INTRODUCTION
HIV-1 remains a global public health problem of unprecedented dimensions. According to the Joint United Nations Programme on HIV/AIDS (UNAIDS), there were 36.7 million (34.0 million-39.8 million) people living with HIV in 2015 [1]. Phylogenetic analysis allows classification of HIV-1 strains into four groups: M, N, O and P. The group M, responsible for the global HIV pandemic, has been further divided into nine subtypes (A-D, F-H, J and K) and 79 circulating recombinant forms (CRFs) to date (http://www.hiv.lanl.gov/content/sequence/HIV/CRFs/CRFs.html). Besides, a proportion of unique recombinant forms (URFs) have also been demonstrated. Actually, new CRFs and URFs continue to be identified and HIV diversity continues to increase [2]. The global distribution of HIV-1 genotypes is extremely complex and dynamic, and specific distributions of genotypes vary among different continents [3]. Globally, the most predominant subtype is subtype C that has spread to different continents, followed by subtype A and B; Subtype B dominates in North America, Western and Central Europe, the Caribbean, Latin America, and Australia [4]. All groups, subtypes and many CRFs have been reported in Africa [5]; The Middle East is mainly affected by subtype B and various CRFs; In India and Ethiopia the epidemics are nearly caused by subtype C. The epidemic in Eastern Europe and Central Asia is dominated by subtype A and B, while in South and Southeast Asia being CRF01_AE. In East Asia the epidemic is dominated by CRF07_BC, CRF08_BC, CRF01_AE and B [4].
Since antiretroviral treatment (ART) has been made available to AIDS patients, the number of patients received the ART has increased rapidly, an estimated 17 million people were accessing life-saving antiretroviral medicines at the end of 2015 according to UNAIDS [1]. The ART has reduced the morbidity and mortality associated with HIV infection, however the great success of ART is now threatened by HIV drug resistance [6]. The new ART recommendations (“treat all”) and the scale-up of pre-exposure prophylaxis [7] using antiretroviral drugs are likely to induce the HIV strains to mutate more quickly under the drug selection pressure. HIV drug resistance has already been observed among treatment-naive patients in China [8–10]. These reports merit attention that the prevalence of antiretroviral drug resistance may compromise the effect of current therapeutic regimens potentially or directly and stress the urgent need to intensify the routine implementation of HIV drug resistance surveys.
Located in the Yangtze River Delta region and northeast of Zhejiang province of China, Jiaxing is a city with very well-developed manufacturing industries attracting large numbers of migrant-workers each year. Since the first case of Jiaxing was detected in 1998, the HIV-1 infection rate has been increasing annually in this city. The HIV epidemiological survey of Jiaxing in 2015 showed that 255 newly diagnosed HIV-1 infected individuals came from 21 provinces as well as municipals, of which the migrants accounted for about 60% of the total infected cases, indicating infections among migrants are a big factor in the HIV-1 epidemic in this city. However, little is known on molecular epidemiology of HIV-1 in Jiaxing, thus our group examined genetic characteristics and antiretroviral resistance mutations among treatment-naive HIV-1 infected patients living in the city.
RESULTS
Subjects included
Of the 99 treatment-naive subjects, 93 (94.0%) pol genes were successfully amplified and sequenced. 81.0% of these subjects were male. Median age was 35 years (range: 16-72). Eighty-three subjects (89.2%) acquired HIV infection through sexual contact. More than half of the participants (56.0%) had the experience of marriage. Nineteen subjects received a high school education degree or above. Median CD4 cells count at sampling was 282 cells/mm3 (range: 12-1266) (Table 1).
Table 1: Demographic characteristics of study subjects a (n=93)
Total n(%)b |
||
|---|---|---|
Gender |
Male |
75(81.0%) |
Female |
18(19.0%) |
|
Household registered |
Local residence in Jiaxing |
40(43.0%) |
others |
53(57.0%) |
|
Age (years) |
<18 |
2(2.2%) |
18-39 |
53(57.0%) |
|
40-59 |
28(30.1%) |
|
≥60 |
10(10.7%) |
|
Marriage Status |
Single |
34(36.6%) |
Married |
28(30.1%) |
|
Divorced/widowed |
24(25.8%) |
|
Unknown |
7(7.5%) |
|
Education |
Middle school and below |
67(72.1%) |
High school and above |
19(20.4%) |
|
Unknown |
7(7.5%) |
|
Occupation |
Workers and peasants |
39(41.9%) |
Commercial sex work |
13(14.0%) |
|
Domestic workers and unemployed |
25(27.0%) |
|
Employees in the service industry |
4(4.3%) |
|
Students |
2(2.1%) |
|
Others and unknown |
10(10.7%) |
|
Route of infection |
Heterosexual transmission |
50(53.7%) |
Homosexual transmission |
33(35.5%) |
|
Intravenous drug use |
4(4.3%) |
|
Unknown |
6(6.5%) |
|
CD4 cell count(cells/mm3) |
≤200 |
27(29.0%) |
>200 |
66(71.0%) |
a, those pol genes were successfully amplified and sequenced; b, values in brackets show the percentages of corresponding cases.
HIV-1 pol phylogenetic diversity
Phylogenetic trees based on pol genes revealed that a majority of the HIV-1 isolates (36, 38.7%) studied belonged to the CRF01_AE, followed by CRF07_BC (32, 34.4%), CRF08_BC (15, 16.1%), subtype B/B’ (5, 5.4%), CRF55_01B (2, 2.1%) (Figure 1, Figure 2). The jpHMM-HIV and bootscanning analyses were performed on three sequences, which did not cluster with any present known references. The results revealed the presence of C/F2/A1 intersubtype recombinant strains with different mosaic structures from those of A1, C and F2 in one sample (CNJX53), the second recombination (CNJX26) was among the CRF and subtype, i.e. CRF01_AE/B, while the third (CNJX55) was among CRFs, i.e. CRF08_BC/CRF07_BC (Figure 3). The sequence analysis revealed a broad diversity in the samples studied.
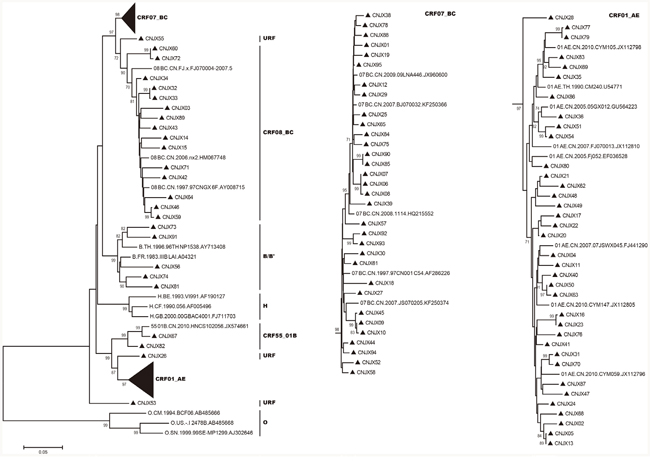
Figure 1: Phylogenetic trees of HIV-1 pol genes were constructed using MEGA 5 based on neighbor-joining methods. The sample sequences and reference HIV-1 genotypes (subtypes B, B’, H, CRF01_AE, CRF07_BC, CRF08_BC, CRF55_01B and group O) were aligned using Bioedit with minor manual adjustments. The statistical robustness of the neighbor-joining tree and reliability of the branching patterns were confirmed by one thousand bootstrap replicates. Only bootstraps greater than 70% are shown at each node. All samples analyzed in the trees are shown by black triangles (▴). URF represents unique recombinant forms. The scale bar represents 5% nucleotide sequence divergence.
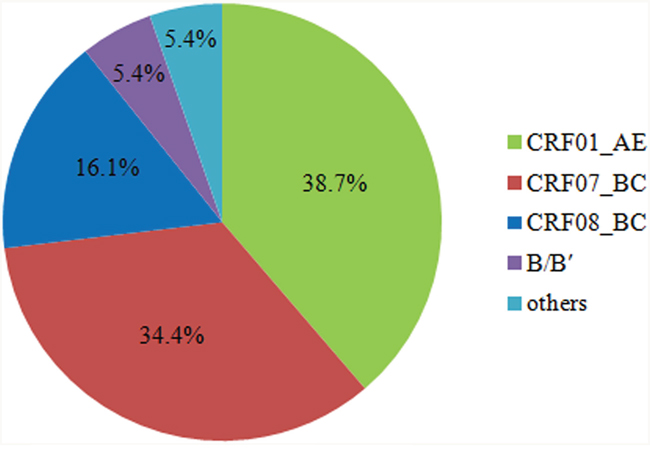
Figure 2: Genotype distribution of HIV-1 strains among treatment-naive individuals in Jiaxing. The partial pol gene fragments of 93 HIV-1 patients were amplified and sequenced. These genotypes were determined by the phylogenetic analysis with reference HIV-1 genotypes. The distribution and percentages of HIV-1 genotypes were shown in pie chart.
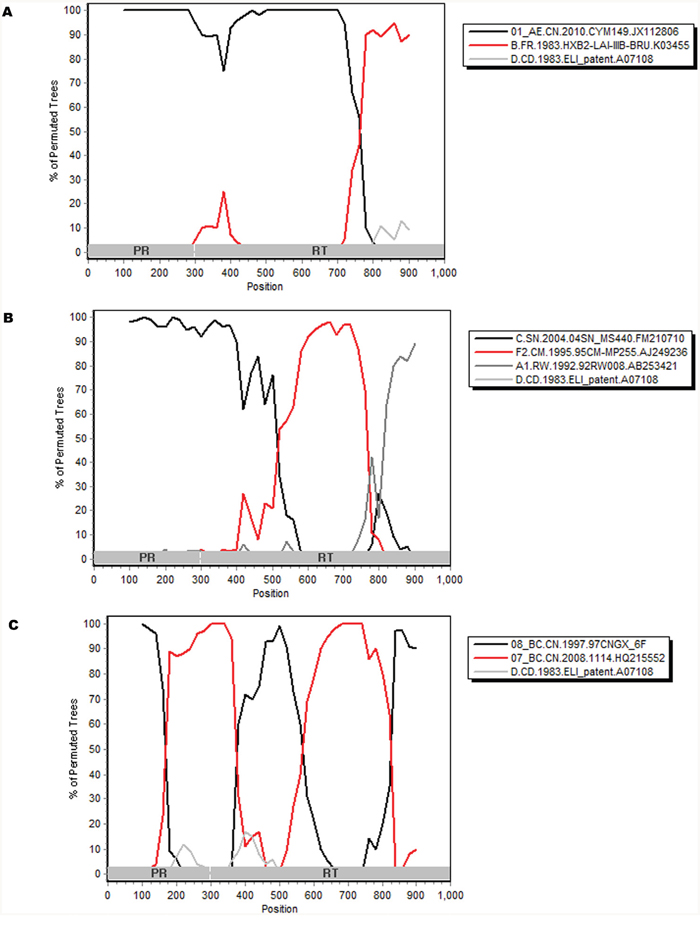
Figure 3. Bootscanning analyses of the partial pol genes of three new unique recombinant strains CNJX26 A., CNJX53 B., CNJX55 C. A bootscanning plot was constructed by using Simplot 3.5.1, which was based on 100 replicates with a 200-bp sliding window moving in steps of 20 bases. The horizontal axis is the nucleotide position from the 5’ end of the analyzed sequence, the vertical axis indicates the percentage supporting the clustering with reference sequences of the analyzed sequence. PR, protease; RT, reverse transcriptase; Subtype references were A1, B, C, D, F2, CRF01_AE, CRF07_BC, CRF08_BC which were shown at the upper right of the figure.
The distribution of HIV-1 strains was uneven among different risk groups. CRF08_BC strains were more likely to be found among heterosexually infected individuals (12, 80%); While most CRF01_AE and CRF07_BC strains were detected in two groups: homosexually infected individuals (20, 55.6%), (10, 31.3%) and heterosexually infected individuals (14, 38.9%), (18, 56.3%) respectively (Table 2).
Table 2: Distribution of HIV-1 genotypes in different risk groups
Risk group |
B/B’ (n=5) |
CRF01_AE (n=36) |
CRF07_BC (n=32) |
CRF08_BC (n=15) |
CRF55_01B (n=2) |
URF b (n=3) |
|---|---|---|---|---|---|---|
Heterosexual contact |
4 (80%) |
14 (38.9%) |
18 (56.3%) |
12 (80%) |
1 (50%) |
1 (33.3%) |
Homosexual contact |
1 (20%) |
20 (55.6%) |
10 (31.3%) |
0 (0%) |
1 (50%) |
1 (33.3%) |
IDU a |
0 (0%) |
0 (0%) |
2 (6.3%) |
1 (6.7%) |
0 (0%) |
1 (33.3%) |
Unknown |
0 (0%) |
2 (5.5%) |
2 (6.3%) |
2 (13.3%) |
0 (0%) |
0 (0%) |
a, Intravenous Drug Use; b, Unique Recombinant Form
Genotypic analysis of HIV-1 drug resistance
Pol genes were successfully amplified from 93 HIV positive samples and were subjected to drug resistance analysis including PI, NRTI and NNRTI resistance-associated mutations through the Stanford HIV Drug Resistance Database. Among our subjects, 46.2% of the amplicons (43/93) contained drug resistance-associated mutations, including 23.7% (22/93) for PI mutations, 1.1% (1/93) for NRTI mutations, and 20.4% (19/93) for NNRTI mutations. PI mutations, such as T74S, L10I, A71T/V, K20I, V11I, were all minor mutations which by themselves showed no significant effect on phenotypes, but might affect the progress of drug resistance or improve replication of viruses containing major mutations [14]. Among samples with reverse transcriptase inhibitors (RTIs) resistance mutations, only one sample harbored NRTI resistance mutation, i.e. T69N, which did not give any resistance to NRTIs; Among samples with NNRTI resistance mutations, the most frequent amino acid substitutions were at the 138th and 179th codon of the reverse transcriptase gene, such as the E138A/K/G (3.2%, 3/93), V179D/E/A/T mutation (9.7%, 9/93), all were rilpivirine (RPV) resistance-associated major or minor mutations [15] (Table 3); Overall, only six individuals (6.5%) showed low to high-level resistance to efavirenz (EFV), etravirine (ETR), rilpivirine (RPV), nevirapine (NVP), all were NNRTIs (Table 4); K103N, conferring a high-level resistance to NVP and EFV, found in three (50%, 3/6) samples, was one of the most common NNRTI-related mutations;
Table 3: HIV-1 antiretroviral resistance mutations identified among different genotypes in treatment-naive subjects
Genotype |
PI (%) a |
PI mutation |
NRTI (%) a |
NRTI mutation |
NNRTI (%) a |
NNRTI mutation |
|---|---|---|---|---|---|---|
CRF01_AE |
19.4%(7/36) |
V11I(1),T74S(1), L10I/V(3), K20I(2) |
2.8%(1/36) |
T69N(1) |
22.2%(8/36) |
V90I(2), V106I(1), V179D(5), K103N(1) |
CRF07_BC |
31.3%(10/32) |
L10I(5), A71V/T(5) |
0%(0/32) |
- |
9.4%(3/32) |
V179D(1), K103N(2) |
CRF08_BC |
6.7%(1/15) |
L10I(1) |
0%(0/15) |
- |
33.3%(5/15) |
V179D(1), E138A/K(3), V90I(1) |
B/B’ |
60%(3/5) |
L10I(1), A71T(3) |
0%(0/5) |
- |
20%(1/5) |
V106I(1) |
CRF55_01B |
0%(0/2) |
- |
0%(0/2) |
- |
100%(2/2) |
V179E(2) |
URF |
33.3%(1/3) |
A71T(1) |
0%(0/3) |
- |
0%(0/3) |
- |
Total |
23.7%(22/93) |
- |
1.1%(1/93) |
- |
20.4%(19/93) |
- |
PI, protease inhibitor; NRTI, nucleoside reverse transcriptase inhibitor; NNRTI, non-nucleoside reverse transcriptase inhibitor; a, the percentage of the samples harboring the PI or NRTI or NNRTI-related mutations in the specific genotype; - : denotes the absence of mutations.
Table 4: Characteristics of patients with primary resistance to antiretroviral drugs
Sample ID |
Sex/age(years)/subtype |
Route of infection/diagnosis year |
Resistance mutation |
Resistance profile |
||||
|---|---|---|---|---|---|---|---|---|
PI |
NRTI |
NNRTI |
Low |
Intermediate |
High |
|||
CNJX14 |
M/42/CRF08_BC |
Hetero/2015 |
- |
- |
E138K,V179AT |
EFV, ETR, NVP |
RPV |
|
CNJX15 |
F/39/CRF08_BC |
Hetero/2015 |
- |
- |
E138A |
RPV |
||
CNJX51 |
F/68/CRF01_AE |
Hetero/2015 |
- |
- |
K103N |
EFV,NVP |
||
CNJX61 |
M/34/CRF07_BC |
IDU/2012 |
- |
- |
K103N |
EFV,NVP |
||
CNJX84 |
M/39/CRF07_BC |
MSM/2015 |
- |
- |
K103N |
EFV,NVP |
||
CNJX89 |
M/56/CRF08_BC |
IDU/2016 |
- |
- |
E138A |
RPV |
||
M, male; F, female; Hetero, Heterosexual contact; MSM, men who have sex with men; IDU, intravenous drug use; EFV, Efavirenz; ETR, Etravirine; RPV, Rilpivirine; NVP, Nevirapine; - : denotes the absence of mutations against the corresponding drugs.
DISCUSSION
Our study firstly detected HIV-1 subtypes and CRFs currently circulating in Jiaxing among treatment-naive individuals. HIV-1 subtype B/B’, CRF01_AE, CRF07_BC, CRF08_BC and CRF55_01B were identified, of which CRF01_AE was the predominant genotype, consistent with the previous study in Zhejiang province [16]. Three URFs, i.e. CRF01_AE/B, C/F2/A1, CRF08_BC/CRF07_BC were also identified, of which two subjects were migrant workers, one was an intravenous drug user from Yunnan province (data not shown). The number of possible URFs detected may be underestimated in this study because only the partial pol gene is sequenced providing limited information on recombination. The identification of CRFs and URFs indicated the high genetic diversity of HIV-1 strains circulating in Jiaxing. Genetic variability of HIV results from the high mutation and recombination rates, together with high replication capacity of the virus [17, 18]. Of note is that recombination requires coinfection or superinfection of viral strains within an individual [19], indicating that coinfection and/or superinfection occur at an alarming speed in Jiaxing. Besides, the migration of HIV-1 infected individuals can also promote viral recombination. Therefore, for Jiaxing, a city with lots of migrant workers flowing into, it is important to identify the genotype to monitor the dynamics and complexity of the HIV epidemic.
Since 1980s, about 40 kinds of antiretroviral drugs have been approved by US Food and Drug Administration, which greatly decreased the morbidity and mortality of HIV infection [20]. However, HIV drug resistance in treatment-naive individuals is increasing worldwide, which may result from the transmission of emergent drug-resistant virus strains to treatment-naive individuals [21]. Our observed drug resistance prevalence (6.5%) in treatment-naive individuals is classified as a moderate level (5%-15%) according to the WHO classification [22], which was similar to the drug resistance prevalence in the recent HIV infections in Zhejiang province, China [8], higher than transmitted drug resistance rates between 2004 (2.9%) and 2005(4.4%) in a nationwide investigation of China when the main transmission route was blood donation [23], lower than those in US and Europe where rates were reported to be 14.6% (2006) [24] and 8.9% (2002-2007) [25], South African with pre-treatment HIV-1 resistance prevalence of 9.0% during 2013 to 2014 [26].
Currently, the most common regimen for HIV patients in Jiaxing contains a combination of a NNRTI plus two NRTIs, such as tenofovir disoproxil fumarate (TDF) / azidothymidine (AZT) + lamivudine (3TC) +EFV/NVP. It was noteworthy that in our study all six patients were resistant to NNRTIs (EFV, ETR, RPV, NVP), while resistance to NRTIs or PIs was not observed. Previous study showed that NNRTIs had a low genetic barrier to drug resistance, a single amino acid change might be sufficient for high-level drug resistance, such as K103N, which had limited effect on viral replication and persisted long after transmission [27, 28]. Besides, NNRTIs usually have longer plasma half-lives than NRTIs [29], making it remain longer in plasma than NRTIs once a patient stops taking drugs, thus induce drug resistance mutations more easily; The above reasons may help explain why all resistant subjects were against NNRTIs. Besides, researches showed that NNRTIs-related mutations increased significantly among newly diagnosed HIV-infected patients in Europe [25, 30], implying us to pay attention to NNRTIs-related mutations. Surveillance of drug resistance mutations among treatment-naive patients will be necessary, thus European and USA guideline panels have already recommended drug-resistance testing prior to treatment [31, 32]. It is hoped that in the future baseline genotypic drug resistance testing can be performed in China to guide doctors select the optimal regimen for a particular patient.
One limitation of our study is that we used standard dideoxynucleotide sequencing, although used as a standard method in HIV drug-resistance testing, fails to identify drug-resistance minority variants that are below 20% of the virus population [33], those minorities can be clinically important as they may grow under drug pressure and lead to treatment failure, thus the method probably underestimates the real prevalence of resistance mutations among treatment-naive patients.
The high genetic diversity of HIV-1 strains and a relative high proportion of antiretroviral resistance mutations among treatment-naive patients pose a serious challenge for HIV prevention and treatment program in Jiaxing; It was noteworthy that most subjects had relatively low levels of education, therefore strengthening propaganda and education of HIV prevention knowledge would be very necessary. This is the first cross-sectional survey describing the genetic characterization of HIV-1 and patterns of PI, NRTI and NNRTI resistance-associated mutations among treatment-naive patients living in Jiaxing, which may provide some implications for future HIV prevention and antiretroviral treatment strategies.
MATERIALS AND METHODS
Study subjects
A total of 104 HIV-1 infected patients living in Jiaxing were enrolled continuously from April 2015 to February 2016. HIV-1 infection in these patients was confirmed by western blot. Of which 5 patients received ART, the remaining 99 subjects were self-reported to be treatment-naive. The epidemiology data were collected by trained interviewers. The study was approved by the Review Board of Jiaxing Municipal Center for Disease Control and Prevention and conducted according to the principles of the World Medical Association Declaration of Helsinki. All subjects provided written informed consent prior to participating in this study. The participants were simultaneously informed of their right to repeal consent by them or their kin, guardians, or caretakers.
HIV-1 pol gene amplification and phylogenetic analysis
Subjects’ whole blood samples were collected and CD4 cell count was measured within 24 hours after sampling. Meanwhile, plasma was separated by centrifugation and stored at -80°C until use. HIV-1 polymerase (pol) gene (containing the full length protease (PR) gene and the first 300 codons of the reverse transcriptase (RT) gene) was amplified, purified and sequenced as previously described [11]. Each sequence was submitted to the Los Alamos National Laboratory HIV Sequence Database (http://www.hiv.lanl.gov/content/index) and COMET HIV-1 [12] to determine HIV genotype, further confirmed by phylogenetic analysis using standard reference sequences. To identify the possible recombination, the sequences were submitted to Jumping Profile Hidden Markov Model (jpHMM-HIV) software [13], and confirmed by bootscanning analyses using Simplot 3.5.1.
Genotypic baseline of HIV-1 drug resistance
For analysis of HIV-1 drug resistance mutation, each sample sequence was submitted to the Stanford HIV Drug Resistance Database (http://hivdb.stanford.edu) to detect drug-resistance mutations including protease inhibitor (PI) major and minor resistance mutations, nucleoside reverse transcriptase inhibitor (NRTI) and non-nucleoside reverse transcriptase inhibitor (NNRTI) resistance mutations. Genotypic susceptibility was interpreted by HIVdb program.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
REFERENCES
1. The Joint United Nations Programme on HIV and AIDS: http://www.unaids.org/en/resources/fact-sheet. Last accessed on 10/30/2016.
2. Hemelaar J. The origin and diversity of the HIV-1 pandemic. Trends Mol Med. 2012; 18: 182-92.
3. Thomson MM, Perez-Alvarez L, Najera R. Molecular epidemiology of HIV-1 genetic forms and its significance for vaccine development and therapy. Lancet Infect Dis. 2002; 2: 461-71.
4. Hemelaar J, Gouws E, Ghys PD, Osmanov S. Global trends in molecular epidemiology of HIV-1 during 2000-2007. AIDS. 2011; 25: 679-89.
5. Peeters M, Toure-Kane C, Nkengasong JN. Genetic diversity of HIV in Africa: impact on diagnosis, treatment, vaccine development and trials. AIDS. 2003; 17: 2547-60.
6. World Health Organization: Global report on early warning indicators of hiv drug resistance. http://www.who.int/hiv/pub/drugresistance/ewi-hivdr-2016/en/. Last accessed on 11/08/2016.
7. World Health Organzation: Guideline on when to start antiretroviral therapy and on pre-exposure prophylaxis for HIV. http://www.who.int/hiv/pub/guidelines/earlyrelease-arv/en/. Last accessed on 09/25/2016.
8. Zhang J, Guo Z, Yang J, Pan X, Jiang J, Ding X, Zhang W, Xia Y, Xu Y, Huang J. Genetic diversity of HIV-1 and transmitted drug resistance among newly diagnosed individuals with HIV infection in Hangzhou, China. J Med Virol. 2015; 87: 1668-76.
9. Jiao Y, Li S, Li Z, Zhang Z, Zhao J, Li L, Wang L, Yin Q, Wang Y, Zeng Z, Shao Y, Ma L. HIV-1 transmitted drug resistance-associated mutations and mutation co-variation in HIV-1 treatment-naive MSM from 2011 to 2013 in Beijing, China. BMC Infect Dis. 2014; 14: 689.
10. Si XF, Huang HL, Wei M, Guan Q, Song YH, Ma PF, Quan Y, Xing H, Shao YM. Prevalence of drug resistance mutations among antiretroviral drug-naive HIV-1-infected patients in China. Chin J Exp Clin Virol. 2004; 18: 308-11.
11. Zhang X, Li S, Li X, Li X, Xu J, Li D, Ruan Y, Xing H, Zhang X, Shao Y. Characterization of HIV-1 subtypes and viral antiretroviral drug resistance in men who have sex with men in Beijing, China. AIDS. 2007; 21: S59-65.
12. Struck D, Lawyer G, Ternes AM, Schmit JC, Bercoff DP. COMET: adaptive context-based modeling for ultrafast HIV-1 subtype identification. Nucleic Acids Res. 2014; 42: e144.
13. Zhang M, Schultz AK, Calef C, Kuiken C, Leitner T, Korber B, Morgenstern B, Stanke M. jpHMM at GOBICS: a web server to detect genomic recombinations in HIV-1. Nucleic Acids Res. 2006; 34: W463-5.
14. D'Aquila RT, International AS-USA, Schapiro JM, Brun-Vezinet F, Clotet B, Conway B, Demeter LM, Grant RM, Johnson VA, Kuritzkes DR, Loveday C, Shafer RW, Richman DD. Drug Resistance Mutations in HIV-1. Top HIV Med. 2002; 10: 21-5.
15. Theys K, Van Laethem K, Gomes P, Baele G, Pineda-Pena AC, Vandamme AM, Camacho RJ, Abecasis AB. Sub-Epidemics Explain Localized High Prevalence of Reduced Susceptibility to Rilpivirine in Treatment-Naive HIV-1-Infected Patients: Subtype and Geographic Compartmentalization of Baseline Resistance Mutations. AIDS Res Hum Retroviruses. 2016; 32: 427-33.
16. Zhang JF, Pan XH, Ding XB, Chen L, Guo ZH, Xu Y, Huang JJ. Molecular epidemiological study on HIV/AIDS under the follow-up program in Zhejiang province in 2009. Chin J Epidemiol. 2013; 34: 44-8.
17. Roberts JD, Bebenek K, Kunkel TA. The accuracy of reverse transcriptase from HIV-1. Science. 1988; 242: 1171-3.
18. Ho DD, Neumann AU, Perelson AS, Chen W, Leonard JM, Markowitz M. Rapid turnover of plasma virions and CD4 lymphocytes in HIV-1 infection. Nature. 1995; 373: 123-6.
19. Robertson DL, Sharp PM, McCutchan FE, Hahn BH. Recombination in HIV-1. Nature. 1995; 374: 124-6.
20. U S Food and Drug Administration: Antiretroviral drugs used in the treatment of HIV infection. http://www.fda.gov/forpatients/illness/hivaids/ucm118915.htm. Last accessed on 11/08/2016.
21. Magiorkinis E, Detsika M, Hatzakis A, Paraskevis D. Monitoring HIV drug resistance in treatment-naive individuals: molecular indicators, epidemiology and clinical implications. Hiv Therapy. 2016; 3: 369-90.
22. Bennett DE, Myatt M, Bertagnolio S, Sutherland D, Gilks CF. Recommendations for surveillance of transmitted HIV drug resistance in countries scaling up antiretroviral treatment. Antivir Ther. 2008; 13: 25-36.
23. Liao L, Xing H, Shang H, Li J, Zhong P, Kang L, Cheng H, Si X, Jiang S, Li X, Shao Y. The prevalence of transmitted antiretroviral drug resistance in treatment-naive HIV-infected individuals in China. J Acquir Immune Defic Syndr. 2010; 53: S10-4.
24. Wheeler WH, Ziebell RA, Zabina H, Pieniazek D, Prejean J, Bodnar UR, Mahle KC, Heneine W, Johnson JA, Hall HI, Variant A, Resistant HIVSG. Prevalence of transmitted drug resistance associated mutations and HIV-1 subtypes in new HIV-1 diagnoses, U.S.-2006. AIDS. 2010; 24: 1203-12.
25. Frentz D, Van de Vijver DA, Abecasis AB, Albert J, Hamouda O, Jorgensen LB, Kucherer C, Struck D, Schmit JC, Vercauteren J, Asjo B, Balotta C, Beshkov D, et al. Increase in transmitted resistance to non-nucleoside reverse transcriptase inhibitors among newly diagnosed HIV-1 infections in Europe. BMC Infect Dis. 2014; 14: 407.
26. Steegen K, Carmona S, Bronze M, Papathanasopoulos MA, van Zyl G, Goedhals D, MacLeod W, Sanne I, Stevens WS. Moderate Levels of Pre-Treatment HIV-1 Antiretroviral Drug Resistance Detected in the First South African National Survey. PLoS One. 2016; 11: e0166305.
27. van de Vijver DA, Wensing AM, Angarano G, Asjo B, Balotta C, Boeri E, Camacho R, Chaix ML, Costagliola D, De Luca A, Derdelinckx I, Grossman Z, Hamouda O, et al. The calculated genetic barrier for antiretroviral drug resistance substitutions is largely similar for different HIV-1 subtypes. J Acquir Immune Defic Syndr. 2006; 41: 352-60.
28. Kikaire B, Khoo S, Walker AS, Ssali F, Munderi P, Namale L, Reid A, Gibb DM, Mugyenyi P, Grosskurth H, Team DT. Nevirapine clearance from plasma in African adults stopping therapy: a pharmacokinetic substudy. AIDS. 2007; 21: 733-7.
29. Bazzoli C, Jullien V, Le Tiec C, Rey E, Mentre F, Taburet AM. Intracellular Pharmacokinetics of Antiretroviral Drugs in HIV-Infected Patients, and their Correlation with Drug Action. Clin Pharmacokinet. 2010; 49: 17-45.
30. Bracciale L, Colafigli M, Zazzi M, Corsi P, Meraviglia P, Micheli V, Maserati R, Gianotti N, Penco G, Setti M, Di Giambenedetto S, Butini L, Vivarelli A, et al. Prevalence of transmitted HIV-1 drug resistance in HIV-1-infected patients in Italy: evolution over 12 years and predictors. J Antimicrob Chemother. 2009; 64: 607-15.
31. AIDSinfo: Guidelines for the Use of Antiretroviral Agents in HIV-1-Infected Adults and Adolescents Available at http://aidsinfo.nih.gov/guidelines. Last accessed on 10/13/2016.
32. Rockstroh JK, Bhagani S, Benhamou Y, Bruno R, Mauss S, Peters L, Puoti M, Soriano V, Tural C, Committee EE. European AIDS Clinical Society (EACS) guidelines for the clinical management and treatment of chronic hepatitis B and C coinfection in HIV-infected adults. HIV Med. 2008; 9: 82-8.
33. Schuurman R, Brambilla D, de Groot T, Huang D, Land S, Bremer J, Benders I, Boucher CA, Group EW. Underestimation of HIV type 1 drug resistance mutations: results from the ENVA-2 genotyping proficiency program. AIDS Res Hum Retroviruses. 2002; 18: 243-8.

