INTRODUCTION
Prostate glands are the largest accessory glands in male genital organs, with pivotal physiological functions. Benign prostatic hyperplasia (BPH) is the most common disease causing urination disorder in medium-elderly males, and its morbidity is increasing with age [1-2]. BPH usually occurs in people beyond the age of 40, and its incidence rate can reach 50% and 83% in people aged 60 and 80 years, respectively [3-4]. According to statistics, approximately half of the BPH patients will undergo moderate or severe lower urinary tract symptom (LUTS) [5] which gravely impacts their living quality. At present, a few factors correlated with BPH clinical progression have been identified, including age, serum prostate specific antigen (PSA), prostate volume, maximum flow rate, postvoid residual urine (PVRV) and international prostate symptom score (I-PSS) [6-7]. Along with the advancement of population aging, BPH has been becoming more prevalent and inducing a series of medical and social-economic problems.
Prostate specific antigen (PSA) is a glucoproteinase, and its active form is a serine protease composed by 237 residues [8]. 70% of PSA have chymotrypsin-like activity, and are closely related to male fertility [9]. In addition, PSA can hydrolyze insulin-like growth factor binding proteins for the release of active free insulin-like growth factors (IGF-Is) which are secreted by epithelial cells and can promote cell’s growth through paracrine [10]. Since IGF-I receptors are distributed on the surfaces of both normal and malignant prostatic cells, it has been hypothesized that PSA possesses certain pathological effects on the growth of prostate cancer cells [11]. It has been confirmed that the expression of PSA is regulated by androgen [12], as well as by its coding gene. The gene PSA is located on the long arm of chromosome 19, and contains 5 exons, 4 introns and 3 promoters. Documents have demonstrated that the polymorphism-158G/A in this gene can potentially affect the transcriptional control of androgen over expression, and thus exert certain influences on PSA expression and prostate development [13-14].
Association researches have previously discussed the effect of PSA gene -158G/A polymorphism on BPH risk, but the findings remained contradictory. Consequently, present meta-analysis was carried out backed by all available studies to systematically clarify such relationship.
RESULTS
Study characteristics
As presented in Figure 1, 95 studies were retrieved from databases after the initial search, and 73 of them were excluded after title and abstract review. Among the rest of 22 studies, 15 more were removed due to letter (n = 1), review (n = 1) and without controls (n = 13). Therefore, a total of 7 independent studies [15-21] were ultimately included in this meta-analysis, containing 758 cases and 752 controls. The principal characteristics of included studies are listed in Table 1. Genotype distribution in controls was corresponded to HWE in all but one study [18].
Table 1: Principal characteristics of the studies included in the meta-analysis
First author-Year |
Country |
Ethnicity |
Control source |
Genotyping method |
Case |
Control |
HWE |
||||
GG |
GA |
AA |
GG |
GA |
AA |
||||||
Alptekin-2012 |
Turkey |
Caucasian |
Population-based |
PCR-RFLP |
2 |
6 |
3 |
1 |
9 |
1 |
0.035 |
Ersekerci-2015 |
Turkey |
Caucasian |
Hospital-based |
PCR-RFLP |
8 |
13 |
7 |
7 |
13 |
10 |
0.495 |
Gunes-2007 |
Turkey |
Caucasian |
Hospital-based |
PCR-RFLP |
34 |
62 |
40 |
15 |
44 |
43 |
0.499 |
Sobti-2008 |
India |
Asian |
Hospital-based |
PCR-RFLP |
57 |
99 |
14 |
74 |
84 |
12 |
0.068 |
Soni-2012 |
India |
Asian |
Population-based |
PCR-RFLP |
32 |
70 |
18 |
57 |
36 |
13 |
0.065 |
Wang-2003 |
Japan |
Asian |
Hospital-based |
PCR-RFLP |
125 |
77 |
14 |
163 |
88 |
15 |
0.497 |
Binnie-2005 |
UK |
Caucasian |
Population-based |
PCR-RFLP |
14 |
63 |
11 |
56 |
NA |
||
PCR-RFLP, polymerase chain reaction-restriction fragment length polymorphism; HWE, Hardy-Weinberg equilibrium; NA, not available.
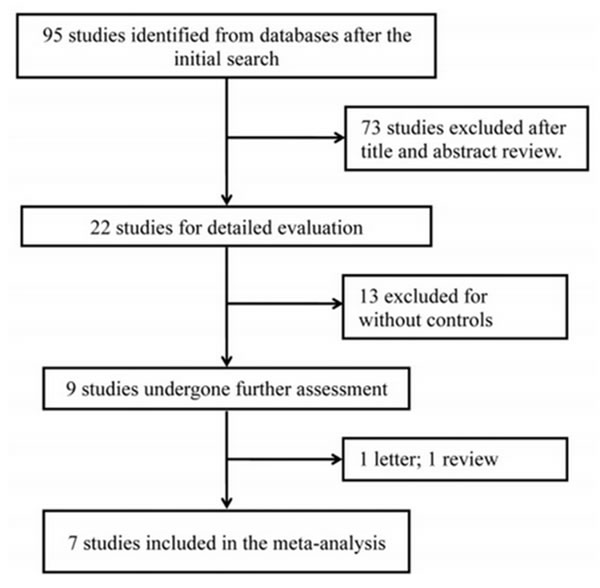
Figure 1: Flow diagram of selecting eligible studies for the meta-analysis.
Meta-analysis results
The meta-analysis results in Table 2 demonstrated that PSA -158G/A polymorphism didn’t affect the susceptibility of BPH significantly under all genetic models in total analysis. However, after subgroup analyses by ethnicity and source of control, PSA -158G/A polymorphism was correlated with decreased susceptibility of BPH in Caucasian group [AA vs. GG: OR = 0.47, 95% CI = 0.25-0.89 (Figure 2); allele A vs. allele G: OR = 0.68, 95% CI = 0.49-0.93], and linked with increased risk in Asian [AA vs. GG: OR = 1.63, 95% CI = 1.02-2.60 (Figure 2); allele A vs. allele G: OR = 1.37, 95% CI = 1.03-1.83] and population-based [AA vs. GG: OR = 2.39, 95% CI = 1.07-5.38; allele A vs. allele G: OR = 1.83, 95% CI = 1.26-2.65 (Figure 3)] groups.
Table 2: PSA -158G/A polymorphism and the susceptibility to benign prostatic hyperplasia.
Genetic comparison |
Group/Subgroup |
OR (95%CI) |
Ph |
|
AA vs.GG |
Ethnicity |
Caucasian |
0.47 (0.25, 0.89) |
0.691 |
Asian |
1.63 (1.02, 2.60) |
0.464 |
||
Source of control |
Population |
2.39 (1.07, 5.38) |
0.775 |
|
Hospital |
0.84 (0.44, 1.63) |
0.089 |
||
Total |
1.07 (0.57, 2.00) |
0.040 |
||
AA+GA vs. GG |
Ethnicity |
Caucasian |
0.64 (0.40, 1.04) |
0.784 |
Asian |
1.72 (0.99, 2.99) |
0.011 |
||
Source of control |
Population |
1.45 (0.46, 4.55) |
0.025 |
|
Hospital |
1.00 (0.64, 1.58) |
0.059 |
||
Total |
1.16 (0.72, 1.86) |
0.002 |
||
AA vs. GG+GA |
Ethnicity |
Caucasian |
0.64 (0.40, 1.03) |
0.336 |
Asian |
1.20 (0.77, 1.88) |
0.987 |
||
Source of control |
Population |
1.41 (0.68, 2.91) |
0.405 |
|
Hospital |
0.80 (0.56, 1.15) |
0.329 |
||
Total |
0.90 (0.65, 1.24) |
0.314 |
||
A vs. G |
Ethnicity |
Caucasian |
0.68 (0.49, 0.93) |
0.549 |
Asian |
1.37 (1.03, 1.83) |
0.102 |
||
Source of control |
Population |
1.83 (1.26, 2.65) |
0.463 |
|
Hospital |
0.94 (0.66, 1.34) |
0.021 |
||
Total |
1.09 (0.77, 1.55) |
0.002 |
||
GA vs. GG |
Ethnicity |
Caucasian |
0.65 (0.35, 1.20) |
0.789 |
Asian |
1.76 (0.97, 3.18) |
0.008 |
||
Source of control |
Population |
1.54 (0.17, 13.70) |
0.086 |
|
Hospital |
1.11 (0.77, 1.58) |
0.211 |
||
Total |
1.28 (0.76, 2.15) |
0.005 |
||
OR, odds ratio; CI, confidence interval; Ph, P-value of heterogeneity test.
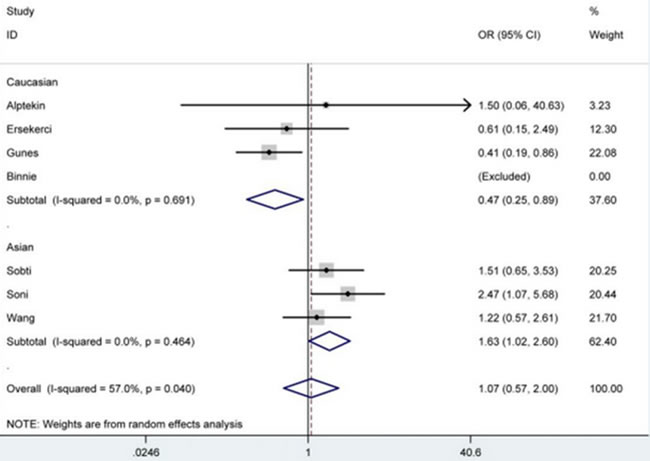
Figure 2: Forest plot of BPH susceptibility associated with PSA -158G/A polymorphism under AA vs. GG model after stratification analysis by ethnicity. The squares and horizontal lines correspond to the study-specific OR and 95% CI. The area of the squares reflects the weight (inverse of the variance). The diamond represents the summary OR and 95% CI.
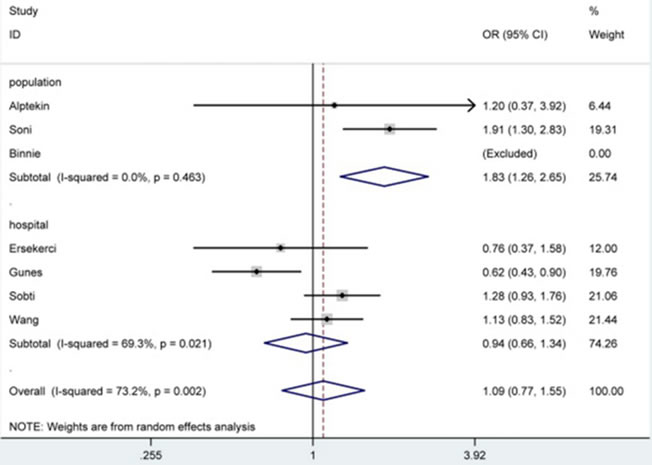
Figure 3: Forest plot of BPH susceptibility associated with PSA -158G/A polymorphism under allele A vs. allele G model after stratification analysis by source of control. The squares and horizontal lines correspond to the study-specific OR and 95% CI. The area of the squares reflects the weight (inverse of the variance). The diamond represents the summary OR and 95% CI.
Heterogeneity analysis
In Q test, significant heterogeneity was detected in AA vs. GG model (P = 0.040), AA+GA vs. GG model (P = 0.002), allele A vs. allele G model (P = 0.002) and GA vs. GG model (P = 0.005). Thus, the random-effects model was selected to evaluate overall results in these four comparisons, while the fixed-effects model was used for the other model (P = 0.314). As for the significant heterogeneity, it was dramatically attenuated or completely removed after stratification analyses by ethnicity and control source. We therefore hypothesized that its plausible origins might include these two aspects.
Sensitivity analysis
We omitted individual studies one at a time and re-calculated overall estimates to detect the impact of each included study on pooled results. Since pooled ORs were not substantially altered, our results were statistically stable and credible.
Evaluation of publication bias
Both Begg’s funnel plot and Egger’s test were utilized to assess the possibility of existing significant publication bias. All funnel plots displayed symmetrical shapes (Figure 4), and Egger’s test also provided evidence for the symmetry (P = 0.150), indicating publication bias was negligible.
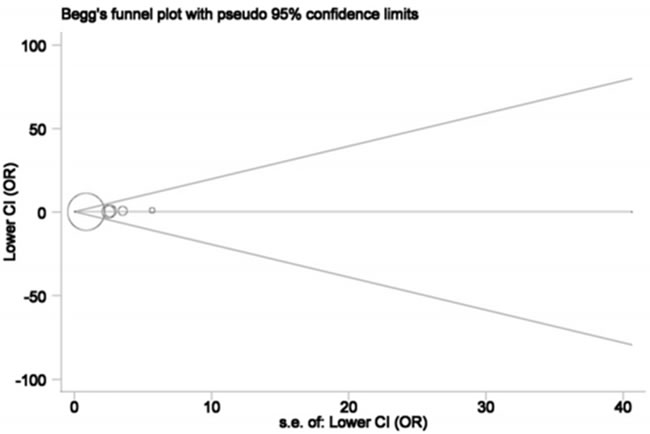
Figure 4: Begg’s funnel plot for publication bias under the model AA vs. GG. Each point represents a separate study for the indicated association. Log(OR), natural logarithm of OR. Horizontal line, mean effect size.
DISSCUSSION
BPH is a chronic progression disease, involving multiple factors and bothering elderly men in general [22-23]. Multiple hypotheses have been put forward for the pathogenesis of BPH, involving effects of hormone and intratesticular non-androgen materials, apoptosis, mesenchyme-epithelium interaction and growth factors. Literature has shown that age and testosterone level could impact the incidence of BPH notably. In addition, different races and genetic backgrounds are also considered as relevant components to BPH development.
PSA, regulated by androgen, is produced by prostate cells, and is generally used as a prostate disease marker to predict the enlargement of prostate volume clinically. PSA gene is polymorphic, and among others, a polymorphism at position -158 (G/A) is one of the most frequently studied objects. In a Turkey population, Gunes et al. found an increasing effect of the GG genotype of PSA -158G/A polymorphism on BPH susceptibility, with an age-adjusted OR of 2.77 (95% CI = 1.277-6.012) [20]. On the contrary, Abha Soni et al. insisted that the genotypes AA and GA of this polymorphism were significantly related to elevated risk of BPH in their study involving 120 BPH cases and 105 controls in Indians [21]. Whereas, a study by Alptekin and colleagues detected no statistically significant difference in the frequencies of genotypes or alleles of PSA -158G/A polymorphism between controls and BPH cases in Turks, suggesting that the polymorphism might not have independent influence on the disease onset [18].
It is necessary to perform a meta-analysis so as to combine those conflicting findings on the relationship between PSA -158G/A polymorphism and BPH susceptibility for a more convincing conclusion in view of distinct discrepancies among those findings. In our study, PSA -158G/A polymorphism showed no significant impact on BPH incidence in total analysis, but we observed with great interest that this polymorphism exerted opposite effects in different subgroups after stratification analysis by ethnicity and source of control. Specifically, under the genetic comparisons of AA vs. GG and A vs. G, PSA -158G/A polymorphism reduced the disease risk in Caucasian group, and increased the risk in Asian and population-based groups. To verify the reliability of the overall estimates from the present meta-analysis, we also implemented a series of tests. In sensitivity analysis, no qualitative alteration in pooled ORs was observed after the omission of any one of included studies, confirming the robustness of final outcomes. Besides, both funnel plot and Egger’s test offered evidences supporting the absence of significant publication bias among selected studies. Hence, our study results had certain powerful in themselves.
Nevertheless, several limitations still needed to be acknowledged. First of all, we only included the articles previously published through online database searching, so possibly relevant reports unpublished or in other sources might be missed. Secondly, only seven eligible studies were ultimately incorporated into the current meta-analysis, with a relatively small sample size, and this situation might affect the precision of our results. Thirdly, not all eligible provided data adjusted for potentially confounding factors, which might introduce some bias into the final results. Fourthly, effects of gene-environment and gene-gene interactions on BPH susceptibility were not considered in this paper owing to inadequate data. Finally, since meta-analysis is a secondary analysis, the overall estimates may be affected by the quality of included studies. Therefore, our findings should be interpreted prudently.
In short, this meta-analysis suggests that PSA -158G/A polymorphism may be a protecting factor against BPH in Caucasian populations, but it may enhance the disease risk in Asians. In view of the shortcomings mentioned above, well-designed studies with larger sample sizes are warranted to verify these findings in future.
MATERIALS AND METHODS
Inclusion and exclusion criteria
Studies were included into this meta-analysis if they met the following criteria: (i) exploring the correlation of PSA gene -158G/A polymorphism with BPH susceptibility; (ii) case-control studies; (iii) sufficient data for computing crude odds ratios (ORs) with their 95% confidence intervals (95% CIs). Papers not satisfying any one of the above standards were excluded from the current study. In addition, removed publications also contained letters and reviews.
Search for eligible studies
Relevant studies were searched from PubMed, EMBASE, Chinese National Knowledge Infrastructure (CNKI) and Web of Science databases. Combinations of the following terms were adopted: “prostate specific antigen” or “PSA” or “KLK3” or “-158G/A”, “polymorphism” or “mutation” or “variant”, and “benign prostatic hyperplasia” or “BPH” or “benign prostate hyperplasia”. References cited in pertinent articles were also screened manually to search for additional relevant papers.
Data extraction
Two reviewers abstracted the following information independently from included studies: first author’s name, year of publication, original country, ethnicity, control source, genotyping method, numbers of cases and controls, genotype frequencies in case and control groups, and P values for Hardy-Weinberg equilibrium (HWE) in controls. All discrepancies over extracted data were resolved via discussion between the two reviewers. If not, the third reviewer would be consulted to reach a final determination.
Statistical analysis
All the statistical tests were conducted using the STATA (version 12.0; StataCorp, College Station, TX) software. The conformity of genotype distribution to HWE in controls was investigated through the goodness of fit chi-square test, and P > 0.05 were regarded as fine compliance to the Law. Pooled ORs with 95% CIs were calculated to estimate the relationship between PSA -158G/A polymorphism and BPH susceptibility under five models: AA vs. GG, AA+GA vs. GG, AA vs. GG+GA, allele A vs. allele G and GA vs. GG. The significance of the pooled ORs was determined by Z-test. Subgroup analyses according to ethnicity and source of control were performed so as to further test such relationship. Heterogeneity among the included studies was checked by chi-square based Q-test, and P > 0.05 in the test suggesting the absence of significant heterogeneity among studies determined that the pooled ORs were computed with fixed-effects model [24]. Otherwise, random-effects model was used (P < 0.05) [25]. Sensitivity analysis was implemented to examine the influence of each single study on pooled results. Begg’s funnel plots and Egger’s test [26] were used to measure potential publication bias across eligible studies.
CONFLICTS OF INTEREST
The authors declare no conflict of interest.
GRANT SUPPORT
This work was supported by National Key Research and Development Plan of China (Grant No. 2016YFC0106300).
REFERENCES
1. Dobrek L, Thor PJ. Benign prostatic hyperplasia - progress in pathophysiology and management. Polski merkuriusz lekarski : organ Polskiego Towarzystwa Lekarskiego. 2015; 39:263-270.
2. Kucerova R, Bienova M, Kral M, Bouchal J, Trtkova KS, Burdova A, Student V, Kolar Z. Androgenetic alopecia and polymorphism of the androgen receptor gene (SNP rs6152) in patients with benign prostate hyperplasia or prostate cancer. Journal of the European Academy of Dermatology and Venereology. 2015; 29:91-96.
3. Karatzas A, Tzortzis V, Giannatou E, Gravas S, Zachos I, Oeconomou A, Melekos M, Tsezou A. Lack of association between the UDP-glucuronosyltransferase 1A1 (UGT1A1) gene polymorphism and the risk of benign prostatic hyperplasia in Caucasian men. Molecular biology reports. 2013; 40:6665-6669.
4. Roehrborn CG, Rosen RC. Medical therapy options for aging men with benign prostatic hyperplasia: focus on alfuzosin 10 mg once daily. Clinical interventions in aging. 2008; 3:511-524.
5. Ray A, Morgan H, Wilkes A, Carter K, Carolan-Rees G. The urolift system for the treatment of lower urinary tract symptoms secondary to benign prostatic hyperplasia: A nice medical technology guidance. Applied health economics and health policy. 2016.
6. Coban S, Doluoglu OG, Keles I, Demirci H, Turkoglu AR, Guzelsoy M, Karalar M, Demirbas M. Age and total and free prostate-specific antigen levels for predicting prostate volume in patients with benign prostatic hyperplasia. The aging male. 2016:1-4.
7. Yamanishi T, Kaga K, Fuse M, Shibata C, Kamai T, Uchiyama T. Six-year follow up of silodosin monotherapy for the treatment of lower urinary tract symptoms suggestive of benign prostatic hyperplasia: What are the factors for continuation or withdrawal? International journal of urology. 2015; 22:1143-1148.
8. Pejcic T, Acimovic M, Dzamic Z, Radovanovic M, Hadzi-Djokic J. Benign prostatic hyperplasia and prostate-specific antigen. Vojnosanitetski pregled. 2015; 72:447-453.
9. Naz RK, Butler TS. Antibodies to prostate-specific antigen in immunoinfertile women and men. Journal of reproductive immunology. 2013; 97:217-222.
10. Wong LI, Labrecque MP, Ibuki N, Cox ME, Elliott JE, Beischlag TV. p,p’-Dichlorodiphenyltrichloroethane (p,p’-DDT) and p,p’-dichlorodiphenyldichloroethylene (p,p’-DDE) repress prostate specific antigen levels in human prostate cancer cell lines. Chemico-biological interactions. 2015; 230:40-49.
11. Zu K, Mucci L, Rosner BA, Clinton SK, Loda M, Stampfer MJ, Giovannucci E. Dietary lycopene, angiogenesis, and prostate cancer: a prospective study in the prostate-specific antigen era. Journal of the National Cancer Institute. 2014; 106:djt430.
12. Kitagawa Y, Ueno S, Izumi K, Kadono Y, Mizokami A, Hinotsu S, Akaza H,Namiki M. Clinical outcomes and nadir prostate-specific antigen (PSA) according to initial PSA levels in primary androgen deprivation therapy for metastatic prostate cancer. World journal of urology. 2016; 34:319-327.
13. Chavan SV, Maitra A, Roy N, Chavan PR. Contribution of allelic variability in prostate specific antigen (PSA) & androgen receptor (AR) genes to serum PSA levels in men with prostate cancer. The Indian journal of medical research. 2014; 139:371-378.
14. Samzadeh M, Hasanzad M, Jamaldini SH, Haghdoost AA, Afshari M, Ziaee SA. Association of G/A polymorphism, rs266882, in AREI region of the prostate-specific antigen gene with prostate cancer risk and clinicopathological features. Urology journal. 2012; 9:691-699.
15. Wang LZ, Sato K, Tsuchiya N, Yu JG, Ohyama C, Satoh S, Habuchi T, Ogawa O, Kato T. Polymorphisms in prostate-specific antigen (PSA) gene, risk of prostate cancer, and serum PSA levels in Japanese population. Cancer letters. 2003; 202:53-59.
16. Sobti RC, Gupta L, Singh SK, Seth A, Kaur P, Thakur H. Role of hormonal genes and risk of prostate cancer: gene-gene interactions in a North Indian population. Cancer genetics and cytogenetics. 2008; 185:78-85.
17. Binnie MC, Alexander FE, Heald C, Habib FK. Polymorphic forms of prostate specific antigen and their interaction with androgen receptor trinucleotide repeats in prostate cancer. The Prostate. 2005; 63:309-315.
18. Alptekin D, Izmirli M, Bayazit Y, Luleyap HU, Yilmaz MB, Soyupak B, Erkoc MA, Tansug Z. Evaluation of the effects of androgen receptor gene trinucleotide repeats and prostate-specific antigen gene polymorphisms on prostate cancer. Genetics and molecular research. 2012; 11:1424-1432.
19. Ersekerci E, Sofikerim M, Taheri S, Demirtas A, Halis F. Genetic polymorphism in sex hormone metabolism and prostate cancer risk. Genetics and molecular research. 2015; 14:7326-7334.
20. Gunes S, Bagci H, Sarikaya S, Bilen CY, Kara N. Prostate-specific antigen and 17-hydroxylase polymorphic genotypes in patients with prostate cancer and benign prostatic hyperplasia. DNA and cell biology. 2007; 26:873-878.
21. Soni A, Bansal A, Mishra AK, Batra J, Singh LC, Chakraborty A, Yadav DS, Mohanty NK, Saxena S. Association of androgen receptor, prostate-specific antigen, and CYP19 gene polymorphisms with prostate carcinoma and benign prostatic hyperplasia in a north Indian population. Genetic testing and molecular biomarkers. 2012; 16:835-840.
22. McConnell JD, Roehrborn CG, Bautista OM, Andriole GL, Jr., Dixon CM, Kusek JW, Lepor H, McVary KT, Nyberg LM, Jr, Clarke HS, Crawford ED, Diokno A, Foley JP, Foster HE, Jacobs SC, Kaplan SA, et al. The long-term effect of doxazosin, finasteride, and combination therapy on the clinical progression of benign prostatic hyperplasia. The New England journal of medicine. 2003; 349:2387-2398.
23. Roehrborn CG. Pathology of benign prostatic hyperplasia. International journal of impotence research. 2008; 20 Suppl 3:S11-18.
24. Mantel N, Haenszel W. Statistical aspects of the analysis of data from retrospective studies of disease. Journal of the National Cancer Institute. 1959; 22:719-748.
25. DerSimonian R, Laird N. Meta-analysis in clinical trials. Controlled clinical trials. 1986; 7:177-188.
26. Egger M, Davey Smith G, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997; 315:629-634.

