomerase activity was markedly enhanced in sarcospheres compared to monolayer cells. This also applied to primary cells from telomerase-expressing osteosarcoma. These data support the theory that telomerase activity can be used to isolate osteosarcoma stem cells. We therefore used a hTERT promoter-GFP lentiviral reporter and established the stable reporter cell lines to assess the cellular telomerase activity. We confirmed that in the three cell lines used the cells can be divided into two subpopulations according to the hTERT promoter activity, which strongly correlated with the expression of the hTERT mRNA and telomerase activity. This enrichment was also observed in four primary telomerase-positive osteosarcomas.
We then tested whether hTERT promoter active cells possess CSC activities such as self-renewal and in vivo transplantation, and if our putative CSCs overlapped with other identified CSC populations in osteosarcoma. The TELpos cells possessed a much stronger capacity to form spheroids than TELneg cells. This ability was not compromised after serial passage, which implies the property of self-renewal. Cell surface markers such as CD117 and Stro-1 have previously been successfully used in isolating osteosarcoma stem cells, and we assessed expression of these in TELpos cells. Phenotypic characterization of TELpos cells showed that they selectively expressed CD117 and Stro-1. Previous studies have reported that ES cell genes play a crucial role in CSCs [29, 30] and TELpos cells were found to overexpress both embryonic stem cell (ES cell)-specific transcription factors Oct4 and Sox2, although Nanog was expressed in both TELpos and TELneg cells without significant difference.
Using a limiting dilution xenograft assay, we confirmed that the sub-population capable of activating the hTERTpromoter/GFP reporter exhibited significantly enhanced tumorigenic activity in vivo. Following either orthotopic or ectopic transplantation, these TELpos cells consistently formed tumors at a lower number of injected cells, and these tumors were able to initiate new tumors after serial transplantation. A further key property of cancer stem cells is the ability to give rise to heterogeneous tumors comprised of tumorigenic and non-tumorigenic cell populations. TELpos cells differentiated toward several mesenchymal phenotypes in vitro, including osteogenic and adipogenic differentiation. Furthermore, TELposcells could produce GFP negative cells both in vitro and in vivo, providing evidence that these cells can give rise to phenotypically diverse progeny.
CSCs have been shown to be more capable of metastasis [31, 32]. Previous research has demonstrated CD117+Stro-1+ osteosarcoma cells to be highly metastatic and enriched for a metastasis-associated marker CXCR4 [14]. In addition, it has been shown that osteosarcoma cells capable of activating a transgenic Oct-4 promoter were more likely to form metastases in the lung [19]. Consistent with these studies, our study established that TELpos cells had higher invasive and metastatic properties compared to TELneg cells, in vitro and in vivo, respectively. TERT has been reported to promote epithelial-mesenchymal transition in gastric cancer cells [33]. It is thus feasible that TELpos osteosarcoma could utilise EMT-related genes to promote metastasis.
Many factors contribute to the drug resistance of CSCs [34, 35]. We found that TELpos cells exhibited a higher resistance to doxorubicin treatment, demonstrating their drug-resistant capability. Doxorubicin’s mechanism of action is via intercalating DNA, causing DDR (DNA damage response) and activating apoptosis [36, 37]. There is increasing evidence that telomerase also participates in DNA repair, and this effect is independent of telomere elongation [38-40]. We speculate that high hTERT expression confers cells with enhanced DNA repair capacity, which results in resistance to doxorubicin.
Different strategies targeting telomerase are currently being tested in several cancers, including osteosarcoma [41, 42]. However, few studies have analyzed whether these therapies are preferentially effective in CSCs. MST312 is a small molecule that has emerged as a promising candidate for anti-telomeric therapy, and it has been reported to be effective in ALDH+ lung cancer stem cells [43, 44]. In our study, we have shown that MST312 has a stronger inhibitory effect on TELpos than TELneg cells. It can inhibit sphere formation in vitro and hinder the in vivo tumor formation of TELpos cells, and it decreases the percentage of TELpos cells in xenografted tumors. These findings provide evidence that a telomerase inhibitor can be preferentially effective in CSCs. It is reported that TERT promotes epithelial proliferation through Myc- and Wnt-related pathways [45], and the Wnt pathway has also been shown to be involved in osteosarcoma pathogenesis [46, 47]. Due to the facts that (i): The MST312 dose used in this study was nontoxic [44]; and (ii): the short duration of treatment, we believe that the effects observed here are mostly due to inhibition of proliferation and self-renewal, and independent of telomere-maintenance.
In contrast to methods using cell surface markers, we found that TELpos cells appear to constitute a relatively large proportion in the tumor mass. This is in accordance with previous research in which osteosarcoma stem cells were isolated using an Oct4/GFP reporter [19]. These data suggest that osteosarcoma contain a large percentage of tumor-initiating cells. Osteosarcoma presents as pleomorphic, malignant, spindle cells that secrete osteoid and express surface antigens commonly associated with mesenchymal stem cells (MSCs), such as CD29, CD44, CD56, CD90 and CD105 [48]. Given these observations, it is likely that osteosarcoma is derived from mesenchymal stem cells. However, it is reported that mesenchymal stem cell have no or very low levels of telomerase activity [50]. In addition, osteosarcoma cell lines such as U2OS and Saos2 [51] and 2/3 of clinical osteosarcoma are telomerase negative [27, 52]. It is thus reasonable that our strategy might not be suitable for those patients where telomerase activity and hTERT expression are undetectable.
Our study is the first to show that osteosarcomas contain a subpopulation of cells identified by high telomerase activity, which possess stem cell-like properties including self-renewal, multi-potency, pulmonary metastatic potential and drug resistance. Further, this study identifies telomerase inhibition as a potential candidate for osteosarcoma stem cell-targeted therapy, which could help to solve the most lethal characteristics of osteosarcoma—recurrence, metastasis and chemotherapy resistance.
Methods
Primary osteosarcoma cells and cell lines
Primary osteosarcoma samples were retrieved from four patients with conventional high-grade osteosarcoma, and were confirmed to be telomerase expressing. Primary osteosarcoma cell lines (OS1-4) were established from tumor lesions taken from surgical biopsies. All the samples were obtained with informed consent, and the study protocol was approved by the institutional review board of Renmin Hospital of Wuhan University. None of specimens had been exposed to chemotherapy or radiotherapy, the diagnosis was confirmed by the histological analysis, and the clinicopathological features are described in Supplementary Table 1. Tumor tissues were processed directly after biopsy. In brief, tumors were washed, cut into small pieces and digested at 37°C with 0.25% trypsin and 0.1% type II collagenase for 30 min and 2 h, respectively. Cells isolated after digestion were plated out into subsequent culture. The telomerase expressing human osteosarcoma cell lines MG63 and MNNG/HOS were purchased from the Shanghai Institute for Biological Sciences of the Chinese Academy of Sciences. 143B was obtained from China Center for Type Culture Collection (CCTCC). All the cells were cultured in RPMI-1640 containing 10% FBS and 1% penicillin/streptomycin. Cells were propagated in a humidified environment at 37˚C with 5% CO2 and 100% humidity. Cell viability was determined using trypan blue staining.
Generation of hTERT-GFP reporter osteosarcoma cell line
The hTERT promoter region (1.5 kb) [53-55]was synthesized and cloned into the multiple cloning site of the lentiviral transcriptional reporter vector pGreenFire1-mCMV-EF1-Puro (TR010PA-P; System Biosciences, Mountain View, CA), with ClaI and EcoRI recognistion site on 5’ and 3’ end, respectively. The promoter region sequence was confirmed by Shanghai Sangon Biotech Co. Ltd. We transfected HEK293-T cells with the constructs and obtained the viral supernatants. To generate stably expressing reporter cell lines, we infected osteosarcoma cells with the viral supernatant and selected the cells with puromycin (5μg/ml) for one week. The transcriptionally inactive pGreenFire-mCMV (TR010PA-P) was used as a negative control and the transcriptionally active pGreenFire-CMV (TR011PA-1) was used as a positive control.
FACScan analysis and sorting
The hTERT-GFP reporter osteosarcoma cells were harvested with fresh 0.25% trypsin solution (Sigma-Aldrich, Inc.) and resuspended in PBS. Cells were maintained on ice until analysis. The GFP expression was assessed and cells were sorted into TELpos and TELneg subpopulation according to their GFP expression status, using a Becton-Dickinson FACSort (San Jose, CA, USA). For analysis of cell surface markers, the cells were harvested and resuspended in PBS/0.5% normal rabbit serum (Sigma-Aldrich, Inc.), and blocked on ice for 15 min. Cells were subsequently labelled with Alexa Fluor® 647 anti-human Stro-1 antibody (BioLegend) and/or PE anti-human CD117 (c-kit) antibody (BioLegend) for 60 min and maintained on ice until analysis. The expression was assessed by flow cytometry and data were analyzed using WinMDI software (Scripps Research Institute, La Jolla, CA, USA).
TRAP-ELISA assay
Telomerase activity was determined using the TeloTAGGG PCR ELISAPLUS kit (Roche, Mannheim, Germany) according to the manufacturer’s protocol. In brief, cells were cultured in monolayer or sphere condition for 14 days and then harvested. 2×105 cells per sample were lysed in 200 μL lysis buffer, and 2 μL cell extract was used for the TRAP reaction. Telomeric repeats were added to biotinylated synthetic primers by telomerase, the elongation product was then amplified by PCR. PCR products were then immobilized to a streptavidin-coated microplate. The immobilized PCR products were detected by an anti-digoxigenin antibody, and then visualized by peroxidase. The absorbance of the samples was measured at 450 nm. Heat-treated cell extract was used as the negative control. A template DNA with the same sequence as a telomerase product with 8 telomeric repeats was used as a positive control. Relative telomerase activities (RTA) were calculated according to protocol.
Drug sensitivity assessment
Osteosarcoma cells (TELpos and TELneg) were cultured for 2 days in complete media to allow cells to recover from sorting stress. Cells (5,000 per well) were cultured in 96-well plates for 1 day and then treated with increasing concentrations of doxorubicin (0-10.0 μmol/L) for 24 hours. Cell viability was measured by CCK8 assay.
Matrigel invasion assay
In vitro invasion assays were performed in transwell chambers (Costar, Cambridge, MA). The upper side of the porous polycarbonate membrane (8.0 µm pore size) was coated with 10 µg/cm2 reconstituted Matrigel basement membrane (Sigma). Cells (2×105/well) were seeded on the upper side of the filter and incubated in RPMI-1640 medium with 0.5% FBS. The lower chamber was filled with complete culture medium containing 10% FBS. After 24h, cells migrated to the lower side were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet. The filters were photographed and cells were counted.
Tumor spheroid assay
The sphere formation assay followed procedures previously described [25]. In brief, cells were plated in six-well ultralow attachment plates (Corning Inc., Corning, NY) at a density of 5,000 cells/well in RPMI-1640 supplemented with B27 Supplement (Invitrogen), 10 ng/mL human EGF (Sigma-Aldrich), and 10 ng/mL human bFGF (Sigma-Aldrich). Cells were incubated at 37˚C in a humidified atmosphere of 95% air and 5% CO2. Fresh aliquots of EGF and bFGF were added every other day. After culture for 14 days, colonies larger than 50 μm in size were regarded as sarcospheres and quantitated by inverted phase contrast microscopy. The spheres were processed to form the next generation of spheres every 14 days.
Animals and transplantation Assay
To determine the in vivo tumorigenicity, BALB/C nude mice about 6 weeks old were purchased and maintained at the Wuhan University Center for Animal Experiment. The care and use of animals has been reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) (approval number: 2011006). After sorting by FACS, live cells were counted by trypan blue staining, and suspended in 10μL of 50% Matrigel/PBS. The mice were randomly divided according to their injected cells and site. Tumors were grown by subcutaneous (MG63) or orthotopic (MNNG/HOS) inoculation with 5 × 103 to 5 × 105 cells. Tumor onset was set at 5 mm diameter, mice were monitored as long as 6 months.
To evaluate the potential to metastasise, we performed orthotopic injections of 143B cells. 1×105 cells in 10μL of 50% Matrigel/PBS were injected into femoral bone marrow cavities of anesthetized BALB/C nude mice. Mice were monitored until 90 days after injection or until tumors reached 10 mm in diameter. At the end of observation point, the mice were anesthetized and X-ray images were taken to detect pulmonary metastases using the KODAK Digital Radiography System. The mice were then sacrificed. Tumor and lung samples were fixed in 4% paraformaldehyde and embedded in paraffin. Tissue sections were stained with haematoxylin and eosin. Data were accumulated from at least three independent experiments.
Differentiation
Osteosarcoma cells were plated in 96-well plates. When cells reached confluency, they were cultured in osteogenic differentiation media for 3 weeks and then stained by alizarin red. For adipogenic differentiation, the cells were cultured in differentiation medium for 5 days. Cells were then fixed with 10% buffered formalin and stained with Oil Red O.
Quantitative real-time PCR
Total RNA was isolated and reverse transcribed. Real-time PCR was then performed using an ABI 7900 System in the presence of SYBR- Green. The following gene-specific primers were used: hTERT (5’ GGAGCAAGTTGCAAAGCATTG -3’, 5’-TCCCACGACGTAGTCCATGTT-3’) and GAPDH (5’ AGAAGGCTGGGGCTCATTTG-3’, 5’-AGGGGCCATCCACAGTCTTC-3’). Target sequences were amplified at 95˚C for 10 min, followed by 40 cycles of 95˚C for 15 s and 60˚C for 1 min. GAPDH was used as endogenous normalization control. All assays were performed in triplicate. The fold change in mRNAs expression was determined according to the method of 2ΔΔCt.
Western Blot
Cell lysates were extracted using RIPA lysis buffer containing protease inhibitor cocktail. Protein concentrations were determined using BCA method. Cell lysates containing 50 μg of protein were loaded and separated on 10% SDS-PAGE gels and subsequently transferred to polyvinylidene difluoride membranes (PVDF). Membranes were blocked in 5% milk solution, incubated at 4˚C overnight with following primary antibodies: rabbit monoclonal Oct4 (Abcam; 1:1,000), rabbit polyclonal Sox2 (Abcam; 1: 1,000), rabbit polyclonal Nanog (Santa Cruz; 1: 2,000), mouse monoclonal β-actin (Santa Cruz; 1: 1,000). They were then washed, and incubated with horseradish peroxidase conjugated secondary antibody at dilution 1:5,000 for 1h at room temperature. Membranes were then washed and developed using ECL Substrate. Densitometry was performed using Image J software.
Testing the efficiency of telomerase inhibitor MST312
Osteosarcoma cell lines were sorted and cultured as described above. The monolayer cultured cells were treated with MST312 for 72 h, with a final concentration of 1µM MST312. For sphere culture, fresh medium with MST312 was used every 3 days, and cultured for 14 days. Xenotransplatation assay was carried out as described above, and 5 × 106 MG63 cells were injected subcutaneously into the flanks of the mice. A total of 8 mice were randomly divided into control (no treatment) and MST312-treated groups, with 4 in each. The MST312 was administered by intraperitoneal injection (40 mg/kg) every other day, starting after injection. Tumor volume was measured every 3-4 days according to the formula: V = length × (width)2/2. All mice were sacrificed 21 days after cell injection and tissues were kept for FACS analysis.
Statistical analyses
Each experiment was performed independently at least three times. Fisher’s Exact Test (one-sided) was used to determine the statistical significance of in vivo tumorigenesis. Otherwise, values were expressed as mean ± SD, and statistical significance (P < 0.05) was determined using Student’s t test.
Acknowledgements
The study was supported in part by Grants from the Hubei Natural Science Foundation of China (2011CHB039), and Fundamental Research Funds for the Central Universities (T201230205). B.M.S. and R.E. were supported by EU FP7 and Breakthrough Breast Cancer funding, respectively.
Disclosure of potential conflict of interest
The authors declare no potential conflicts of interest.
References
1. Ottaviani G, Jaffe N. The epidemiology of osteosarcoma. Cancer Treat Res. 2009;152:3-13.
2. Buckley JD, Pendergrass TW, Buckley CM, Pritchard DJ, Nesbit ME, Provisor AJ, Robison LL. Epidemiology of osteosarcoma and Ewing’s sarcoma in childhood: a study of 305 cases by the Children’s Cancer Group. Cancer. 1998;83:1440-1448.
3. Savage SA, Mirabello L. Using epidemiology and genomics to understand osteosarcoma etiology. Sarcoma. 2011;2011:548151.
4. Yin K, Liao Q, Zhong D, Ding J, Niu B, Long Q, Ding D. Meta-analysis of limb salvage versus amputation for treating high-grade and localized osteosarcoma in patients with pathological fracture. Exp Ther Med. 2012;4:889-894.
5. Allison DC, Carney SC, Ahlmann ER, Hendifar A, Chawla S, Fedenko A, Angeles C, Menendez LR. A meta-analysis of osteosarcoma outcomes in the modern medical era. Sarcoma. 2012;2012:704872.
6. Dokos C, Groll AH. Heterogeneity in osteosarcoma cell lines and the biological effects of the chemical structure of 25-hydroxyvitamin D(3) and 1 a,25 dihydroxyvitamin D(3). J Orthop Res. 2012;30:2052; author reply 2052-2053.
7. Olstad OK, Gautvik VT, Reppe S, Rian E, Jemtland R, Ohlsson C, Bruland OS, Gautvik KM. Molecular heterogeneity in human osteosarcoma demonstrated by enriched mRNAs isolated by directional tag PCR subtraction cloning. Anticancer Res. 2003;23:2201-2216.
8. Valent P, Bonnet D, De Maria R, Lapidot T, Copland M, Melo JV, Chomienne C, Ishikawa F, Schuringa JJ, Stassi G, Huntly B, Herrmann H, Soulier J, Roesch A, Schuurhuis GJ, Wohrer S, et al. Cancer stem cell definitions and terminology: the devil is in the details. Nat Rev Cancer. 2012;12:767-775.
9. Hussein D, Punjaruk W, Storer LC, Shaw L, Othman R, Peet A, Miller S, Bandopadhyay G, Heath R, Kumari R, Bowman KJ, Braker P, Rahman R, Jones GD, Watson S, Lowe J, et al. Pediatric brain tumor cancer stem cells: cell cycle dynamics, DNA repair, and etoposide extrusion. Neuro Oncol. 2011;13:70-83.
10. Nitta E, Suda T. [Cancer stem cell concepts--lesson from leukemia]. Nihon Rinsho. 2009;67:1863-1867.
11. Shenoy AK, Fisher RC, Butterworth EA, Pi L, Chang LJ, Appelman HD, Chang M, Scott EW, Huang EH. Transition from colitis to cancer: high Wnt activity sustains the tumor-initiating potential of colon cancer stem cell precursors. Cancer Res. 2012;72:5091-5100.
12. Wang D, Zhu H, Zhu Y, Liu Y, Shen H, Yin R, Zhang Z, Su Z. CD133(+)/CD44(+)/Oct4(+)/Nestin(+) stem-like cells isolated from Panc-1 cell line may contribute to multi-resistance and metastasis of pancreatic cancer. Acta Histochem. 2012.
13. Visvader JE, Lindeman GJ. Cancer stem cells: current status and evolving complexities. Cell Stem Cell. 2012;10:717-728.
14. Adhikari AS, Agarwal N, Wood BM, Porretta C, Ruiz B, Pochampally RR, Iwakuma T. CD117 and Stro-1 identify osteosarcoma tumor-initiating cells associated with metastasis and drug resistance. Cancer Res. 2010;70:4602-4612.
15. Ying M, Liu G, Shimada H, Ding W, May WA, He Q, Adams GB, Wu L. Human osteosarcoma CD49f(-)CD133(+) cells: impaired in osteogenic fate while gain of tumorigenicity. Oncogene. 2012.
16. Yang M, Yan M, Zhang R, Li J, Luo Z. Side population cells isolated from human osteosarcoma are enriched with tumor-initiating cells. Cancer Sci. 2011;102:1774-1781.
17. Rainusso N, Man TK, Lau CC, Hicks J, Shen JJ, Yu A, Wang LL, Rosen JM. Identification and gene expression profiling of tumor-initiating cells isolated from human osteosarcoma cell lines in an orthotopic mouse model. Cancer Biol Ther. 2011;12:278-287.
18. Wang L, Park P, Zhang H, La Marca F, Lin CY. Prospective identification of tumorigenic osteosarcoma cancer stem cells in OS99-1 cells based on high aldehyde dehydrogenase activity. Int J Cancer. 2011;128:294-303.
19. Levings PP, McGarry SV, Currie TP, Nickerson DM, McClellan S, Ghivizzani SC, Steindler DA, Gibbs CP. Expression of an exogenous human Oct-4 promoter identifies tumor-initiating cells in osteosarcoma. Cancer Res. 2009;69:5648-5655.
20. Blackburn EH, Greider CW, Szostak JW. Telomeres and telomerase: the path from maize, Tetrahymena and yeast to human cancer and aging. Nat Med. 2006;12:1133-1138.
21. Hahn WC. Role of telomeres and telomerase in the pathogenesis of human cancer. J Clin Oncol. 2003;21:2034-2043.
22. Shay JW, Wright WE. Telomeres and telomerase in normal and cancer stem cells. FEBS Lett. 2010;584:3819-3825.
23. Ju Z, Rudolph KL. Telomeres and telomerase in cancer stem cells. Eur J Cancer. 2006;42:1197-1203.
24. Vicente-Duenas C, Romero-Camarero I, Sanchez-Garcia I. Understanding telomerase in cancer stem cell biology. Cell Cycle. 2012;11:1479-1480.
25. Gibbs CP, Kukekov VG, Reith JD, Tchigrinova O, Suslov ON, Scott EW, Ghivizzani SC, Ignatova TN, Steindler DA. Stem-like cells in bone sarcomas: implications for tumorigenesis. Neoplasia. 2005;7:967-976.
26. Singh SK, Hawkins C, Clarke ID, Squire JA, Bayani J, Hide T, Henkelman RM, Cusimano MD, Dirks PB. Identification of human brain tumour initiating cells. Nature. 2004;432:396-401.
27. Sanders RP, Drissi R, Billups CA, Daw NC, Valentine MB, Dome JS. Telomerase expression predicts unfavorable outcome in osteosarcoma. J Clin Oncol. 2004;22:3790-3797.
28. Fujiwara-Akita H, Maesawa C, Honda T, Kobayashi S, Masuda T. Expression of human telomerase reverse transcriptase splice variants is well correlated with low telomerase activity in osteosarcoma cell lines. Int J Oncol. 2005;26:1009-1016.
29. Basu-Roy U, Seo E, Ramanathapuram L, Rapp TB, Perry JA, Orkin SH, Mansukhani A, Basilico C. Sox2 maintains self renewal of tumor-initiating cells in osteosarcomas. Oncogene. 2012;31:2270-2282.
30. Wang L, Park P, Lin CY. Characterization of stem cell attributes in human osteosarcoma cell lines. Cancer Biol Ther. 2009;8:543-552.
31. Baccelli I, Trumpp A. The evolving concept of cancer and metastasis stem cells. J Cell Biol. 2012;198:281-293.
32. Duarte S, Momier D, Baque P, Casanova V, Loubat A, Samson M, Guigonis JM, Staccini P, Saint-Paul MC, De Lima MP, Carle GF, Pierrefite-Carle V. Preventive cancer stem cell-based vaccination reduces liver metastasis development in a rat colon carcinoma syngeneic model. Stem Cells. 2013;31:423-432.
33. Liu Z, Li Q, Li K, Chen L, Li W, Hou M, Liu T, Yang J, Lindvall C, Bjorkholm M, Jia J, Xu D. Telomerase reverse transcriptase promotes epithelial-mesenchymal transition and stem cell-like traits in cancer cells. Oncogene. 2012.
34. Elliott A, Adams J, Al-Hajj M. The ABCs of cancer stem cell drug resistance. IDrugs. 2010;13:632-635.
35. Singh A, Settleman J. EMT, cancer stem cells and drug resistance: an emerging axis of evil in the war on cancer. Oncogene. 2010;29:4741-4751.
36. Davidson D, Grenier J, Martinez-Marignac V, Amrein L, Shawi M, Tokars M, Aloyz R, Panasci L. Effects of the novel DNA dependent protein kinase inhibitor, IC486241, on the DNA damage response to doxorubicin and cisplatin in breast cancer cells. Invest New Drugs. 2012;30:1736-1742.
37. Kawakami K, Nishida H, Tatewaki N, Nakajima Y, Konishi T, Hirayama M. Persimmon leaf extract inhibits the ATM activity during DNA damage response induced by Doxorubicin in A549 lung adenocarcinoma cells. Biosci Biotechnol Biochem. 2011;75:650-655.
38. Ray S, Karamysheva Z, Wang L, Shippen DE, Price CM. Interactions between telomerase and primase physically link the telomere and chromosome replication machinery. Mol Cell Biol. 2002;22:5859-5868.
39. Stellwagen AE, Haimberger ZW, Veatch JR, Gottschling DE. Ku interacts with telomerase RNA to promote telomere addition at native and broken chromosome ends. Genes Dev. 2003;17:2384-2395.
40. Masutomi K, Yu EY, Khurts S, Ben-Porath I, Currier JL, Metz GB, Brooks MW, Kaneko S, Murakami S, DeCaprio JA, Weinberg RA, Stewart SA, Hahn WC. Telomerase maintains telomere structure in normal human cells. Cell. 2003;114:241-253.
41. Fujimori J, Matsuo T, Shimose S, Kubo T, Ishikawa M, Yasunaga Y, Ochi M. Antitumor effects of telomerase inhibitor TMPyP4 in osteosarcoma cell lines. J Orthop Res. 2011;29:1707-1711.
42. Xu T, Rao Y, Zhu W, Guo F. Inhibition of cell growth and telomerase activity in osteosarcoma cells by DN-hTERT. J Huazhong Univ Sci Technolog Med Sci. 2006;26:601-603.
43. Serrano D, Bleau AM, Fernandez-Garcia I, Fernandez-Marcelo T, Iniesta P, Ortiz-de-Solorzano C, Calvo A. Inhibition of telomerase activity preferentially targets aldehyde dehydrogenase-positive cancer stem-like cells in lung cancer. Mol Cancer. 2011;10:96.
44. Seimiya H, Oh-hara T, Suzuki T, Naasani I, Shimazaki T, Tsuchiya K, Tsuruo T. Telomere shortening and growth inhibition of human cancer cells by novel synthetic telomerase inhibitors MST-312, MST-295, and MST-1991. Mol Cancer Ther. 2002;1:657-665.
45. Choi J, Southworth LK, Sarin KY, Venteicher AS, Ma W, Chang W, Cheung P, Jun S, Artandi MK, Shah N, Kim SK, Artandi SE. TERT promotes epithelial proliferation through transcriptional control of a Myc- and Wnt-related developmental program. PLoS Genet. 2008;4:e10.
46. Li C, Shi X, Zhou G, Liu X, Wu S, Zhao J. The canonical Wnt-beta-catenin pathway in development and chemotherapy of osteosarcoma. Front Biosci. 2013;18:1384-1391.
47. Lin CH, Guo Y, Ghaffar S, McQueen P, Pourmorady J, Christ A, Rooney K, Ji T, Eskander R, Zi X, Hoang BH. Dkk-3, a secreted wnt antagonist, suppresses tumorigenic potential and pulmonary metastasis in osteosarcoma. Sarcoma. 2013;2013:147541.
48. Siclari VA, Qin L. Targeting the osteosarcoma cancer stem cell. J Orthop Surg Res. 2010;5:78.
49. Mohseny AB, Szuhai K, Romeo S, Buddingh EP, Briaire-de Bruijn I, de Jong D, van Pel M, Cleton-Jansen AM, Hogendoorn PC. Osteosarcoma originates from mesenchymal stem cells in consequence of aneuploidization and genomic loss of Cdkn2. J Pathol. 2009;219:294-305.
50. Zimmermann S, Voss M, Kaiser S, Kapp U, Waller CF, Martens UM. Lack of telomerase activity in human mesenchymal stem cells. Leukemia. 2003;17:1146-1149.
51. Jegou T, Chung I, Heuvelman G, Wachsmuth M, Gorisch SM, Greulich-Bode KM, Boukamp P, Lichter P, Rippe K. Dynamics of telomeres and promyelocytic leukemia nuclear bodies in a telomerase-negative human cell line. Mol Biol Cell. 2009;20:2070-2082.
52. Terasaki T, Kyo S, Takakura M, Maida Y, Tsuchiya H, Tomita K, Inoue M. Analysis of telomerase activity and telomere length in bone and soft tissue tumors. Oncol Rep. 2004;11:1307-1311.
53. Wick M, Zubov D, Hagen G. Genomic organization and promoter characterization of the gene encoding the human telomerase reverse transcriptase (hTERT). Gene. 1999;232:97-106.
54. Takakura M, Kyo S, Kanaya T, Hirano H, Takeda J, Yutsudo M, Inoue M. Cloning of human telomerase catalytic subunit (hTERT) gene promoter and identification of proximal core promoter sequences essential for transcriptional activation in immortalized and cancer cells. Cancer Res. 1999;59:551-557.
55. Cong YS, Wen J, Bacchetti S. The human telomerase catalytic subunit hTERT: organization of the gene and characterization of the promoter. Hum Mol Genet. 1999;8:137-142.

