INTRODUCTION
The combination of surgical operation with radiotherapy and adjuvant intravesical administration is the conventional treatment strategy for human bladder cancer (HBC) [1]. The ideal intravesical agent should be sensitive to HBC cells, and could be absorbed easily in the carcinoma cells but exert fewer side effects on normal tissue cells. Resveratrol (3,5,4’-trihydroxy-trans-stilbene, RV, Supplementary Figure 1A), a natural dietary polyphenol, possesses anti-cancer and other beneficial pharmacological activities [2–4]. Furthermore, the lipophilic characteristic of RV caused by its basic structural skeleton of the central carbon-carbon double bond conjugated with two benzene rings (Supplementary Figure 1A), which would lead it more easily to be absorbed by the bladder mucosa cells, and thus could reach the effective drug concentration in the carcinoma cells and exert better biological activities [5, 6]. With the characteristics of adjuvant intravesical therapy for bladder cancer, RV may be a viable candidate, especially in bladder cancer perfusion chemotherapy.
Since Jang et al. demonstrated that RV possessed cancer chemopreventive activity [3], studies on the bioactivity of RV have increased rapidly [7–9], and RV’s anti-tumor effect represents some of the most convincing and intriguing [7–10]. It was reported that RV could restore PTEN expression by targeting oncomiRs of the miR-17 family in prostate cancer [11], and also could inhibit STAT3 activation, enhancing autophagy and apoptosis in rat orthotopic glioblastoma [12]. Short-term exposure RV could cause growth inhibition and apoptosis of HBC EJ cells in vitro and in vivo [13]. And as we all know, RV could be metabolized rapidly and produce various metabolites such as RV glucuronide or/and RV sulfate conjugates (Supplementary Figure 1) [14–18]. It was found that RV could be metabolized to RV sulfates in human breast cancer MB-MDA-231 and ZR-75-1 cells [14], human medulloblastoma UW228-3 [17], human glioblastoma LN-18 and U251 cells [19, 20]. However, RV glucuronide was found as the main metabolite in rat glioblastoma RG2 and C6 cells, and showed discrepant metabolic patterns between human and rat glioblastoma cells [20]. So far, little work has been carried out to explore the metabolism of RV in HBC EJ and T24 cells. Thus, how RV exerts its bioactivity in bladder cancer becomes an interesting issue, either by RV parent compound or its metabolites, or both RV and its metabolites synergistically exert the beneficial effect? To clarify this ambiguity, we analyzed RV’s metabolic pattern in HBC T24 and EJ cells, then biotransformed its major metabolite in vitro and tested its bioactivity to ascertain the effective bioactive form of RV, and further checked the safety of the active compound at the therapeutic dosage to evaluate RV’s clinic medicinal value.
RESULTS
Responses of BC cells to RV
To explore the biological activity and the effective dosage of RV in HBC T24 and EJ cells, MTT assay was carried out. As shown in Figure 1A (left), after incubation with 100μM RV for 6h, 12h, 24h, 48h and 72h, the inhibition ratio of T24 cells was 15.3±0.3 %, 13.6±0.3 %, 16.5±1.8 %, 58.5±1.5 % and 76.6±1.6 %, respectively. While the inhibition ratio of EJ cells was 2.4±0.3 %, 2.5±0.2 %, 15.1±1.1 %, 20.1±1.5 % and 37.3±1.6 % after incubation with 100μM RV for 6h, 12h, 24h, 48h and 72h, respectively. The above results showed that RV could induce a significant time-dependent growth inhibition to T24 cells, but the proliferation of EJ cells was less suppressed (Figure 1A) [21]. Meanwhile, Figure 1A (right) also presented a concentration-dependent inhibition in T24 and EJ cells after incubation with 0, 20μM, 40μM, 60μM, 80μM, 100μM, 150μM and 200μM RV, respectively.
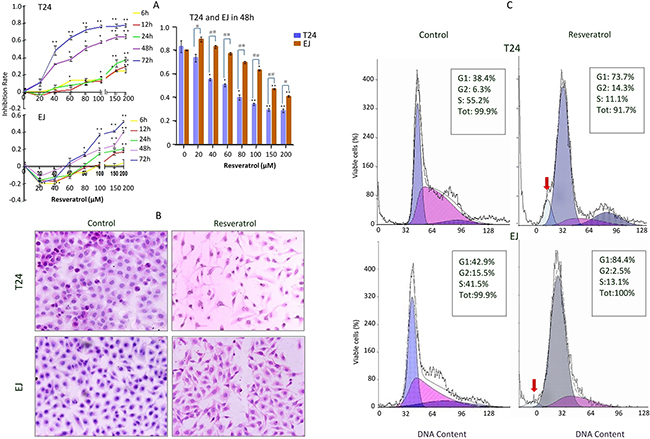
Figure 1: Chemosensitivity evaluation of resveratrol to T24 and EJ cells. A. Effect of resveratrol treatment on human bladder cancer (HBC) T24 and EJ cells. Cells were incubated with different concentrations (0, 20, 40, 60, 80, 100, 150 and 200μM) resveratrol for different time periods (0, 6, 12, 24, 48 and 72h), respectively, and then the cells number was determined by MTT as described in the Materials and Methods. Data are presented as means ± S.D. of three independent experiments. Bars means standard errors, *P<0.05, **P<0.001 reveal significant difference between RV-treatment and Control HBC cells. #P<0.05, ##P<0.001 show significant different between T24 RV-treatment cells and EJ RV-treatment cells. B. HE morphological staining performed on T24 and EJ cells without (Control) and with 100μM RV (Resveratrol) incubation for 48 hours (100×). Cells at a density of 4×105 cells per well were placed in dishes with coverslips, then T24 and EJ cells were treated without (Control) and with (Resveratrol) 100μM resveratrol treatment for 48h. Cells coverslips were harvested for examination and T24 cells exhibited more obviously spindle-shaped change than EJ cells. C. Flow cytometry analysis on the fractionation of cell cycles and apoptotic cells in T24 and EJ cell populations without (Control) and with (Resveratrol) 100μM resveratrol incubation for 48 hours. Red arrows, indicate the peak of apoptotic cells. Data revealed a presentative experiment in triplicate with similar results.
The RV-sensitivity of HBC cells was further evaluated by hematoxylin and eosin (HE) staining, as shown in Figure 1B, we found the majority of T24 cells presented spindle-shaped, segments of cell bodies, and detached from the culture plate after exposure to 100μM RV for 48h. But compared with T24 cells, there was no obvious morphologic change in EJ cells. And the 100μM-RV 48h-treatment was used for the further experiments. Flow cytometry (FCM) analyses showed that the G1 and S fractions were 38.4% and 55.2% in normally cultured T24 cells, but changed to 73.7% and 11.1% after 100μM RV treatment (Figure 1C). The percentages of G1 and S phase of EJ cells were 42.9% and 41.5% under normal culture condition and become 84.4% and 13.1% after 100μM-RV treatment for 48h. The above results indicated RV could induce G1 phase cell cycle arrest in HBC cells (Figure 1C).
RV monosulfate (RVS) was the major metabolite in HBC cells
To identify the RV’s metabolite(s) in HBC T24 and EJ cells, the cells and the conditioned media were collected after 100μM RV incubation for 48h, then were purified with solid phase extraction (SPE) to eliminate the interferer, and subsequently were analyzed by high performance liquid chromatography (HPLC), liquid chromatography-mass spectrometry (LC-MS) and high-resolution mass spectrometry (HRMS). For the metabolic profile of the T24 cells is similar to EJ cells (Supplementary Table 1, Supplementary Figure 2), we only showed the identification of T24 cells and its culture media (Figure 2). The T24 cell-free media which only contained RV was incubated for 48h as the background control. HPLC analyses showed that only one compound, trans-RV, was found in the standard group (Figure 2A(a), M1); two compounds, trans-RV and cis-RV, were presented in the control group (Figure 2A(b), M1, M2); and three major compounds could be detected both in T24 cell lysates (Figure 2A(c), M1, M2, M3) and condition media of T24 cells (Figure 2A(d), M1, M2, M3), which were later proved to be trans-RV, cis-RV and RVS according to their retention time and molecular weight. Compared with the above data, the T24 cells treated without RV showed no compound peak as control (Figure 2A(e)).
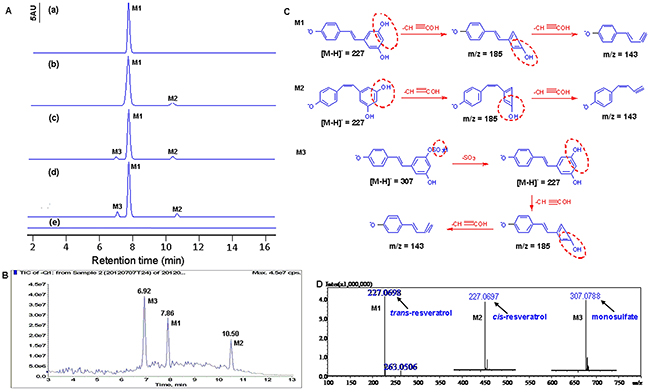
Figure 2: Identification of resveratrol’s metabolites in HBC T24 cells. A. HPLC chromatography analysis. (a) trans-resveratrol standard was dissolved in methanol and analyzed by HPLC (M1, tR=7.86); (b) The culture media incubation with resveratrol without T24 cells for 48h (M1, tR=7.86; M2, tR=7.02); (c) The T24 cells lysate was analyzed after incubation with 100μM resveratrol for 48h (M1, tR=7.89; M2, tR=7.06; M3, tR=10.38); (d) The supernatant of T24 cells was analyzed after incubation with 100μM resveratrol for 48h (M1, tR=7.87; M2, tR=7.01; M3, tR=10.41). (e) The T24 cells treated without resveratrol as Control. B. MS analyses of resveratrol metabolites in T24 cells. Total ion chromatogram (TIC) of the supernatant of T24 cells treated with 100μM resveratrol for 48h. Peak M1, M2 and M3 indicated retention time corresponding to different mass composition of metabolites; C. Proposed mechanism for the decomposition of the m/z 227 [M-H]- ion of resveratrol, the decomposition of the m/z 227 [M-H]- and m/z 307 [M-H]- ion of metabolites. D. Shimadzu LC-MS-IT-TOF-based HRMS analysis of resveratrol metabolites in T24 cells. Arrows labeled as M1, M2 and M3 indicated the exact [M-H]- molecular ion weight of 227.0698 (C14H11O3, calculated m/z 227.0708), 227.0697 (C14H11O3, calculated m/z 227.0708), 307.0788 (C14H11SO6, calculated m/z 307.0276), respectively. In Figure 3, M1 represents trans-resveratrol, M2 represents cis-resveratrol and M3 represents resveratrol monosulfate (RVS), respectively.
The above three major molecules were identified by LC-MS/MS using a combination of full and selected ion scanning techniques. Total ion chromatogram (TIC) of the T24 cells treated with 100μM RV for 48h were listed in Figure 2B, the peaks of M1, M2 and M3 could be detected from the chromatograms that represent trans-RV (M1), cis-RV (M2) and RVS (M3), respectively.
For further identify the RV metabolite(s) in HBC T24 and EJ cells, a combination of full and selected ion scanning of MS coupled with LC techniques was used. As illustrated in Figure 2C, the [M-H]– spectrum of M1 characterized by its molecular ion at m/z 227 which generated a series of fragment ions at 185 and 143. The fragment ion at m/z 185 was generated from m/z 227 after loss of 42amu (C2H2O), and the m/z 185 was further fragmented to m/z 143 after the loss of 42amu (C2H2O), which corresponded to trans-RV (Figure 2C, M1). Another metabolite (M2) was considered as an isomeric RV with mass spectral features identical to M1, and the fragment ions showed m/z at 185 and 143 attributed to cis-RV (Figure 2C, M2). The [M-H]- ion of M3 showed the dissociation molecule ions of m/z 307 and 227, respectively, the ion corresponding to RV (m/z 227) after losing 80amu, a sulfate moiety, from the RVS, then the m/z 227 was fragmented to m/z 185 for the further loss of 42amu (C2H2O) from RV, which appeared to be the main characteristic fragmentation pathway of RVS (Figure 2C, M3), and was also reported somewhere else [17, 22].
HRMS was applied to further confirm the RV metabolite(s), which showed the [M-H]– molecular ion exact mass as 227.0698 (C14H11O3, calculated m/z 227.0708), 227.0697 (C14H11O3, calculated m/z 227.0708) and 307.0788 (C14H11SO6, calculated m/z 307.0276), which was consistent with the report of LC-MS/MS and corresponded to trans-RV, cis-RV and RVS, respectively (Figure 2D) [17].
RV metabolic process in HBC cells
To evaluate the correlation between RV metabolism and its pharmaceutical activity, the RV metabolites in T24 and EJ cells and their supernatant were analyzed by HPLC, and the cell morphology was evaluated by HE staining (Figure 3). The supernatant and lysate of T24 cells were collected for HPLC analysis after 100μM RV treatment for 3h, 6h, 9h, 12h, 24h and 48h, respectively (Figure 3A). The HPLC results showed that RVS peak was observed as early as 3h after drug treatment (Figure 3C, 3D), but the T24 cells showed neither growth arrest nor morphological change until 24h-100μM RV treatment (Figure 3B), and the RVS didn’t dominate in the supernatant and lysate (Figure 3C, 3D), although T24 cells showed distinctly growth inhibition in 48h-RV treatment (Figure 3B), which suggested that the RV metabolism was preceded to growth inhibition in T24 cells. Meanwhile, EJ cells showed a similar RV metabolic pattern with T24 cells. However, compared with T24 cells, EJ cells still did show neither obvious growth arrest nor morphological change till 48h RV-treatment (Supplementary Figure 2).
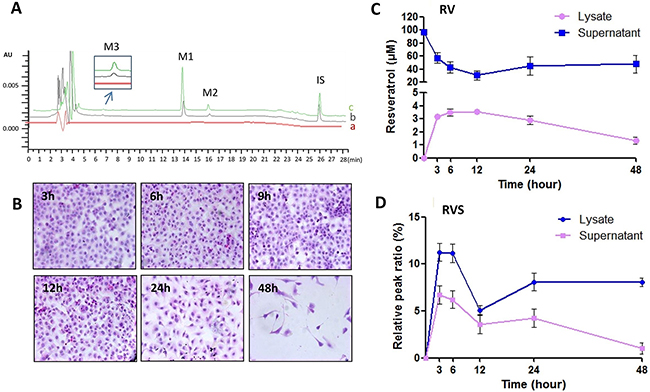
Figure 3: RV metabolic pattern in HBC T24 cells. A. Representative HPLC/DAD analysis of resveratrol in T24 cells. (a) HBC T24 cells treated without RV as Control; (b, c) T24 cells treated with 100μM resveratrol for 48h, cell supernatant (b) and cell lysates (c) spiked with 1,8-dihydroxy anthraquinone (internal standard, IS). Peaks: M1. trans-resveratrol, tR=13.82min; M2. cis-resveratrol, tR=15.97min; M3. resveratrol monosulfate (RVS), tR=6.67min; IS. 1,8-dihydroxy anthraquinone, tR=25.93min (Internal standard/IS). B. Morphologic changes were evaluated by HE staining (100×), and T24 cells showed neither observable growth arrest nor morphological change until 24h resveratrol incubation. C & D. Quantification of RV and RVS in T24 cells. Resveratrol concentrations in the cell lysates and supernatant after 100μM resveratrol treatment for 3, 6, 12, 24 and 48h, respectively.
RV upregulated SULT1A1 expression
Sulfation is an important metabolic pathway for xenobiotics and is catalyzed by the cytosolic sulfotransferases (SULTs). SULT1A1 appears to be an important phenol SULT because of its abundance and distribution in many tissues and wide substrate specificity [23, 24]. As shown in Figure 4, SULT1A1 expressed in normally cultured HBC T24 and EJ cells, and the densitometry scan of Western blots revealed that the SULT1A1 expression in RV-treated T24 cells (RV) increased about 1.7-fold and increased about 1.3-fold higher than that in RV-treated EJ cells (Figure 4A). The above results were further confirmed by PCR. RT-PCR showed the expression of SULT1A1 in RV-treated T24 and EJ cells was upregulated approximately 2-fold and 1.5-fold higher (Figure 4B), and was enhanced about 2.3-fold and 2.2-fold in real-time PCR (Figure 4C), respectively. ICC staining showed that SULT1A1 expressed in the cytoplasm of T24 and EJ cells, and the expression was both upregulated after 100μM RV treatment (Figure 4D).
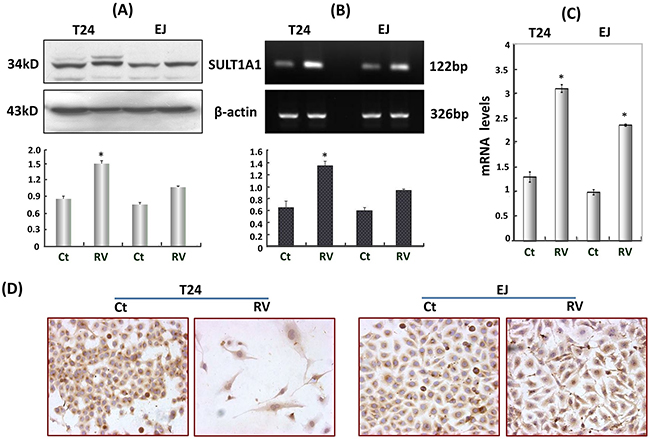
Figure 4: Resveratrol upregulated SULT1A1 expression in T24 and EJ cells. A. Western blots, B. RT-PCR, C. Real-time PCR and D. ICC (100×) all showed that SULT1A1 was upregulated in T24 and EJ cells after resveratrol treatment. Ct, RV represented HBC cells treated without and with 100μM resveratrol incubation for 48h, respectively. * P<0.05, represents statistical significance between RV-treatment HBC cells and the normally cultured HBC cells, respectively.
Decreased anticancer effects of RVS in HBC cells
RVS was the main metabolite in T24 cells, and SULT1A1 distributed widely in many tissues. RVS was prepared with the homogenate of rat livers in vitro and identified by HPLC and LC/MS. The MS/MS analysis showed RVS was prepared successfully (Table 1), and HPLC analysis revealed that the RVS was the major component and about 84.13% of parent trans-resveratrol was biotransformed according to the chromatogram peak area (Figure 5A). T24 cells were treated with a final concentration of 100μM RVS mixture for 48h, but different from 100μM RV treatment, T24 cells showed neither distinct growth suppression nor the signs of cell apoptosis (Figure 5B, 5C).
Table 1: LC-MS/MS analysis of the compounds of resveratrol biotransformation
MS1 Ions (m/z) |
MS2 Product Ions |
Identification |
|
|---|---|---|---|
m/z |
Fragment Loss |
||
227 |
185 |
[M-H-42]- |
standard resveratrol |
143 |
[M-H-84]- |
||
227 |
185 |
[M-H-42]- |
trans-resveratrol |
143 |
[M-H-84]- |
||
227 |
185 |
[M-H-42]- |
cis-resveratrol |
143 |
[M-H-84]- |
||
307 |
227 |
[M-H-80]- |
resveratrol monosulfate |
185 |
[M-H-80-42]- |
||
143 |
[M-H-80-84]- |
||
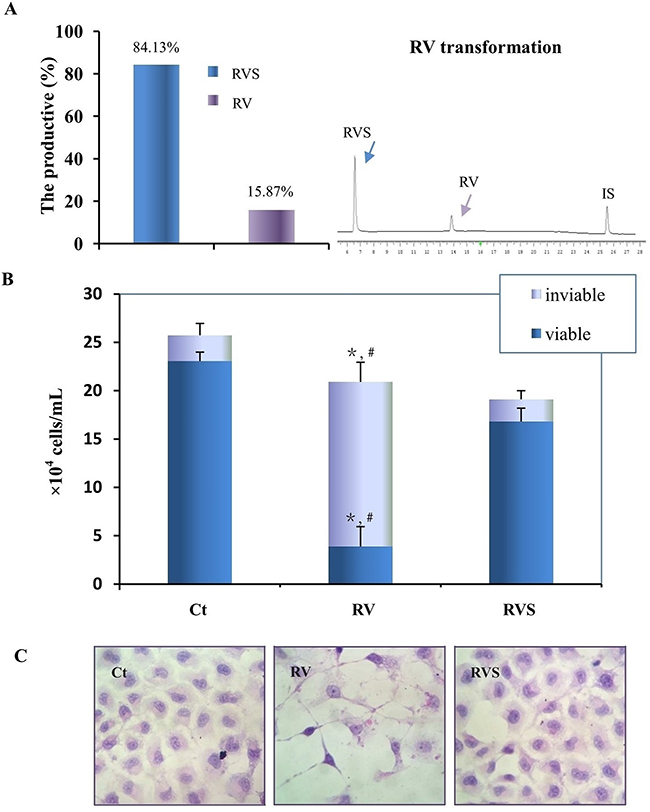
Figure 5: Biotransformation and bioactivity evaluation of RVS on T24 cells. A. Quantification of the biotransformation efficiency of RVS (Left) by representative HPLC analysis (Right). RV, RVS and IS represent resveratrol, resveratrol monosulfate and 1,8-dihydroxy anthraquinone (Internal standard), respectively. B. Cells number was determined by Trypan Blue exclusion after normal culture (Ct), 100μM trans-resveratrol (RV), and 84.13% resveratrol monosulfate/15.87% trans-resveratrol mixture (RVS) incubation for 48h, respectively. The column indicates the number of viable cells. *, #, RV treatment compared with Ct and RVS treatment, respectively (P<0.01). C. Morphologic evaluation of T24 cells incubated with normally culture (Ct), 100μM trans-resveratrol (RV), and resveratrol monosulfate/trans-resveratrol mixture (RVS) for 48h by HE staining (200×).
RV showed almost no side-effect to PBC cells
To explore whether RV has side-effect on primary cultured normal rat bladder epithelial cells (PBC), MTT and HE assay were carried out. The total number of PBC cells were about 250 000 (range: 250 000 cells/bladder to 350 000 cells/bladder) collected from twelve bladders. MTT results showed that the proliferation of PBC cells was not inhibited by 100μM-RV treatment, even tolerated as high concentration as 200μM-RV treatment (Figure 6A). After 100μM RV treatment for 48 hours, the condition media of the PBC cells were clear and the cells number increased. Compared with 48h-100μM RV treated T24 cells (Figure 5C), HE staining showed that PBC cells neither observable growth arrest nor morphological change after 48h-100μM RV incubation (Figure 6B), and the effective antitumor dosage 100μM RV displayed almost no side effect on PBC cells.
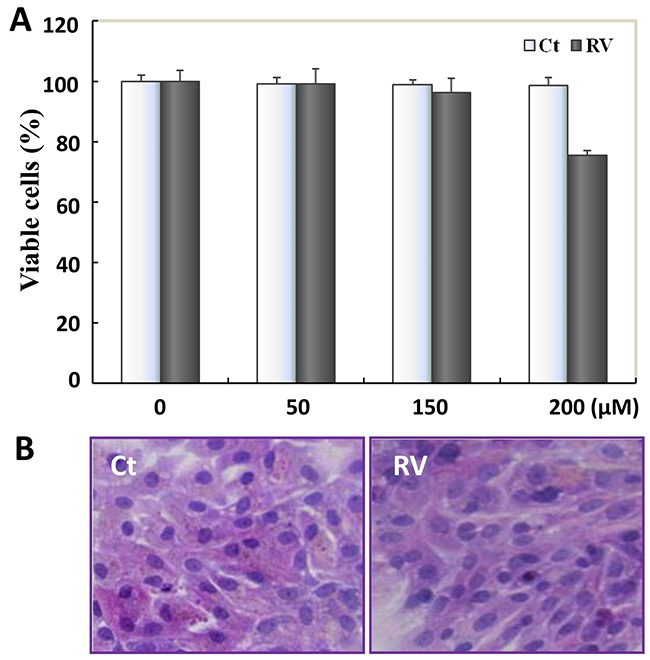
Figure 6: The safety evaluation of resveratrol to PBC cells. A. Resveratrol’s effect on the cell viability of the primary cultured normal rat bladder epithelial cells (PBC). Cells number was determined by MTT after 48h-100μM resveratrol incubation, data were expressed as means ± S.D. (n=3) (*, P<0.05). B. HE morphological staining performed on PBC cells with 100μM resveratrol treatment for 48h. PBC cells showed neither observable growth arrest nor morphological change. Ct, represented PBC cells were cultured in normal culture media; RV, represented PBC cells were treated with 100μM resveratrol treatment for 48h.
DISCUSSION
Exploring RV’s bioactive form has received more attention for “Resveratrol Paradox”, i.e., RV’s low bioavailability but high pharmacological activity [9, 18]. So far, the confirmation trial of RV’s metabolic active pattern in vitro and in vivo is still limited. Some researchers reported that piceatannol, which could be biotransformed from RV by cytochrome P450 CYP 1A1, 1A2 and 1B1 in vitro [25, 26], possessed more powerful biological activity [25, 27], but piceatannol as phase I metabolite was seldom detected in RV’s metabolites [18, 28–30]. Though the vast majority of studies have been performed using RV parent compound, some researchers proposed that RV phase II metabolites might also possess the pharmacological activity for RV’s low bioavailability [31, 32]. In the treatment of colon cancer cells, Aires et al. found RVS could inhibit colon cancer cells growth and accumulate cancer cells in S phase, but RV glucuronides (RV-3-O-glucuronide or RV-4’-O-glucuronide, Supplementary Figure 1) could not suppress cancer cells proliferation [31]. Almost at the same time, Polycarpou et al. evaluated the actions of RV and its metabolites on the growth of colon cancer cells in vitro. The results showed that RV could cause S phase arrest in all three cell lines (CCL-228, Caco-2 and HCT-116), RV 3-O-glucuronide and RV 4’-O-glucuronide caused G1 arrest in CCL-228 and Caco-2 cells, but RVS had no effect on cell cycle [32]. In addition to colon cancer, RVS also showed less bioactivity in human brain tumors (medulloblatoma UW228-3 and glioblastoma U251) and breast cancer cells (MB-MDA-231, ZR-75-1) [14, 17]. The above studies elucidated that RV parent compound could inhibit cancer cells growth, but whether RV sulfates or RV glucuronides playing the corresponding bioactivity would depend on the cancer cell types and the protocols used in the experiments.
RV showed the beneficial antitumor bioactivity in HBC cells [13, 33, 34], but how RV metabolized and whether RV or its metabolites (sulfates or glucuronides) exerted the corresponding effect on bladder carcinoma have not been reported so far. In this content, we found that HBC T24 and EJ cells showed different sensitivity to RV. Therefore, a comparison of RV metabolic patterns in RV sensitive and RV low sensitive carcinoma cells would be helpful to figure the issue out. In virtue of HPLC, LC-MS/MS and HRMS, we found RV was mainly metabolized into RVS in both HBC T24 and EJ cells (Figure 2), but it appeared that only a very small fraction of RV was metabolized to RVS after incubation with HBC cells, and the vast majority remained as the parent compound at 48h (Figure 3). Meanwhile, RVS was found as early as at 3h-time point both in T24 and EJ cells incubated with RV, but the T24 cells showed growth arrest until 24h-RV incubation and EJ cells still showed less morphological change (Figure 3, Supplementary Figure 2). According to the above results, the total amount of RV sulfate in cell lysate and supernatant didn’t increase significantly compared with RV parent form, and the the metabolism was preceded to growth inhibition, which implied that the RV metabolites might not dominate significant anticancer bioactivity in HBC T24 cells.
RV was unstable, and its cis-form (cis-RV) was found both in the cell lysates and supernatant media of T24 cells (Figure 2A, Figure 3), but cis-RV did not exert bioactivity [17], so the operation was carried out under yellow light to avoid possible photochemical side-reactions. It is also known that resveratrol is a phytoestrogen, and phenol red in the culture media might show hormonal background, so we compared the cell culture of DMEM added with/without phenol red, to measure whether phenol red has interference effect on RV metabolism in HBC cells. As shown in Supplementary Figure 3, phenol red showed no interference impact on both RV metabolism and HBC cells growth, so we chose phenol red as indicator in this experiment.
For further evaluating the bioactivity of RV metabolites in HBC cells, the RVS was prepared by RV biotransformation in vitro [17, 35, 36], and was used to treat RV-sensitive T24 cells. Compared with the 100μM RV treatment, the T24 cells incubated with RVS mixture (84.13% RVS) did not show significant apoptotic characteristics and growth inhibition, thus, it indicated that RVS possessed less pharmaceutical effect on T24 cells. RV’s pharmacological properties may result from activating or inhibiting the corresponding signaling pathways through cellular receptors [37]. It was reported that RV could inhibit the phosphorylation of Akt and decrease the expression of miR-21 in T24 and 5637 cells, thus induced HBC cells apoptosis via miR-21 regulation of the Akt/Bcl-2 signaling pathway [33]. Bai et al. also found RV could efficiently trigger HBC cells apoptosis through the modulation of Bcl-2 family proteins and activation of caspase 9 and caspase 3 followed by poly (ADP-ribose) polymerase degradation [21, 33]. In addition, the recent studies have demonstrated RV could activate SIRT1 in vitro by lowering its Michaelis constant (KM) [6, 38, 39], and more excited, Howitz et al. found RV could activate in vivo Sirt1 at very low concentration (nanomolar range) [38], which indicated RV could exert its multiple bioactivities though the low bioavailability. The above findings support our data that RV itself is more directly responsible for the antitumor activity, and RVS probably only be the metabolic form excreted from the HBC cells/tissues.
Drug metabolism is mainly classified into phase I and phase II reactions. So far, several polymorphic enzymes such as UDP-glucuronosyltransferases (UGTs), SULTs, N-acetyltransferases (NATs) are involved in the phase II metabolism of the xenobiotics in humans [40–42]. Since both T24 and EJ cells generate the same RV metabolite, and RVS is the major metabolite in HBC cells, which is catalyzed by SULTs, so the level of SULTs expression may positively or negatively influence RV’s bioavailability. On the other hand, the role of genetic polymorphism on HBC risk has been investigated by several studies [43–46]. In 1999, Vineis et al. has reported that a possible association between metabolic polymorphisms and susceptibility to cancer [47]. Furthermore, similar results were found in the environmental exposures research, Hung et al. in 2004 investigated the effects of multiple genes including NAT, GST and SULT families, which showed clearly that the polymorphisms of these families may modulate individual response to bladder carcinogens and cancer susceptibility [48]. On the basis of overall limited evidence in human carcinogenicity data, the studies carried out that different population may be due to variability in the individual susceptibility to bladder carcinogens. Epidemiological and experimental evidence favors the important effects of gender, race and age on incidence and mortality of bladder cancer [49]. Particularly, a few surveys have investigated that several polymorphic enzymes are involved in the metabolism of the bladder carcinogens in humans. SULT1A1 is one of the families of genes encoding the polymorphic enzymes which are involved in the metabolism of bladder carcinogens in humans. It has been reported that the role of SULT1A1 in both the bioactivation and detoxification of various dietary and environmental mutagens may depend on the tissue or organ [50, 51]. It has been demonstrated that SULTs have substrate-dependent effects and they exhibit marked differences in tissue distribution as well as their sensitivity to thermal inactivation and inhibitor [51]. Moreover, previous studies have shown gender-specific differences for SULT activity [52–55]. Nowell et al. found that higher platelet phenol SULT activity in women than in men [56]. Additionally, in support of this, Klasaaen et al. observed a higher SULT1A1 mRNA expression in adult male rats than adult female rats [57]. Additionally, some studies provide epidemiologic evidence of a reduced bladder cancer risk in individuals with the SULT1A1 His213 allele genotypes which have been linked with an increased risk for cancer [58, 59]. All these studies elucidated that the gene variants which contribution in the inter-individual variations of genetic susceptibility to HBC could actually affect the metabolism of the relevant exposures in HBC. In this study, RV showed different sensitivity to the HBC T24 and EJ cells, comparing the expression levels of SULT1A1 which turned out to be closely but not directly related to the metabolic activity of RV, therefore, the different RV-responses of HBC T24 and EJ cells would be a possible association between metabolic polymorphisms and individual genetic susceptibility to cancer.
Since RV parent form was more directly responsible for RV’s pharmacological activity, the safety profile of RV should be considered for further clinical application [13, 39, 60]. It was reported that RV was administered orally to male rats for 28d at a dose of 20mg/(kg X d), 1000 times the amount consumed by a 70kg person taking 1.4g of RV/d, but produced no adverse effect as assessed by growth, hematology, clinical chemistry, and histopathology [61]. In humans, a phase I study showed that ingestion of a single dose of RV (0.5g, 1g, 2.5g or 5g; 10 subjects per group) did not cause serious adverse events [61]. Recently, Anton SD and co-workers have conducted a double-blind, randomized, placebo-controlled trial to examine the safety in 32 overweight, older adults (mean age, 73±7 years). Compared with placebo, short-term (90 days) RV supplementation at doses of 300 mg/day and 1000mg/day does not adversely affect blood chemistries and is well tolerated in overweight, older individuals [62]. The above findings support the research of RV in larger clinical trials.
So far, the safety evaluation of RV on bladder has not been reported, therefore, the normal bladder epithelial cells were also treated with RV here, and the effective dosage of trans-RV (100μM) on bladder carcinoma cells showed no adverse effect on normal bladder cells. Especially, the bladder is a well-defined cavity organ in the anatomical location, so regional intravesical instillation is highly conducive to HBC therapy. At diagnosis, nearly 80% of bladder carcinomas are superficial and usually treated with drug adjuvant intravesical therapies after cystectomy to delay or prevent recurrence [63]. The good lipophilicity of RV makes it easy to be well absorbed by bladder endothelial cells via simple diffusion, and treatment with the effective doses of RV was well tolerated by the PBC cells, therefore, in view of the excellent safety of RV [61, 62, 64–66], intravesical instillation of RV would provide the potential clinical value for bladder carcinoma prevention and treatment.
CONCLUSIONS
In conclusion, HBC T24 cells showed higher sensitivity to RV than EJ cells, but both of them produced the same metabolite, RVS. RV parent form, rather than its metabolite (RVS), was primarily responsible for the anti-bladder carcinoma activity. RV’s associated metabolic enzyme SULT1A1 was upregulated after RV treatment in different sensitive HBC cells, and that indicated that neither RV’s metabolic pattern nor the related metabolic enzyme SULT1A1 was correlated with the different RV sensitivities of HBC T24 and EJ cells. In addition, RV showed almost no side-effect on the rat normal bladder epithelial cells at the therapeutic dosage, so compared to RV metabolites, trans-RV would be a potential pharmacological medicine in bladder cancer clinical prevention and therapy. In consideration of variable responses of T24 and EJ cells to RV, therefore, individual cancer types should be regarded as an important factor in medicine application. Meanwhile, in view of RV’s bioavailability was affected by different administration routes [67], the appropriate administration routes and drug delivery carriers should be focused on to improve RV’s bioavailability.
MATERIALS AND METHODS
Cell culture and treatment
HBC T24 and EJ (human bladder transitional cell carcinoma) were purchased from the Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China)/American Type Collection (ATCC, Rockville, USA). Cells were cultured in high glucose Dulbecco’s modified Eagles medium (DMEM; Invitrogen Co., Grand Island, NY, USA) which were supplemented with 10% fetal bovine serum (Gibco Life Science, Grand Island, NY, USA) and 1% penicillin-streptomycin (Gibco Invitrogen Corp., Grand Island, NY, USA) under standard conditions at 37°C in a humidified atmosphere containing 5% CO2 and 95% air. The cells were plated in 100 mm dishes (Nunc A/S, Roskilde, Denmark) at the density of 50,000/ml and incubated for 24h before further experiments. For morphologic evaluation and immunocytochemistry (ICC) staining, the coverslips were put into the dishes before initial cell seeding and collected after treatment with/without RV (Sigma Chem Co., St. Louis, MO, USA) in the experiments.
To efficiently dissolve RV, dimethyl sulfoxide (DMSO; Sigma Chem Co., St. Louis, MO, USA) was chosen as a solvent. For trans-RV was sensitive to natural light and ultraviolet light [16], a stock solution of 100mM trans-RV was wrapped in aluminum foil for protection against light and stored at -20°C. It would be diluted with culture media to the optimum working concentrations just before use. 0.2% DMSO was used to incubate with the cells as the background control, which caused no measurable effect on cell growth.
Primary urinary bladder transitional cell culture
The healthy Wistar rats were obtained from the Experimental Animals Center of Dalian Medical University, which were fed in cages under controlled conditions maintained at 22°C with a 12h light/dark period. All experimental protocols had been reviewed and approved by the ethics committee of Dalian Medical University for the protection of human subjects and experimental animals before conducting the project. The rats were sacrificed using CO2 asphyxia, and the bladders were freshly removed and treated with trypsin and ethylene diamine tetraacetic acid (EDTA), then the primary cultured normal rat bladder epithelial cells (PBC) were selectively harvested for the further experiments [68]. The cells were cultured in high glucose DMEM supplemented with 10% fetal bovine serum and 1% penicillin-streptomycin, under standard conditions at 37°C in a humidified atmosphere containing 5% CO2 and 95% air.
Sensitivity evaluation of RV
Cell viability were determined by 3-(4,5-dimethy- lthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT; Sigma-Aldrich Co., USA) assay [69]. HBC T24 and EJ cells were seeded in 96-well plates with DMEM medium supplemented 10% fetal bovine serum. After overnight incubation at 37°C, the cells were treated with various concentrations of RV (0-200μM), and 0.2% DMSO was used as background control. After incubation with RV or DMSO for 6h, 12h, 24h and 48h, respectively, the cell absorbance data were measured by a spectrophotometer (Thermo Fisher Scientific, USA) at 490nm. After being treated with 100μM RV for 48h, the cells were collected for flow cytometry (FCM) analysis [70]. Meanwhile, cell-bearing coverslips were harvested and fixed properly for hematoxylin and eosin (HE) and ICC staining. To establish the confidential conclusions, each of the experimental groups was set in triplicate and the experiments were repeated at least three times.
Sample preparation and HPLC analysis
After treatment with 100μM RV, the cell-free culture media and the cells were harvested, respectively. The cells were washed three times with PBS (phosphate buffered saline solution, pH 7.4) and lysed by sonication [71]. Then the cell lysates and their condition media were centrifuged at 10,000g for 5min, and followed by SPE [72]. In brief, the samples were loaded onto the Cleanert PEP-SPE cartridges (60mg; Agela Technol Inc. PA, USA), which were previously activated with methanol (Fisher Sci, Fair Lawn, NJ, USA) and ultrapure water purified with a Milli-Q water purification system (Millipore, Bedford, MA, USA), then the cartridges were subsequently washed with ultrapure water. The samples absorbed in the cartridge were eluted with methanol, and the eluate was dried by nitrogen spraying. At last, the residues were dissolved in 200μl of methanol for HPLC and LC-MS analysis. In all cases, sample manipulation was performed in the dark to minimize the possible photochemical isomerization of trans-RV to its cis-form [17, 73].
Altogether four kinds of samples were subjected to HPLC analysis: Sample 1, RV standards; Sample 2, RV-containing media as background control; Sample 3, the culture media incubated with T24 or EJ cells after 100μM RV treatment for 48h; Sample 4, the T24 or EJ cells treated with 100μM RV for 48h. The determination of the samples was performed on the Agilent 1200 HPLC system (Agilent Technologies, Santa Clara, CA, USA) consisted of an Agilent 1260 binary pump and 1260 dual wavelength UV-Vis detector. The detection was carried out at a wavelength 303nm and the column oven was set at 30°C [64]. Chromatographic separation of the samples was performed on a Cosmosil C18-AR-II column (5μm, 4.6mm×250mm; Nacalai Tesque, Japan) preceded by a C18 guard column (5μm, 4.6mm×10mm), with a mobile phase consisted of 5mM ammonium acetate (mobile phase A, Alfa Aesar, A Johnson Matthey Company, Ward Hill, MA, USA) and methanol (mobile phase B) at a flow rate of 1ml/min. The mobile phases were degassed by sonication for 15min at room temperature before use. A gradient elution was carried out as follows: 0min, 45% B; 25min, 60% B; 30min, 45% B; 40min, 45% B. Subsequently, equilibrate the column for 10min before the next injection. Samples were filtered with a 0.45μm filter membrane (Millipore, Bedford, MA, USA) and a 10μl aliquot was injected.
For quantitative analysis, cells were scraped off, washed three times with PBS (pH 7.4), and lysed with 416μl PBS and 84μl IS (1, 8-dihydroxyanthraquinone, 200μg/ml) by sonication. Cells cultured for 48h in medium with the same working concentration of DMSO (0.2%) were used as a background control. The collected culture medium and cell lysates were centrifuged at 12, 000 rpm/min for 10 min at 4°C. The supernatant was collected, and then purified with SPE. The eluates were evaporated to a final volume of 400μl for HPLC analysis. Chromatographic condition: The analyses were performed on the HITACHI Chromaster 5000 HPLC system (Hitachi High-Technologies Corporation, Tokyo, Japan) consisted of a HITACHI 5110 pump, 5210 auto sampler and 5430 diode array detector. The detection was carried out at a wavelength 303nm and 5310 column oven was set at 30°C. All the separation of the samples was carried out on a Cosmosil C18-AR-II column (5μm, 4.6mm×250mm; Nacalai Tesque, Japan) with a mobile phase consisted of 20% acetonitrile (mobile phase A, acetic acid adjusted pH 3.5) and 80% acetonitrile (mobile phase B, acetic acid adjusted pH 3.5) at a flow rate of 1ml/min. The mobile phase consisted of two phases, phases A was 20% acetonitrile and phase B was 80% acetonitrile (acetic acid adjusted pH 3.5). The gradient elution mode was carried out as follows: 0-14 min, linear gradient from A: B (0: 100, v/v) to A: B (60: 40, v/v); 14-20 min, the liner gradient from A: B (60:40, v/v) to A: B (0: 100, v/v), the mobile phase was hold on A: B (0: 100, v/v). Each run was followed by equilibration time of 15min before the next injection [67].
Identification of RV metabolite(s) by LC-MS/MS and HRMS
To further identify the metabolite(s) of RV in T24 cells, the extracted samples were analyzed by direct online LC-MS/MS under the chromatographic series (Agilent Technol Inc., Santa Clara, CA, USA) coupled to an Applied Biosystems API 3200 QTrap tandem mass spectrometer (Applied Biosystem/MDS SCIEX, Foster City, CA, USA). The MS and MS/MS data were obtained by the Applied Biosystem/MDS SCIEX analyst software (Version 1.4.1). A Cosmosil C18-AR-II column (5μm, 4.6mm×250mm; Nacalai Tesque, Japan) with a guard column was used for chromatographic separation. 5mM ammonium acetate was used as solvent A, and methanol as solvent B with the following gradient at a flow rate of 500μl/min: 45-60% B linear (0-25min), 60-45% B linear (25-30min), 45% B linear (30-40min).
The MS determination of the metabolites was operated in a negative ion mode. To obtain maximum sensitivity, the ion spray interface and the mass spectrometric parameters were optimized before use. Full-scan data acquisition was performed by scanning over the range of m/z 100-600 in profile mode, using a cycle time of 2s and a pause between scans of 2ms [17]. The identification of the samples was based on their retention time and ion fragmentations in the MS and MS/MS mode.
For further confirmation of the metabolites, HRMS analysis was performed on the LC-ESI-IT-TOF-MS (Shimadzu Co., Kyoto, Japan) in negative ion mode at a resolution of 10,000 FWHM. Before the sample was injected onto a Shim-pack VP-ODS column (5μm, 2.0×150 mm; Shimadzu Co., Kyoto, Japan), the accurate masses were corrected by using the standard sample sodium trifluoroacetate. The column temperature was kept at 40°C, and the flow rate was 0.6ml/min. The mobile phase consisted of two phases: phase A was acetonitrile/10mM acetic acid water solution (95:5, v/v, pH 3.0), and phase B was 10mM acetic acid water solution (pH 3.0). A gradient elution was carried out as the following proportions (v/v) of phase A and B: 0min (95/5), 5min (70/30), 5.5min (40/60), 12.5(40/60), 12.6min (95/5). The column was equilibrated with 5% phase B for 5min before the next run. MS data were processed with LC-MS solution ver. 3.4 software (Shimadzu, Japan).
Bladder-associated metabolic enzyme determination by ICC and western blots
The main metabolite of RV in HBC T24 and EJ cells was RVS, since sulfation was regulated by phase II metabolic enzyme sulfotransferases (SULTs), and SULT1A1 took part in phenol compounds metabolism [23, 24], thus evaluating the potential influence of RV to SULT1A1 appeared to be essential. To evaluate RV’s related metabolic enzyme SULT1A1 in T24 and EJ cells, ICC staining and Western blots were performed. For ICC staining, cells on coverslips were collected from different experimental groups treated with/without RV. The first antibody of rabbit anti-human SULT1A1 (Protein Tech Group, Inc., Chicago, USA) was used in the dilution rates of 1:120. Other solutions were prepared by ICC staining kit (HistotainTM-plus Kits SP-9000, Zymed, USA).
For Western blots analysis, total cellular proteins were prepared from the treated cells. 50μg of the sample proteins were separated by electrophoresis in 10% sodium dodecylsulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to polyvinylidene difluoride membrane (Amersham Biosci, Buckinghamshire, UK). The membrane was blocked with 5% skimmed milk in TBS-T (10mM Tris-Cl, PH 8.0, 150mM NaCl and 0.5% Tween 20) at 37°C for 2h, followed by incubation with the first antibodies in the appropriate concentrations (SULT1A1, 1:1000; β-actin, 1:3000, Protein Tech Group, Inc., Chicago, USA) at 4°C overnight, and then incubated with HRP-conjugated anti-rabbit IgG (Zymed Lab Inc., San Francisco, CA, USA) at 37°C for 1.5h. The immunolabeling was detected using the enhanced chemiluminescence system (Roche Diagnostics GmbH, Mannheim, Germany) and visualized using the UVP Bio-spectrum Imaging System (UVP, Inc, Upland, CA, USA). β-Actin was used as an internal quantitative control in densitometry analysis. After removing the labeling signal by incubation with stripping buffer (62.5mM Tris-Cl, pH 6.7, 100mM 2-mercaptoethanol, 2% SDS) at 55°C for 30min, the membrane was re-probed with β-actin by the same experimental procedures until all of the parameters were examined.
RT-PCR and real-time PCR
For semi-quantitative determination of SULT1A1 mRNA expression, total cellular RNA was extracted from cells with Trizol reagent (Life Technologies, Grand Island, NY, USA). Reverse transcription was performed on RNA samples, and this was followed by PCR for SULT1A1 and β-actin, according to the manufacturer’s recommendations (Takara, Dalian Branch, Dalian, China). The sequences of PCR primers were as follows: SULT1A1, Forward: 5’-GCAACGCAAAGGATGTGGCA-3’, Reverse: 5’-TCC GTAGGACACTTCTCCGA-3’. β-actin, Forward: 5’-GC ATGGAGTCCTGTGGCAT-3’, Reverse: 5’-CATGAAG CATTTGCGGTGG-3’ [17]. The PCR products were resolved on 1% agarose gel containing ethidium bromide (0.5μg/ml), and photographed with the UVP Bio-spectrum Imaging System. The PCR products generated from the same reverse transcription solutions by a pair of β-actin primers were used as internal quantitative controls.
For quantitative real-time PCR, RNA samples (1μg) were reversely transcribed in a final volume of 20μl containing Prime Script RT reagents (Takara, Dalian Branch, Dalian, China). Reaction mixtures were then incubated at 37°C for 15min and 85°C for 5s, and kept at 4°C. The primers were as follows: SULT1A1, Forward: 5’-GCAACGCAAAGGATGTGGC-3’, Reverse: 5’-TC CCTTTTCGGGTTCTCCTTC-3’. GAPDH, Forward: 5’-GAAGGTCGGAGTCAACGGAT-3’, Reverse: 5’-CC TGGAAGATGGTGATGGG-3’. 25μl reaction mixtures were prepared by adding 2×SYBR Premix Ex TaqTM II (Takara, Dalian Branch, Dalian, China), 10μM forward and reverse primers (Takara, Dalian Branch, Dalian, China), 2μl of cDNA template, and a suitable amount of distilled H2O. Amplification and detection were performed with the Thermal Cycler Dice Real Time System (TaKaRa; Code TP800). Each reaction was performed in triplicate, and ‘no-template’ controls were included in each experiment.
RV biotransformation and anticancer evaluation
For sulfonating RV, the homogenate of rat livers with ice-buffer (250mM sucrose, 10mM HEPES, 3mM 2-mercaptoethanol, pH7.4) was centrifuged at 10,000×g for 20min in 4°C, followed by supernatant was centrifuged at 100,000×g for 60min in 4°C to get cytosolic proteins which stored in -80°C until use (less than 6 months) [35]. Subsequently, a mixture (200μl) containing 100μl cytosolic proteins isolated from rat livers, 5mM RV, 2mM 3’-phosphoadenosine 5’-phosphosulfate (PAPS, Sigma Chem Co., St. Louis, MO, USA), 1mM dithiothreitol (DTT) and 20mM Mops buffer was prepared and incubated at 37°C for 2h. The reaction was terminated by standing the mixture-containing thin-wall tube in the boiling water for 5min. After the suspension was centrifuged at 10,000×g for 5min, 10μl of the supernatant was subjected to HPLC and LC/MS analysis for separating RV and its biotransformation metabolite(s), and the aliquots of the remaining part was used to treat T24 cells in the total concentration of 100μM. The cells treated with the chemical solution for liver lysate preparation, the liver lysate alone and the combination of RV were used as background controls, respectively. The cellular response(s) were checked with the parameters mentioned above.
Statistical analyses
Data were given as the mean ± standard deviation, and statistical analyses were performed using SPSS 13.0 and GraphPad Prism 5. MTT data were analyzed with one-way ANOVA. It was considered statistically significant if the p-value is less than 0.05.
ACKNOWLEDGMENTS
We thank Drs. Ming-Zhe Gao and Peng Gao in the Laboratory of High Resolution Separation/Analysis and Metabonomics of Dalian institute of Chemical Physics, Chinese Academy of Sciences, for their assistance with the HPLC, LC-MS/MS and HRMS analysis.
CONFLICTS OF INTEREST
The authors declare no conflict of interest.
GRANT SUPPORT
The work was supported by the grants from National Natural Science Foundation of China (No. 81072063, 81450016 and 81672945), and the Science Project of Liaoning Province (No. 201602234). In part by an oversea fellowship from China Scholarship Council (No. 201408210085).
REFERENCES
1. Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011; 61: 69-90. doi: 10.3322/caac.20107.
2. Baur JA, Pearson KJ, Price NL, Jamieson HA, Lerin C, Kalra A, Prabhu VV, Allard JS, Lopez-Lluch G, Lewis K, Pistell PJ, Poosala S, Becker KG, et al. Resveratrol improves health and survival of mice on a high-calorie diet. Nature. 2006; 444: 337-42. doi: 10.1038/nature05354.
3. Jang M, Cai L, Udeani GO, Slowing KV, Thomas CF, Beecher CW, Fong HH, Farnsworth NR, Kinghorn AD, Mehta RG, Moon RC, Pezzuto JM. Cancer chemopreventive activity of resveratrol, a natural product derived from grapes. Science. 1997; 275: 218-20. doi: 10.1126/science.275.5297.218.
4. Milne JC, Lambert PD, Schenk S, Carney DP, Smith JJ, Gagne DJ, Jin L, Boss O, Perni RB, Vu CB, Bemis JE, Xie R, Disch JS, et al. Small molecule activators of SIRT1 as therapeutics for the treatment of type 2 diabetes. Nature. 2007; 450: 712-16. doi: 10.1038/nature06261.
5. Francioso A, Mastromarino P, Masci A, d’Erme, Mosca L. Chemistry, stability and bioavailability of resveratrol. Med Chem. 2014; 10: 237-45. doi: 10.2174/15734064113096660053.
6. Gambino J, Ingles M, Olaso G, Lopez-Grueso R, Bonet-Costa V, Gimeno-Mallench L, Mas-Bargues C, Abdelaziz KM, Gomez-Cabrera MC, Vina J, Borras C. Properties of resveratrol: in vitro and in vivo studies about metabolism, bioavailability, and biological effects in animal models and humans. Oxid Med Cell Longev. 2015; 2015: 837042. doi: org/10.1155/2015/837042.
7. Brown VA, Patel KR, Viskaduraki M, Crowell JA, Perloff M, Booth TD, Vasilinin G, Sen A, Schinas AM, Piccirilli G, Brown K, Steward WP, Gescher AJ, et al. Repeat dose study of the cancer chemopreventive agent resveratrol in healthy volunteers: safety, pharmacokinetics, and effect on the insulin-like growth factor axis. Cancer Res. 2010; 70: 9003-11. doi: 10.1158/0008-5472.
8. Stefanska B, Karlic H, Varga F, Fabianowska-Majewska K, Haslberger A. Epigenetic mechanisms in anti-cancer actions of bioactive food components--the implications in cancer prevention. Br J Pharmacol. 2012; 167: 279-97. doi: 10.1111/j.1476-5381.2012.02002.x.
9. Zhou R, Fukui M, Choi HJ, Zhu BT. Induction of a reversible, non-cytotoxic S-phase delay by resveratrol: implications for a mechanism of lifespan prolongation and cancer protection. Br J Pharmacol. 2009; 158: 462-74. doi: 10.1111/j.1476-5381.2009.00268.x.
10. Gescher A, Steward WP, Brown K. Resveratrol in the management of human cancer: how strong is the clinical evidence? Ann N Y Acad Sci. 2013; 1290: 12-20. doi: 10.1111/nyas.12205.
11. Dhar S, Kumar A, Rimando AM, Zhang X, Levenson AS. Resveratrol and pterostilbene epigenetically restore PTEN expression by targeting oncomiRs of the miR-17 family in prostate cancer. Oncotarget. 2015; 6: 27214-26. doi: 10.18632/oncotarget.4877.
12. Xue S, Xiao-Hong S, Lin S, Jie B, Li-Li W, Jia-Yao G, Shun S, Pei-Nan L, Mo-Li W, Qian W, Xiao-Yan C, Qing-You K, Peng Z, et al. Lumbar puncture-administered resveratrol inhibits STAT3 activation, enhancing autophagy and apoptosis in orthotopic rat glioblastomas. Oncotarget. 2016; 7: 75790-9. doi: 10.18632/oncotarget.12414.
13. Wu ML, Li H, Yu LJ, Chen XY, Kong QY, Song X, Shu XH, Liu J. Short-term resveratrol exposure causes in vitro and in vivo growth inhibition and apoptosis of bladder cancer cells. PLoS One. 2014; 9: e89806. doi: 10.1371/journal.pone.0089806.
14. Miksits M, Wlcek K, Svoboda M, Kunert O, Haslinger E, Thalhammer T, Szekeres T, Jäger W. Antitumor activity of resveratrol and its sulfated metabolites against human breast cancer cells. Planta Med. 2009; 75: 1227-30. doi: 10.1055/s-0029-1185533.
15. Patel KR, Andreadi C, Britton RG, Horner-Glister E, Karmokar A, Sale S, Brown VA, Brenner DE, Singh R, Steward WP, Gescher AJ, Brown K. Sulfate metabolites provide an intracellular pool for resveratrol generation and induce autophagy with senescence. Sci Transl Med. 2013; 5: 205ra133. doi: 10.1126/scitranslmed.3005870.
16. Patel KR, Brown VA, Jones DJ, Britton RG, Hemingway D, Miller AS, West KP, Booth TD, Perloff M, Crowell JA, Brenner DE, Steward WP, Gescher AJ, et al. Clinical pharmacology of resveratrol and its metabolites in colorectal cancer patients. Cancer Res. 2010; 70: 7392-99. doi: 10.1158/0008-5472.
17. Shu XH, Li H, Sun Z, Wu ML, Ma JX, Wang JM, Sun Y, Fu YS, Chen XY, Kong QY, Liu J. Identification of metabolic pattern and bioactive form of resveratrol in human medulloblastoma cells. Biochem Pharmacol. 2010; 79: 1516-25. doi: 10.1016/j.bcp.2010.01.022.
18. Walle T. Bioavailability of resveratrol. Ann N Y Acad Sci. 2011; 1215: 9-15. doi: 10.1111/j.1749-6632.2010.05842x.
19. Shu XH, Li H, Sun XX, Wang Q, Sun Z, Wu ML, Chen XY, Li C, Kong QY, Liu J. Metabolic patterns and biotransformation activities of resveratrol in human glioblastoma cells: relevance with therapeutic efficacies. PLoS One. 2011; 6: e27484. doi: 10.1371/journal.pone.0027484.
20. Sun Z, Shi S, Li H, Shu XH, Chen XY, Kong QY, Liu J. Evaluation of resveratrol sensitivities and metabolic patterns in human and rat glioblastoma cells. Cancer Chemother Pharmacol. 2013; 72: 965-73. doi: 10.1007/s00280-013-2274-y.
21. Lin X, Wu G, Huo WQ, Zhang Y, Jin FS. Resveratrol induces apoptosis associated with mitochondrial dysfunction in bladder carcinoma cells. Int J Urol. 2012; 19: 757-64. doi: 10.1111/j.1442-2042.2012.03024.x.
22. Wang D, Hang T, Wu C, Liu W. Identification of the major metabolites of resveratrol in rat urine by HPLC-MS/MS. J Chromatogr B Analyt Technol Biomed Life Sci. 2005; 829: 97-106. doi: 10.1016/j.jchromb.2005.09.040.
23. Miksits M, Wlcek K, Svoboda M, Thalhammer T, Ellinger I, Stefanzl G, Falany CN, Szekeres T, Jaeger W. Expression of sulfotransferases and sulfatases in human breast cancer: impact on resveratrol metabolism. Cancer Lett. 2010; 289: 237-45. doi: 10.1016/j.canlet.2009.08.020.
24. Murias M, Miksits M, Aust S, Spatzenegger M, Thalhammer T, Szekeres T, Jaeqer W. Metabolism of resveratrol in breast cancer cell lines: impact of sulfotransferase 1A1 expression on cell growth inhibition. Cancer Lett. 2008; 261: 172-82. doi: 10.1016/j.canlet.2007.11.008.
25. Piver B, Fer M, Vitrac X, Merillon JM, Dreano Y, Berthou F, Lucas D. Involvement of cytochrome P450 1A2 in the biotransformation of trans-resveratrol in human liver microsomes. Biochem Pharmacol. 2004; 68: 773-82. doi: 10.1016/j.bcp.2004.05.008.
26. Potter GA, Patterson LH, Wanogho E, Perry PJ, Butler PC, Ijaz T, Ruparelia KC, Lamb JH, Farmer PB, Stanley LA, Burke MD. The cancer preventative agent resveratrol is converted to the anticancer agent piceatannol by the cytochrome P450 enzyme CYP1B1. Br J Cancer. 2002; 86: 774-8. doi: 10.1038/sj.bjc.6600197.
27. Piotrowska H, Kucinska M, Murias M. Biological activity of piceatannol: leaving the shadow of resveratrol. Mutat Res. 2012; 750: 60-82. doi: 10.1016/j.mrrev.2011.11.001.
28. Tome-Carneiro J, Larrosa M, Gonzalez-Sarrias A, Tomas-Barberan FA, Garcia-Conesa MT, Espin JC. Resveratrol and clinical trials: the crossroad from in vitro studies to human evidence. Curr Pharm Des. 2013; 19: 6064-93. doi: 10.2174/13816128113199990407.
29. Wenzel E, Somoza V. Metabolism and bioavailability of trans-resveratrol. Mol Nutr Food Res. 2005; 49: 472-81. doi: 10.1002/mnfr.200500010.
30. Kroon PA, Iyer A, Chunduri P, Chan V, Brown L. The cardiovascular nutrapharmacology of resveratrol: pharmacokinetics, molecular mechanisms and therapeutic potential. Curr Med Chem. 2010; 17: 2442-55. doi: 10.2174/092986710791556032.
31. Aires V, Limagne E, Cotte AK, Latruffe N, Ghiringhelli F, Delmas D. Resveratrol metabolites inhibit human metastatic colon cancer cells progression and synergize with chemotherapeutic drugs to induce cell death. Mol Nutr Food Res. 2013; 57: 1170-81. doi: 10.1002/mnfr.201200766.
32. Polycarpou E, Meira LB, Carrington S, Tyrrell E, Modjtahedi H, Carew MA. Resveratrol 3-O-D-glucuronide and resveratrol 4’-O-D-glucuronide inhibit colon cancer cell growth: evidence for a role of A3 adenosine receptors, cyclin D1 depletion, and G1 cell cycle arrest. Mol Nutr Food Res. 2013; 57: 1708-17. doi: 10.1002/mnfr.201200742.
33. Bai Y, Mao QQ, Qin J, Zheng XY, Wang YB, Yang K, Shen HF, Xie LP. Resveratrol induces apoptosis and cell cycle arrest of human T24 bladder cancer cells in vitro and inhibits tumor growth in vivo. Cancer Sci. 2010; 101: 488-93. doi: 10.1111/j.1349-7006.2009.01415.x.
34. Stocco B, Toledo K, Salvador M, Paulo M, Koyama N, Torqueti Toloi MR. Dose-dependent effect of resveratrol on bladder cancer cells: chemoprevention and oxidative stress. Maturitas. 2012; 72: 72-8. doi: 10.1016/j.maturitas.2012.02.004.
35. Purchartova K, Engels L, Marhol P, Sulc M, Kuzma M, Slamova K, Elling L, Křen V. Enzymatic preparation of silybin phase II metabolites: sulfation using aryl sulfotransferase from rat liver. Appl Microbiol Biotechnol. 2013; 97: 10391-8. doi: 10.1007/s00253-013-4794-0.
36. Sharan S, Iwuchukwu OF, Canney DJ, Zimmerman CL, Nagar S. In vivo-formed versus preformed metabolite kinetics of trans-resveratrol-3-sulfate and trans-resveratrol-3-glucuronide. Drug Metab Dispos. 2012; 40: 1993-2001. doi: 10.1124/dmd.112.046417.
37. Varoni EM, Lo Faro AF, Sharifi-Rad J, Iriti M. Anticancer molecular mechanisms of resveratrol. Front Nutr. 2016; 3: 8. doi: 10.3389/fnut.2016.00008.
38. Howitz KT, Bitterman KJ, Cohen HY, Lamming DW, Lavu S, Wood JG, Zipkin RE, Chung P, Kisielewski A, Zhang LL, Scherer B, Sinclair DA. Small molecule activators of sirtuins extend Saccharomyces cerevisiae lifespan. Nature. 2003; 425: 191-6. doi: 10.1038/nature01960.
39. Sinclair DA, Guarente L. Small-molecule allosteric activators of sirtuins. Annu Rev Pharmacol Toxicol. 2014; 54: 363-80. doi: 10.1146/annurev-pharmtox-010611-134657.
40. Allali-Hassani A, Pan PW, Dombrovski L, Najmanovich R, Tempel W, Dong A, Loppnau P, Martin F, Thornton J, Edwards AM, Bochkarev A, Plotnikov AN, Vedadi M, et al. Structural and chemical profiling of the human cytosolic sulfotransferases. PLoS Biol. 2007; 5: e97. doi: 10.1371/journal.pbio.0050097.
41. Lindsay J, Wang LL, Li Y, Zhou SF. Structure, function and polymorphism of human cytosolic sulfotransferases. Curr Drug Metab. 2008; 9: 99-105. doi: 10.2174/138920008783571819.
42. Jancova P, Anzenbacher P, Anzenbacherova E. Phase II drug metabolizing enzymes. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub. 2010; 154: 103-16. doi: 10.5507/bp.2010.017.
43. Strange RC, Fryer AA. The glutathione S-transferases: influence of polymorphism on cancer susceptibility. IARC Sci Publ. 1999; 148: 231-49.
44. Kellen E, Hemelt M, Broberg K, Golka K, Kristensen VN, Hung RJ, Matullo G, Mittal RD, Porru S, Povey A, Schulz WA, Shen J, Buntinx F, et al. Pooled analysis and meta-analysis of the glutathione S-transferase P1 Ile 105Val polymorphism and bladder cancer: a HuGE-GSEC review. Am J Epidemiol. 2007; 165: 1221-30. doi: 10.1093/aje/kwm003.
45. Engel LS, Taioli E, Pfeiffer R, Garcia-Closas M, Marcus PM, Lan Q, Boffetta P, Vineis P, Autrup H, Bell DA, Branch RA, Brockmoller J, Daly AK, et al. Pooled analysis and meta-analysis of glutathione S-transferase M1 and bladder cancer: a HuGE review. Am J Epidemiol. 2002; 156: 95-109. doi: 10.1093/aje/kwf018.
46. Sanderson S, Salanti G, Higgins J. Joint effects of the N-acetyltransferase 1 and 2 (NAT1 and NAT2) genes and smoking on bladder carcinogenesis: a literature-based systematic HuGE review and evidence synthesis. Am J Epidemiol. 2007; 166: 741-51. doi: 10.1093/aje/kwm167.
47. Vineis P, Malats N. Strategic issues in the design and interpretation of studies on metabolic polymorphisms and cancer. IARC Sci Publ. 1999; 51-61.
48. Hung RJ, Boffetta P, Brennan P, Malaveille C, Hautefeuille A, Donato F, Gelatti U, Spaliviero M, Placidi D, Carta A, Carlo AS, Porru S. GST, NAT, SULT1A1, CYP1B1 genetic polymorphisms, interactions with environmental exposures and bladder cancer risk in a high-risk population. Int J Cancer. 2004; 110: 598-604. doi: 10.1002/ijc.20157.
49. Madeb R, Messing EM. Gender, racial and age differences in bladder cancer incidence and mortality. Urol Oncol. 2004; 22: 86-92. doi: 10.1016/S1078-1439(03)00139-X.
50. Glatt H, Engelke CE, Pabel U, Teubner W, Jones AL, Coughtrie MW, Andrae U, Falany CN, Meinl W. Sulfotransferases: genetics and role in toxicology. Toxicol Lett. 2000; 112-113: 341-8. doi: 10.1016/S0378-4274(99)00214-3.
51. Glatt H, Boeing H, Engelke CE, Ma L, Kuhlow A, Pabel U, Pomplun D, Teubner W, Meinl W. Human cytosolic sulphotransferases: genetics, characteristics, toxicological aspects. Mutat Res. 2001; 482: 27-40. doi: 10.1016/S0027-5107(01)00207-X.
52. Brittelli A, De Santi C, Raunio H, Pelkonen O, Rossi G, Pacifici GM. Interethnic and interindividual variabilities of platelet sulfotransferases activity in Italians and Finns. Eur J Clin Pharmacol. 1999; 55: 691-5. doi: 10.1007/s002280050695.
53. Pacifici GM, Gucci A, Giuliani L. Testosterone sulphation and glucuronidation in the human liver: interindividual variability. Eur J Drug Metab Pharmacokinet. 1997; 22: 253-8. doi: 10.1007/BF03189815.
54. Pacifici GM, D’Alessandro C, Gucci A, Giuliani L. Sulphation of the heterocyclic amine 1,2,3,4-tetrahydroisoquinoline in the human liver and intestinal mucosa: interindividual variability. Arch Toxicol. 1997; 71: 477-81. doi: 10.1007/s002040050415.
55. Runge-Morris MA. Sulfotransferase gene expression in rat hepatic and extrahepatic tissues. Chem Biol Interact. 1994; 92: 67-76. doi: 10.1016/0009-2797(94)90054-X.
56. Nowell S, Ambrosone CB, Ozawa S, MacLeod SL, Mrackova G, Williams S, Plaxo J, Kadlubar FF, Lang NP. Relationship of phenol sulfotransferase activity (SULT1A1) genotype to sulfotransferase phenotype in platelet cytosol. Pharmacogenetics. 2000; 10: 789-97. doi: 10.1097/00008571-200012000-00004.
57. Klaassen CD, Liu L, Dunn RT 2nd. Regulation of sulfotransferase mRNA expression in male and female rats of various ages. Chem Biol Interact. 1998; 109: 299-313. doi: 10.1016/S0009-2797(97)00141-5.
58. Zheng L, Wang Y, Schabath MB, Grossman HB, Wu X. Sulfotransferase 1A1 (SULT1A1) polymorphism and bladder cancer risk: a case-control study. Cancer Lett. 2003; 202: 61-9.
59. Bamber DE, Fryer AA, Strange RC, Elder JB, Deakin M, Rajagopal R, Fawole A, Giliseen RA, Campbell FC, Coughtrie MW. Phenol sulphotransferase SULT1A1*1 genotype is associated with reduced risk of colorectal cancer. Pharmacogenetics. 2001; 11: 679-85. doi: 10.1097/00008571-200111000-00006.
60. Szekeres T, Fritzer-Szekeres M, Saiko P, Jager W. Resveratrol and resveratrol analogues--structure-activity relationship. Pharm Res. 2010; 27: 1042-48. doi: 10.1007/s11095-010-0090-1.
61. Juan ME, Vinardell MP, Planas JM. The daily oral administration of high doses of trans-resveratrol to rats for 28 days is not harmful. J Nutr. 2002; 132: 257-60. doi: 10.1016/j.jfranklin.2009.03.004.
62. Anton SD, Embry C, Marsiske M, Lu X, Doss H, Leeuwenburgh C, Manini TM. Safety and metabolic outcomes of resveratrol supplementation in older adults: results of a twelve-week, placebo-controlled pilot study. Exp Gerontol. 2014; 57: 181-7. doi: 10.1016/j.exger.2014.05.015.
63. Lotan Y, Amiel G, Boorjian SA, Clark PE, Droller M, Gingrich JR, Guzzo TJ, Inman BA, Kamat AM, Karsh L, Nielsen ME, Smith ND, Shariat SF, et al. Comprehensive handbook for developing a bladder cancer cystectomy database. Urol Oncol. 2013; 31: 812-26. doi: 10.1016/j.urolonc.2011.09.004.
64. Qian G, Leung SY, Lu G, Leung KS. Optimization and validation of a chromatographic method for the simultaneous quantification of six bioactive compounds in Rhizoma et Radix Polygoni Cuspidati. J Pharm Pharmacol. 2008; 60: 107-13. doi: 10.1211/jpp.60.1.0014.
65. Boocock DJ, Faust GE, Patel KR, Schinas AM, Brown VA, Ducharme MP, Booth TD, Crowell JA, Perloff M, Gescher AJ, Steward WP, Brenner DE. Phase I dose escalation pharmacokinetic study in healthy volunteers of resveratrol, a potential cancer chemopreventive agent. Cancer Epidemiol Biomarkers Prev. 2007; 16: 1246-52. doi: 10.1158/1055-9965.EPI-07-0022.
66. Patel KR, Scott E, Brown VA, Gescher AJ, Steward WP, Brown K. Clinical trials of resveratrol. Ann N Y Acad Sci. 2011; 1215: 161-9. doi: 10.1111/j.1749-6632.2010.05853.x.
67. Shu XH, Wang LL, Li H, Song X, Shi S, Gu JY, Wu ML, Chen XY, Kong QY, Liu J. Diffusion efficiency and bioavailability of resveratrol administered to rat brain by different routes: therapeutic implications. Neurotherapeutics. 2015; 12: 491-501. doi: 10.1007/s13311-014-0334-6.
68. Hashimoto Y, Kitagawa HS. In vitro neoplastic transformation of epithelial cells of rat urinary bladder by nitrosamines. Nature. 1974; 252: 497-9. doi: 10.1038/252497a0.
69. Ko JC, Su YJ, Lin ST, Jhan JY, Ciou SC, Cheng CM, Lin YW. Suppression of ERCC1 and Rad51 expression through ERK1/2 inactivation is essential in emodin-mediated cytotoxicity in human non-small cell lung cancer cells. Biochem Pharmacol. 2010; 79: 655-64. doi: 10.1016/j.bcp.2009.09.024.
70. Pozo-Guisado E, Alvarez-Barrientos A, Mulero-Navarro S, Santiago-Josefat B, Fernandez-Salguero PM. The antiproliferative activity of resveratrol results in apoptosis in MCF-7 but not in MDA-MB-231 human breast cancer cells: cell-specific alteration of the cell cycle. Biochem Pharmacol. 2002; 64: 1375-86. doi: 10.1016/s0006-2952(02)01296-0.
71. Wang D, Xu Y, Liu W. Tissue distribution and excretion of resveratrol in rat after oral administration of Polygonum cuspidatum extract (PCE). Phytomedicine. 2008; 15: 859-66. doi: 10.1016/j.phymed.2008.02.009.
72. Mercolini L, Addolorata Saracino M, Bugamelli F, Ferranti A, Malaguti M, Hrelia S, Raqqi MA. HPLC-F analysis of melatonin and resveratrol isomers in wine using an SPE procedure. J Sep Sci. 2008; 31: 1007-14. doi: 10.1002/jssc.200700458.
73. Vian MA, Tomao V, Gallet S, Coulomb PO, Lacombe JM. Simple and rapid method for cis- and trans-resveratrol and piceid isomers determination in wine by high-performance liquid chromatography using chromolith columns. J Chromatogr A. 2005; 1085: 224-9. doi: 10.1016/j.chroma.2005.05.083.

