INTRODUCTION
Colorectal cancer (CRC) is the third most common cancer and the second leading cause of cancer-related death all around the world with an estimated 1.2 million new cases and a half million deaths each year [1]. The 5-year survival for patients with early-stage CRC is nearly 90%, so this disease could be potentially cured if early diagnosed [2]. Colonoscopy is widely used in clinical practice, which is regarded as the gold standard for detecting CRC. However, it has several limitations such as invasive nature, high cost and a bothering bowel preparation. Additionally, its success depends on the skill and experience of the operators. Thus, its widespread application for CRC in large-scale screening is hampered [3]. On the other hand, less-invasive diagnostic methods, such as fecal occult blood testing (FOBT) [4] and carcinoembryonic antigen (CEA) screening [5] in blood are of limited value owning to poor sensitivity and specificity [6, 7]. In view of this clinical challenge, novel, reliable and non-invasive biomarkers for early diagnosis of CRC are pressing needed.
MicroRNAs (miRNA), a class of small non-coding RNAs of 19-22 nucleotides in length, function as post-transcriptional regulators by directly cleaving target messenger RNA (mRNA) or translational repression [8]. Numerous studies have demonstrated that miRNAs can be stably detected in serum or plasma and have the potential to be new biomarkers for early diagnosis of various types of cancers [9–12]. Additionally, several studies have revealed that some miRNAs are able to discriminate CRC patients from healthy controls and serve as serum biomarkers for CRC screening with high accuracy [13, 14]. However, these results lack consistencies because of different research methods and tested populations between laboratories.
In the present study, we performed a four-phase study to screen miRNAs in CRC serum samples. In the screening phase, we systematically analyze the miRNA expression profile in 3 peripheral serum pools from 30 CRC cases and 1 pooled sample from 10 controls using miRCURY platform. Differentially expressed miRNAs were subjected to individual qRT–PCR confirmation in 30 CRC patients and 30 healthy controls in the training stage. Then, in the validation phase, the identified miRNAs were further detected in additional samples, including 136 CRC patients and 90 healthy subjects. Meanwhile, the diagnostic potential and effectiveness of these miRNAs for the detection of CRC was evaluated in an independent cohort of 30 CRC serum samples and 10 NCs. Furthermore, the expression profile of identified miRNAs was assessed in the CRC tissue. Serum exosomal miRNAs were further analyzed to explore the potential form of the identified miRNAs in serum of CRC patients.
RESULTS
Characteristics of study subjects
A total of 334 serum samples from 196 patients with CRC and 138 healthy subjects were analyzed in a four-step approach that included: a screening phase, a training phase, a testing phase, and an external validation phase. The demographics and clinical features of the study subjects were shown in Table 1. Schematic representation of the experiment was illustrated in Figure 1. There were no significant differences between CRC patients and normal controls in the distribution of age or gender (p > 0.05).
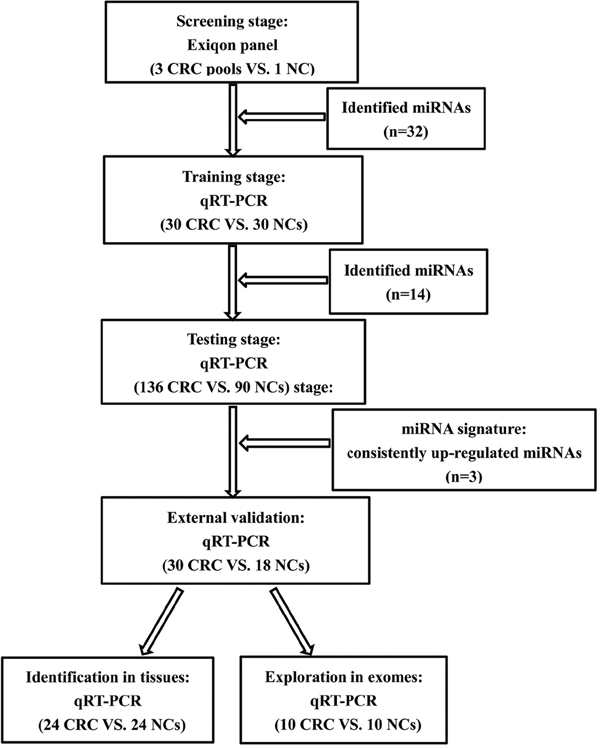
Figure 1: The flow chart of the experiment design. CRC: colorectal cancer; NC: normal control.
Table 1: Clinical characteristics of 196 CRC patients and 138 normal controls
Variables |
Training stage (n = 60) |
Testing stage (n = 226) |
External validation stage |
|||
|---|---|---|---|---|---|---|
Cases (%) |
Controls |
Cases (%) |
Controls |
Cases (%) |
Controls |
|
Number |
30 |
30 |
136 |
90 |
30 |
18 |
Gender |
||||||
Male |
18(60) |
18(60) |
82(60.3) |
46(51.1) |
23(76.7) |
10(55.6) |
Female |
12(40) |
12(40) |
54(39.7) |
44(48.9) |
7(23.3) |
8(44.4) |
Age |
||||||
<60 |
9(30) |
22(73.3) |
55(40.4) |
48(53.3) |
19(63.3) |
11(61.1) |
≥60 |
21(70) |
8(26.7) |
81(59.6) |
42(46.7) |
11(36.7) |
7(38.9) |
Location |
||||||
Colon |
10(33.3) |
61(44.9) |
13(43.3) |
|||
Rectum |
20(66.7) |
75(55.1) |
17(56.7) |
|||
Differentiation grade |
||||||
Middle-Low |
28(93.3) |
127(93.4) |
29(96.7) |
|||
High |
2(6.7) |
9(6.6) |
1(3.3) |
|||
TNM stage |
||||||
I |
9(30) |
26(19.1) |
4(13.3) |
|||
II |
11(36.7) |
60(44.1) |
18(60) |
|||
III |
10(33.3) |
50(36.8) |
8(26.7) |
|||
IV |
0(0) |
0(0) |
0(0) |
|||
Profiling of serum miRNAs for CRC detection
We utilized the Exiqon miRCURY-Ready-to-Use-PCR-Human-panel-I + II-V1.M to screen the expression levels of 168 miRNAs in 3 serum pools from 30 CRC cases and 1 pooled sample from 10 controls. miRNAs that had a Ct < 37 and 5 Ct less than the negative control (No Template Control, NTC) and showed more than 1.5 fold altered expression between all 3 pooled CRC samples and the NC pool sample were selected as candidates. Among the 168 serum miRNAs detected, 32 miRNAs (29 up-regulated miRNAs and 3 down-regulated miRNAs; Supplementary Table 1) were found to be differentially expressed in CRC, which were further analysed by qRT-PCR in the subsequent validation step.
Validation of miRNAs in serum by qRT-PCR
We next performed qRT-PCR assay to confirm the expression of 32 candidate miRNAs in 30 CRC patients and 30 NCs in the training stage. Fourteen miRNAs were found differentially expressed and selected for validation in a larger population consisting of 136 serum samples of CRC patients and 90 NCs in the testing phase (Supplementary Table 1). Consistent with the results from the training phase, 3 out of the 14 miRNAs (miR-19a-3p, miR-21-5p and miR-425-5p) showed significantly higher serum levels in CRC group than in control group. The differential expression of the three miRNAs in the 166 CRC samples compared to the 120 controls enrolled in the training and testing sets was shown in Figure 2.
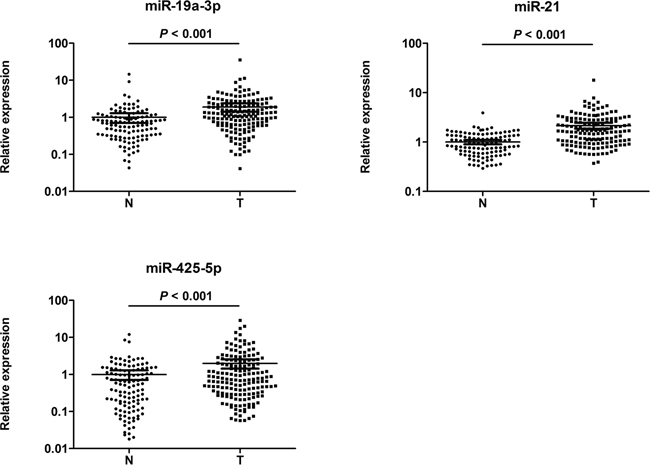
Figure 2: Expression levels of the three miRNAs in the serum of 166 CRC patients and 120 NCs (in the training and validation phases). A: miR-19a-3p; B: miR-21-5p; C: miR-425-5p; N: normal controls; T: tumor. Horizontal line: mean with 95% CI. The y axis represents relative expression of miRNAs normalized to cel-miR-39. P-values were calculated using the nonparametric Mann– Whitney U-test.
Diagnostic value of the candidate miRNAs
We generated ROC curves to evaluate the performance of the three miRNAs in discriminating the CRC patients from NCs. Logistic regression model for CRC prediction was applied on the data from the combination of training and validation phases. The AUC for serum miR-19a-3p, miR-21-5p and miR-425-5p was 0.685, 0.773 and 0.614, respectively when the training and validation samples were combined (Supplementary Figure 1). The combination of the three miRNAs presented further improvement in AUC of 0.783 (95% CI, 0.730 to 0.837) (Figure 3A) for patients with CRC. We also explored the diagnostic value of the three-miRNA panel in the two stages separately. For the training phase, the AUC of the three-miRNA panel was 0.886 (95% CI, 0.803 to 0.968) (Figure 3B). For the validation phase, the miRNA panel showed a AUC value of 0.768 (95% CI, 0.706 to 0.831) (Figure 3C).
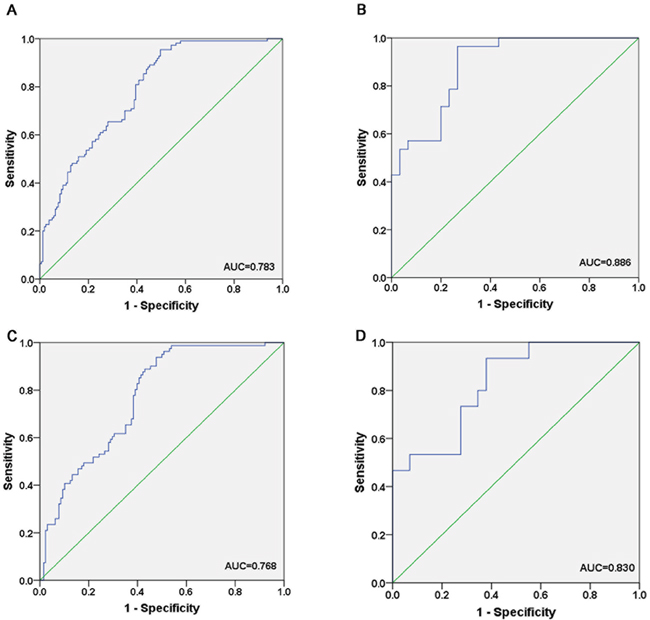
Figure 3: Receiver-operating characteristic (ROC) curves for the three-miRNA panel to discriminate CRC patients from NCs. A. the combined two phases of training and validation phases (166 CRC VS. 120 NCs); B. training phase (30 CRC VS. 30 NCs); C. validation phase (136 CRC VS. 90 NCs); D. external validation (30 CRC VS. 18 NCs). AUC: area under the curve.
To further validate the diagnostic performance of the three-miRNA panel, we measured the expression of miR-19a-3p, miR-21-5p and miR-425-5p by using qRT-PCR in an independent cohort of 30 CRC serum samples and 18 NCs. The results showed that all the three miRNAs were consistently overexpressed in the CRC group compared to NCs. Furthermore, the combination of these three miRNAs yielded an AUC of 0.830 (95% CI, 0.708 to 0.952) (Figure 3D), supporting the diagnostic value of this miRNA panel. We further evaluated the expression levels of miR-19a-3p, miR-21-5p and miR-425-5p with different TNM stages in the total training and validation phases. Unfortunately, none of the three miRNAs was significantly associated with clinical TNM stage (data not shown).
In addition, to evaluate whether these miRNAs were consistently differentially expressed in colon and rectal cancers, we conducted a subgroup analysis. As a result, miR-19a-3p, miR-21-5p and miR-425-5p were all up-regulated both in colon and rectal cancers. Moreover, miR-122-5p was found upregulated in colon cancers while miR-92a-3p upregulated in rectal cancers. The expression levels and diagnostic value of miR-122-5p, miR-19a-3p, miR-21-5p and miR-425-5p in colon cancers were shown in Supplementary Figure 2. The elevated expression and diagnostic performance of miR-19a-3p, miR-21-5p, miR-425-5p and miR-92a-3p in rectal cancers were shown in Supplementary Figure 3. miR-122-5p and miR-92a-3p might have the specificity in colon and rectal cancers, respectively.
Expression of serum miRNAs in tissue samples
The three miRNAs identified in serum were further detectable using qRT–PCR in an additional of 24 pairs of FFPE tissue samples. Consistent with our serum results, the expression of miR-19a-3p, miR-21-5p, and miR-425-5p was found to be significantly higher in colon and rectal cancer tissues, respectively compared to those in normal tissues (Figure 4).
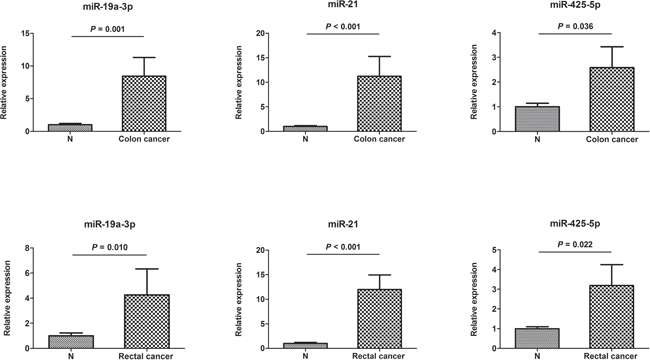
Figure 4: Expression of the three miRNAs in the tumor tissues of 12 colon and 12 rectal cancer patients. N: adjacent nontumor tissues. Error bar: standard error. The y axis represents relative expression of miRNAs normalized to U6.
We also detected the expression levels of miR-122-5p and miR-92a-3p. The expression of the two miRNAs was significantly increased in colon and rectal cancer group compared with normal group, respectively (Supplementary Figure 4).
Expression of the identified miRNAs in serum exosomes
The expression of exosomal miR-19a-3p, miR-21-5p, and miR-425-5p was assessed by qRT–PCR in 10 serum samples from CRC patients and 10 NCs to explore the potential form of the identified miRNAs in serum of CRC patients. Compared to NCs, all the three miRNAs were up-regulated in CRC serum exosomes. Additionally, the three miRNAs were consistently expressed in colon and rectal cancers (Figure 5).
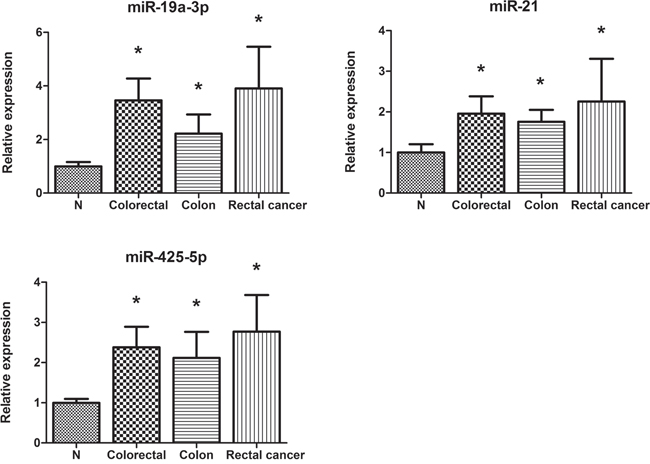
Figure 5: Expression of the three miRNAs in the serum exosomes of 10 CRC patients and 10 NCs. Error bar: standard error.
The expression of miR-122-5p and miR-92a-3p were also found to be significantly higher in CRC serum exosomes compared to NCs. And the two miRNAs were also consistently expressed in colon and rectal cancers (Supplementary Figure 5).
Comparison of miRNAs in peripheral and arterial serum
As blood flows from arterial to venous circulation, we assessed expression levels of the miRNAs in 5 arterial serum samples and compared those in the matched peripheral serum samples from the same individual. Among the three miRNAs, miR-21-5p showed higher expression levels in arterial plasma in more than a half of the subjects. However, due to the relatively small sample size, the results were not statistically significant (Supplementary Figure 6).
DISCUSSION
In this study, we analyzed a cohort of 334 serum samples from 196 patients with CRC and 138 healthy subjects in a four-step approach, and found that the expression levels of miR-19a-3p, miR-21-5p and miR-425-5p could predict CRC. A few recent studies have revealed the diagnostic role of circulating miRNAs in CRC [13, 14, 18]. Unfortunately, to date, independent studies have failed to develop clinically actionable miRNA biomarkers for a variety of reasons including population and sample diversity, especially the discovery and validation methodologies used in these reports. In the current study, we utilized Exiqon miRNA qPCR panels which appeared to show better sensitivity and linearity with measurements of miRNAs in relatively low abundance than TaqMan platform [19]. This platform was used in the screening phase to analyze differential expression profiling of serum miRNAs in 3 CRC and 1 NC pooled samples. The screening stage was followed by the training and testing phases of RT–qPCR validation in our study. We found that up-regulated levels of miR-19a-3p, miR-21-5p and miR-425-5p in serum could accurately discriminate patients with CRC from healthy controls. To ensure the reliability and reproducibility of the diagnostic value of the three -miRNA signature, we intensively validated our findings in the external validation stage. Notably, the expression of miR-19a-3p, miR-21-5p and miR-425-5p in CRC tissue samples was consistent with our serum results, which suggested the important roles for these miRNAs in tumorigenesis and progression of the disease.
Serum miR-19a-3p has been explored as a potentially noninvasive biomarker for colorectal cancer in two additional studies [14, 20]. As a member of the miR-17-92 cluster, miR-19a-3p could promote the proliferation and metastasis of colon cancer cells [21]. Another study reported that miR-19a-3p could enhance invasion and metastasis by targeting Transglutaminase-2 in CRC cells [22]. Serum miR-19a-3p could predict resistance to FOLFOX chemotherapy in advanced colorectal cancer cases [23]. miR-21-5p has been reported as a reliable diagnostic biomarker for colorectal cancer in several additional studies that is in accordance with our finding [13, 18, 24, 25]. Recent findings indicate that miR-21-5p might function as oncogene in various cancers, including CRC. miR-21-5p could regulate several different target genes and pathways involving tumor proliferation, invasion and metastasis and play an important role in 5-FU resistance and affect the radiosensitivity of CRC cells [26–28]. Especially, serum miR-21-5p level was found to be a robust independent predictor of the presence of CRC [29]. However, there was no significant association between serum miR-21-5p levels and clinical stage in patients with CRC in our study. miR-425-5p has been reported to be implicated in tumorigenesis in many cancer types [30–32]. However, the mechanism of miR-425-5p in CRC is still unclear. Zhang et al. showed that miR-425-5p could regulate chemoresistance in CRC cells via regulating Programmed Cell Death 10 [33]. In addition, miR-425-5p has been reported to promote tumorigenicity and aggressiveness in breast cancer and gastric cancer [31, 34]. miR-425-5p was discovered for the first time to be valuable biomarker of CRC in our study. The function of miR-425-5p in CRC is needed to be further investigated. We also found that serum miR-122-5p was up-regulated in colon cancer. miR-122-5p, a liver-specific miRNA that is abundant in the liver and plays an important role in regulating hepatocyte development and differentiation [35]. miR-122-5p has also been reported to regulate tumorigenesis in hepatocellular carcinoma and have tumor-suppressive function in breast cancer [36, 37]. Additionally, miR-92a-3p was upregulated in rectal cancer not colon cancer serum. miR-92a-3p was evaluated as a diagnostic biomarker in CRC in previous study [14, 38, 39]. As a member of large cluster miR-17-92, miR-92a-3p has an important role as an oncogene in some cancers [40, 41]. Upregulation of the miR-92a-3p has been shown to disrupt the functions of several important factors of growth and division in colorectal cells [42]. Schee K et al. suggested that miR-92a-3p was involved in the metastasis of CRC by targeting the CDH1 gene [43]. miR-122-5p and miR-92a-3p may be as potentially biomarker for discriminating colon cancer from rectal cancer. Interestingly, the five miRNAs expression were found to be consistently higher in colon and rectal cancer tissues, while the analysis of miRNA expression by The Cancer Genome Atlas (TCGA) project identified no clear distinctions between rectal cancers and colon cancers tissues [44], which suggested that there might be some difference of the miRNA expression profile between serum and tissue samples in CRC. Of course, further research of these miRNAs in CRC formation and development is needed.
Certainly, these circulating miRNAs identified in our study were also associated with some other cancers. High serum miR-19a-3p expression correlates with worse prognosis of patients with non-small cell lung cancer [45]. miR-21-5p acts as a broad-spectrum biomarker for many other solid cancers, including gastric, pancreatic, breast, and prostate cancers [11, 46–48]. One advantage characteristic for the serum miR-21-5p level is its relatively higher sensitivity and specificity in diagnosis of CRC than both NCs and other cancer patients [49, 50]. miR-425-5p has been identified as a potential biomarker in renal cell carcinoma, lung squamous cell carcinoma, breast cancer and bladder cancer [30, 51, 52]. An up-regulation of circulating miR-425-5p has been observed in head and neck cancer patients after radiotherapy in the blood plasma compared with primary HNSCC (head and neck squamous cell carcinoma) cells [53]. This is a common concern with many other miRNA biomarkers, because of their simultaneous dysregulation in multiple cancers. Belonging to circulatory system, arterial blood is more difficult to obtain. We assessed the expression levels of the three miRNAs in arterial serum of only 5 patients. Unfortunately, no result achieved statistical significance due to the small sample size. Apparently, more efforts were needed to investigate the specificity of circulating miRNAs.
Given the fact that the majority of circulating miRNAs come from tumor tissue [54], we evaluated the expression of the three identified serum miRNAs in CRC tissues. It turned out that miR-19a-3p, miR-21-5p and 425-5p were found to be highly upregulated in CRC tissues compared to those in normal tissues. The findings may at least in part verify the theory. Exosomes are small membrane vesicles of approximately 30–140 nm that embed protein, lipids, mRNAs, and miRNAs, depending on the origin of the secreting cells [55]. A recent study discovered that miRNA expression in serum and saliva was predominantly derived from exosomes [56]. Moreover, recent studies demonstrated that exosomal miRNAs in serum may be promising diagnostic biomarkers for the detection of CRC [57]. We further explored serum exosomal miRNAs to identify the potential form of the three miRNAs in serum. miR-19a-3p, miR-21-5p, and miR-425-5p were up-regulated in CRC serum exosomes. miR-122-5p and miR-92a-3p were also found to be significantly higher in CRC exosomes compared with NCs. Other studies have showed that the majority of miRNA signal in serum is primarily contributed by free-floating RNAs. More studies are needed to resolve this issue.
In conclusion, a three-miRNA panel in the serum of CRC patients which could serve as a non-invasive biomarker in the detection of CRC was identified and validated. Our serum based miRNA biomarkers may be clinically applicable for noninvasive screening of patients with CRC.
MATERIALS AND METHODS
Study design, patients and controls
A total of 196 histopathologically conformed CRC patients and 138 normal controls (NCs) were enrolled between 2012 and 2014 at the First Affiliated Hospital of Nanjing Medical University (training and testing stage) and the Third Affiliated Hospital of Soochow University (external validation stage). People who showed no evidence of disease (including cancer, precancerous lesion and chronic diseases) were selected as normal controls. An additional of twenty-four paired formalin fixed paraffin embedded (FFPE) sections of CRC and matched adjacent gastric mucosa tissues were obtained. All the procedures were approved by Institutional Review Boards of the First Affiliated Hospital of Nanjing Medical University. Written informed consent was taken from each participant.
In this study, we carried out a 4-stage study for CRC. In the initial screening stage (Figure 1), thirty peripheral serum samples from CRC patients and 10 from NCs were randomly chosen and pooled as 3 CRC samples and 1 NC sample (10 samples were pooled as 1 pool sample, colon-to-rectal ratio 33 to 23) for miRNA microarrays. Approximately 20–25 ng RNA isolated from each pool of serum samples was reverse transcribed to cDNA by using the miRCURY Locked Nucleic Acid (LNA™) Universal Reverse Transcription (RT) microRNA PCR, Polyadenylation and cDNA synthesis kit (Exiqon miRNA qPCR panel, Vedbaek, Denmark) following the manufacturer’s protocol. Microarrays were scanned on 7900HT real-time PCR system (Applied Biosystems, Foster City, CA, USA) with Exiqon miRCURY-Ready-to-Use PCR-Human-panel-I + II-V1.M (Exiqon miRNA qPCR panel, Vedbaek, Denmark), which could detect 168 miRNAs in plasma/serum to identify differently expressed miRNAs. Melting curve analyses were performed at the end of the PCR cycles. Detectable miRNAs were those with a Ct < 37 and 5 Ct less than the negative control (No Template Control, NTC). An RNA spike-in (UniSp6) and a DNA spike-in (Sp3) were used as technical controls to evaluate if the technical performance of all samples is similar. The Ct values were normalized based on the average of the normalizer assays in the panel and this included miR-423-5p and miR-93-5p. The formula used to calculate the normalized Ct values is: normalized Ct (Δ Ct) = average Ct (assay) – average Ct (normalizer assays). The relative expression levels of miRNAs between CRC patients and NCs were calculated using 2-ΔΔCt method.
In the training stage, the differentially expressed miRNAs discovered via screening phase was confirmed using qRT-PCR in 30 CRC samples and 30 NCs. Then, the miRNAs identified by the training stage were further evaluated by qRT-PCR in the testing phase in serum samples including 136 CRC patients and 90 NCs. For the external validation set, we subjected 30 cases and 18 controls to evaluate the diagnostic value of the three-miRNA signature in CRC. The selected miRNAs were further verified in 12 pairs of formalin-fixed paraffin-embedded (FFPE) colon cancer tissue specimens and 12 pairs of rectal ones and adjacent nontumor tissues from surgery patients. In addition, exosomal miRNAs from 10 patients and NCs were analyzed to investigate the potential form of the miRNAs in the peripheral serum.
Whole blood samples (5ml) were collected from each patient and normal controls with overnight fasting. In CRC patients, blood samples were collected within one week before initial treatment. All samples were processed within 1 hour and stored in 4°C, then subjected to centrifuge at 1,500 r.p.m. for 10 min within 12 h after collection. Then, cell-free serum was further resolved by centrifugation at 1,2000 r.p.m. for 2 min to guarantee complete removal of cell debris. The serum sample was stored in an RNase-free eppendorf tube at -80°C until use.
RNA extraction
RNA was isolated from 200μl serum using mirVana Paris Kit (Ambion, Austin, TX, USA) according to the manufacturer’s protocol. To allow for normalization of sample-to-sample variation in RNA extraction procedures, synthetic C. elegans miRNA cel-miR-39 (5 nM/L, 5μl RiboBio, Guangzhou, China) was spiked into each denatured sample after combining the serum sample with denaturing solution (Ambion, Austin, TX, USA). Total RNA was extracted from FFPE specimens using the High Pure FFPE RNA Micro Kit (Ambion, Austin, TX, USA). RNA was eluted with 100μl of RNase-free water and stored at − 80°C for further use. The ultraviolet spectrophotometer was used to evaluate the concentration and purity of the total RNA. The concentration of serum RNA ranged from 15.73 ng/μl to 48.62 ng/μl.
Isolation of exosomes
ExoQuick Exosome Precipitation Solution (System Biosciences, Mountain View, Calif)was used to isolate exosomes from serum according to the manufacturer’s protocol. Briefly, 200μl serum was mixed with 100μl ExoQuick exosome precipitation solution and then kept at 4°C for 30 min, followed by centrifugation at 13,000 rpm for 2 min. After the supernatants were removed, the exosome pellets were retained for further RNA extraction.
Quantitative reverse transcription polymerase chain reaction (qRT-PCR)
The amplification of miRNA was performed using the specific primers of reverse transcription (RT) and polymerase chain reaction (PCR) from Bulge-Loop™ miRNA qRT-PCR Primer Set (RiboBio, Guangzhou, China) as previously described [15, 16]. The quantification of PCR product was evaluated by the level of fluorescence in emitted by SYBR Green (SYBR® Premix Ex Taq™ II, TaKaRa). RT and PCR were performed as previously described [15]. RT reactions were carried out at 42°C for 60 min followed by 70°C for 10 min. The qRT-PCR was run on a LightCycler® 480 Real-Time PCR System (Roche Diagnostics, Mannheim, Germany) in 384-well plates at 95°C for 20 sec, followed by 40 cycles of 95°C for 10 sec, 60°C for 20 sec and then 70°C for 10 sec. The melting analysis was added finally to evaluate the specificity of PCR products. The expression of miRNAs in tissue specimens, serum samples and exosomes were calculated using the comparative 2-ΔΔCt method relative to RNU6B (U6) and cel-miR-39 [17].
Statistical analysis
Differential miRNAs expression between CRC patients and NCs was analyzed using Mann-Whitney test. The association between miRNAs and the clinical characteristics was estimated by the one-way ANOVA or χ2 test. Receiver operating characteristic (ROC) curves and the area under the ROC curve (AUC) were used to evaluate the diagnostic value of the candidate miRNAs for CRC. Logistic regression model for CRC prediction was applied on the data from the training and validation phases. SPSS (version 15.0, SPSS Inc., Chicago, IL, USA) software was used to perform all statistical analysis. Differences were considered significant when p < 0.05.
ACKNOWLEDGMENTS AND FUNDING
This work was supported by the National Natural Science Foundation of China [Grant number: 81672400, 81672788, 81370516].
CONFLICTS OF INTEREST
The authors declare no competing financial interests.
REFERENCES
1. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015; 65: 87-108.
2. Akkoca AN, Yanik S, Ozdemir ZT, Cihan FG, Sayar S, Cincin TG, Cam A, Ozer C. TNM and Modified Dukes staging along with the demographic characteristics of patients with colorectal carcinoma. Int J Clin Exp Med. 2014; 7: 2828-2835.
3. Lieberman D. Colorectal Cancer Screening With Colonoscopy. JAMA INTERN MED. 2016; 176:903-904.
4. Bretthauer M. Colorectal cancer screening. J Intern Med. 2011; 270: 87-98.
5. De Maio G, Rengucci C, Zoli W, Calistri D. Circulating and stool nucleic acid analysis for colorectal cancer diagnosis. World J Gastroenterol. 2014; 20: 957-967.
6. Ransohoff DF, Sox HC. Clinical Practice Guidelines for Colorectal Cancer Screening: New Recommendations and New Challenges. JAMA. 2016; 315: 2529-2531.
7. Knudsen AB, Zauber AG, Rutter CM, Naber SK, Doria-Rose VP, Pabiniak C, Johanson C, Fischer SE, Lansdorp-Vogelaar I, Kuntz KM. Estimation of Benefits, Burden, and Harms of Colorectal Cancer Screening Strategies: Modeling Study for the US Preventive Services Task Force. JAMA. 2016; 315: 2595-2609.
8. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004; 116: 281-297.
9. Zhou X, Zhu W, Li H, Wen W, Cheng W, Wang F, Wu Y, Qi L, Fan Y, Chen Y, Ding Y, Xu J, Qian J, Huang Z, Wang T, Zhu D, Shu Y, Liu P. Diagnostic value of a plasma microRNA signature in gastric cancer: a microRNA expression analysis. Sci Rep. 2015; 5: 11251.
10. Zhang C, Wang C, Chen X, Yang C, Li K, Wang J, Dai J, Hu Z, Zhou X, Chen L, Zhang Y, Li Y, Qiu H, Xing J, Liang Z, Ren B, Yang C, Zen K, Zhang CY. Expression profile of microRNAs in serum: a fingerprint for esophageal squamous cell carcinoma. Clin Chem. 2010; 56: 1871-1879.
11. Liu R, Chen X, Du Y, Yao W, Shen L, Wang C, Hu Z, Zhuang R, Ning G, Zhang C, Yuan Y, Li Z, Zen K, Ba Y, Zhang CY. Serum microRNA expression profile as a biomarker in the diagnosis and prognosis of pancreatic cancer. Clin Chem. 2012; 58: 610-618.
12. Geng Q, Fan T, Zhang B, Wang W, Xu Y, Hu H. Five microRNAs in plasma as novel biomarkers for screening of early-stage non-small cell lung cancer. Respir Res. 2014; 15: 149.
13. Yamada A, Horimatsu T, Okugawa Y, Nishida N, Honjo H, Ida H, Kou T, Kusaka T, Sasaki Y, Yagi M, Higurashi T, Yukawa N, Amanuma Y, Kikuchi O, Muto M, Ueno Y, Nakajima A, Chiba T, Boland CR, Goel A. Serum miR-21, miR-29a, and miR-125b Are Promising Biomarkers for the Early Detection of Colorectal Neoplasia. Clin Cancer Res. 2015; 21: 4234-4242.
14. Zheng G, Du L, Yang X, Zhang X, Wang L, Yang Y, Li J, Wang C. Serum microRNA panel as biomarkers for early diagnosis of colorectal adenocarcinoma. Br J Cancer. 2014; 111: 1985-1992.
15. Zhao DS, Chen Y, Jiang H, Lu JP, Zhang G, Geng J, Zhang Q, Shen JH, Zhou X, Zhu W, Shan QJ. Serum miR-210 and miR-30a expressions tend to revert to fetal levels in Chinese adult patients with chronic heart failure. Cardiovasc Pathol. 2013; 22: 444-450.
16. Du Y, Zhu M, Zhou X, Huang Z, Zhu J, Xu J, Cheng G, Shu Y, Liu P, Zhu W, Wang T. miR-20a enhances cisplatin resistance of human gastric cancer cell line by targeting NFKBIB. Tumour Biol. 2016; 37: 1261-1269.
17. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001; 25: 402-408.
18. Wang J, Huang SK, Zhao M, Yang M, Zhong JL, Gu YY, Peng H, Che YQ, Huang CZ. Identification of a circulating microRNA signature for colorectal cancer detection. PLoS One. 2014; 9: e87451.
19. Jensen SG, Lamy P, Rasmussen MH, Ostenfeld MS, Dyrskjot L, Orntoft TF, Andersen CL. Evaluation of two commercial global miRNA expression profiling platforms for detection of less abundant miRNAs. BMC Genomics. 2011; 12: 435.
20. Zekri AR, Youssef AS, Lotfy MM, Gabr R, Ahmed OS, Nassar A, Hussein N, Omran D, Medhat E, Eid S, Hussein MM, Ismail MY, Alenzi FQ, Bahnassy AA. Circulating Serum miRNAs as Diagnostic Markers for Colorectal Cancer. PLoS One. 2016; 11: e0154130.
21. Yu G, Li H, Wang X, Wu T, Zhu J, Huang S, Wan Y, Tang J. MicroRNA-19a targets tissue factor to inhibit colon cancer cells migration and invasion. Mol Cell Biochem. 2013; 380: 239-247.
22. Cellura D, Pickard K, Quaratino S, Parker H, Strefford JC, Thomas GJ, Mitter R, Mirnezami AH, Peake NJ. miR-19-Mediated Inhibition of Transglutaminase-2 Leads to Enhanced Invasion and Metastasis in Colorectal Cancer. Mol Cancer Res. 2015; 13: 1095-1105.
23. Chen Q, Xia HW, Ge XJ, Zhang YC, Tang QL, Bi F. Serum miR-19a predicts resistance to FOLFOX chemotherapy in advanced colorectal cancer cases. Asian Pac J Cancer Prev. 2013; 14: 7421-7426.
24. Basati G, Emami Razavi A, Abdi S, Mirzaei A. Elevated level of microRNA-21 in the serum of patients with colorectal cancer. Med Oncol. 2014; 31: 205.
25. Liu GH, Zhou ZG, Chen R, Wang MJ, Zhou B, Li Y, Sun XF. Serum miR-21 and miR-92a as biomarkers in the diagnosis and prognosis of colorectal cancer. Tumour Biol. 2013; 34: 2175-2181.
26. Asangani IA, Rasheed SA, Nikolova DA, Leupold JH, Colburn NH, Post S, Allgayer H. MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene. 2008; 27: 2128-2136.
27. Deng J, Lei W, Fu JC, Zhang L, Li JH, Xiong JP. Targeting miR-21 enhances the sensitivity of human colon cancer HT-29 cells to chemoradiotherapy in vitro. Biochem Biophys Res Commun. 2014; 443: 789-795.
28. Zhang J, Xiao Z, Lai D, Sun J, He C, Chu Z, Ye H, Chen S, Wang J. miR-21, miR-17 and miR-19a induced by phosphatase of regenerating liver-3 promote the proliferation and metastasis of colon cancer. Br J Cancer. 2012; 107: 352-359.
29. Menendez P, Padilla D, Villarejo P, Palomino T, Nieto P, Menendez JM, Rodriguez-Montes JA. Prognostic implications of serum microRNA-21 in colorectal cancer. J Surg Oncol. 2013; 108: 369-373.
30. Wang J, Li Z, Ge Q, Wu W, Zhu Q, Luo J, Chen L. Characterization of microRNA transcriptome in tumor, adjacent, and normal tissues of lung squamous cell carcinoma. J Thorac Cardiovasc Surg. 2015; 149: 1404-1414 e1404.
31. Zhang Z, Li Y, Fan L, Zhao Q, Tan B, Li Z, Zang A. microRNA-425-5p is upregulated in human gastric cancer and contributes to invasion and metastasis in vitro and in vivo. Exp Ther Med. 2015; 9: 1617-1622.
32. Fleming NH, Zhong J, da Silva IP, Vega-Saenz de Miera E, Brady B, Han SW, Hanniford D, Wang J, Shapiro RL, Hernando E, Osman I. Serum-based miRNAs in the prediction and detection of recurrence in melanoma patients. Cancer. 2015; 121: 51-59.
33. Zhang Y, Hu X, Miao X, Zhu K, Cui S, Meng Q, Sun J, Wang T. MicroRNA-425-5p regulates chemoresistance in colorectal cancer cells via regulation of Programmed Cell Death 10. J Cell Mol Med. 2016; 20: 360-369.
34. Di Leva G, Piovan C, Gasparini P, Ngankeu A, Taccioli C, Briskin D, Cheung DG, Bolon B, Anderlucci L, Alder H, Nuovo G, Li M, Iorio MV, Galasso M, Santhanam R, Marcucci G, Perrotti D, Powell KA, Bratasz A, Garofalo M, Nephew KP, Croce CM. Estrogen mediated-activation of miR-191/425 cluster modulates tumorigenicity of breast cancer cells depending on estrogen receptor status. PLoS Genet. 2013; 9: e1003311.
35. Morita K, Taketomi A, Shirabe K, Umeda K, Kayashima H, Ninomiya M, Uchiyama H, Soejima Y, Maehara Y. Clinical significance and potential of hepatic microRNA-122 expression in hepatitis C. Liver Int. 2011; 31: 474-484.
36. Ergun S, Ulasli M, Igci YZ, Igci M, Kirkbes S, Borazan E, Balik A, Yumrutas O, Camci C, Cakmak EA, Arslan A, Oztuzcu S. The association of the expression of miR-122-5p and its target ADAM10 with human breast cancer. Mol Biol Rep. 2015; 42: 497-505.
37. Tsai WC, Hsu SD, Hsu CS, Lai TC, Chen SJ, Shen R, Huang Y, Chen HC, Lee CH, Tsai TF, Hsu MT, Wu JC, Huang HD, Shiao MS, Hsiao M, Tsou AP. MicroRNA-122 plays a critical role in liver homeostasis and hepatocarcinogenesis. J Clin Invest. 2012; 122: 2884-2897.
38. Ng EK, Chong WW, Jin H, Lam EK, Shin VY, Yu J, Poon TC, Ng SS, Sung JJ. Differential expression of microRNAs in plasma of patients with colorectal cancer: a potential marker for colorectal cancer screening. Gut. 2009; 58: 1375-1381.
39. Huang Z, Huang D, Ni S, Peng Z, Sheng W, Du X. Plasma microRNAs are promising novel biomarkers for early detection of colorectal cancer. Int J Cancer. 2010; 127: 118-126.
40. Ren P, Gong F, Zhang Y, Jiang J, Zhang H. MicroRNA-92a promotes growth, metastasis, and chemoresistance in non-small cell lung cancer cells by targeting PTEN. Tumour Biol. 2016; 37: 3215-3225.
41. Arques O, Chicote I, Puig I, Tenbaum SP, Argiles G, Dienstmann R, Fernandez N, Caratu G, Matito J, Silberschmidt D, Rodon J, Landolfi S, Prat A, Espin E, Charco R, Nuciforo P, Vivancos A, Shao W, Tabernero J, Palmer HG. Tankyrase Inhibition Blocks Wnt/beta-Catenin Pathway and Reverts Resistance to PI3K and AKT Inhibitors in the Treatment of Colorectal Cancer. Clin Cancer Res. 2016; 22: 644-656.
42. Schee K, Boye K, Abrahamsen TW, Fodstad O, Flatmark K. Clinical relevance of microRNA miR-21, miR-31, miR-92a, miR-101, miR-106a and miR-145 in colorectal cancer. BMC Cancer. 2012; 12: 505.
43. Lin Q, Chen T, Lin Q, Lin G, Lin J, Chen G, Guo L. Serum miR-19a expression correlates with worse prognosis of patients with non-small cell lung cancer. J Surg Oncol. 2013; 107: 767-771.
44. Liu C, Zhang Y, Chen H, Jiang L, Xiao D. Function analysis of rs9589207 polymorphism in miR-92a in gastric cancer. Tumour Biol. 2016; 37: 4439-4444.
45. Li BS, Zhao YL, Guo G, Li W, Zhu ED, Luo X, Mao XH, Zou QM, Yu PW, Zuo QF, Li N, Tang B, Liu KY, Xiao B. Plasma microRNAs, miR-223, miR-21 and miR-218, as novel potential biomarkers for gastric cancer detection. PLoS One. 2012; 7: e41629.
46. Asaga S, Kuo C, Nguyen T, Terpenning M, Giuliano AE, Hoon DS. Direct serum assay for microRNA-21 concentrations in early and advanced breast cancer. Clin Chem. 2011; 57: 84-91.
47. Yaman Agaoglu F, Kovancilar M, Dizdar Y, Darendeliler E, Holdenrieder S, Dalay N, Gezer U. Investigation of miR-21, miR-141, and miR-221 in blood circulation of patients with prostate cancer. Tumour Biol. 2011; 32: 583-588.
48. Kanaan Z, Rai SN, Eichenberger MR, Roberts H, Keskey B, Pan J, Galandiuk S. Plasma miR-21: a potential diagnostic marker of colorectal cancer. Ann Surg. 2012; 256: 544-551.
49. Wang B, Zhang Q. The expression and clinical significance of circulating microRNA-21 in serum of five solid tumors. J Cancer Res Clin Oncol. 2012; 138: 1659-1666.
50. Ge YZ, Xin H, Lu TZ, Xu Z, Yu P, Zhao YC, Li MH, Zhao Y, Zhong B, Xu X, Zhou LH, Wu R, Xu LW, Wu JP, Li WC, Zhu JG, Jia RP. MicroRNA expression profiles predict clinical phenotypes and prognosis in chromophobe renal cell carcinoma. Sci Rep. 2015; 5: 10328.
51. Scheffer AR, Holdenrieder S, Kristiansen G, von Ruecker A, Muller SC, Ellinger J. Circulating microRNAs in serum: novel biomarkers for patients with bladder cancer? World J Urol. 2014; 32: 353-358.
52. Summerer I, Niyazi M, Unger K, Pitea A, Zangen V, Hess J, Atkinson MJ, Belka C, Moertl S, Zitzelsberger H. Changes in circulating microRNAs after radiochemotherapy in head and neck cancer patients. Radiat Oncol. 2013; 8: 296.
53. Zen K, Zhang CY. Circulating microRNAs: a novel class of biomarkers to diagnose and monitor human cancers. Med Res Rev. 2012; 32: 326-348.
54. Cancer Genome Atlas N. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012; 487: 330-337.
55. Simpson RJ, Lim JW, Moritz RL, Mathivanan S. Exosomes: proteomic insights and diagnostic potential. Expert Rev Proteomics. 2009; 6: 267-283.
56. Turchinovich A, Weiz L, Langheinz A, Burwinkel B. Characterization of extracellular circulating microRNA. Nucleic Acids Res. 2011; 39: 7223-7233.
57. Ogata-Kawata H, Izumiya M, Kurioka D, Honma Y, Yamada Y, Furuta K, Gunji T, Ohta H, Okamoto H, Sonoda H, Watanabe M, Nakagama H, Yokota J, Kohno T, Tsuchiya N. Circulating exosomal microRNAs as biomarkers of colon cancer. PLoS One. 2014; 9: e92921.

