INTRODUCTION
Cancer has become one of the most important diseases threatening human health and life [1–3]. Exploring early diagnosis and treatment are critical for the research and clinical treatment of cancers [4, 5]. Recently, researchers focus on the molecular mechanism and new tumor biomarkers associated with tumor screening, diagnosis, prognosis, and evaluation of treatment efficacy [6–8]. However, the exact mechanism of cancers is still unknown. Therefore, to identify sensitive and specific biomarkers for prognosis of patients with cancers is urgently needed.
Long non-coding RNAs (lncRNAs) are a class of non-coding transcripts longer than 200 nucleotides [9, 10]. By comparing their expression of tumors and normal cells, lncRNAs are abnormally expressed in the various tumors, functioning as oncogenes or tumor suppressors [11–14].
Maternally expressed gene 3 (MEG3) is the first lncRNA to be found to have tumor suppressor function, which is expressed in many human normal tissues [15]. The lost expression of MEG3 has been found in many human tumors, such as bladder cancer [16], cervical carcinoma [17], hepatocellular cancers [15], and meningiomas [18]. In addition, hypermethylation of promoter or intergenic differentially methylated region (DMRs) upstream of MEG3 gene has been found to exert a vital role in the silence of MEG3 expression in tumors [19]. Moreover, MEG3 could inhibit cell proliferation in non-small lung cancer by inducing the expression of P53 [20, 21]. Together, lncRNA MEG3 may not only act as a potential therapeutic target, but also as a novel prognostic biomarker in cancer. However, no meta-analysis was been conducted assess the association between MEG3 and the survival of patients with cancers. Therefore, this meta-analysis evaluated the value of the MEG3 with tumor metastasis, progression, and survival.
RESULTS
Study characteristics
As shown in Figure 1, we searched 321 articles in the databases. After screening the titles and abstracts, 28 full-text articles were assessed for eligibility. Then because of no usable data or incomplete data, 19 papers were excluded. As a result, a total of 9 articles were in the current meta-analysis [22–30]. Eight different types of cancer were evaluated in this meta-analysis, with 1 non-small cell lung cancer (NSCLC), 2 gastric cancer (GC), 1 tongue squamous cell carcinoma (TSCC), 1 non-functioning pituitary adenomas (NFPAs), hepatocellular carcinoma (HCC), 1 osteosarcoma, 1 prostate cancer (PC), 1 bladder cancer (BC). In these included studies, the level of MEG3 expression was determined in collected tumor tissues.
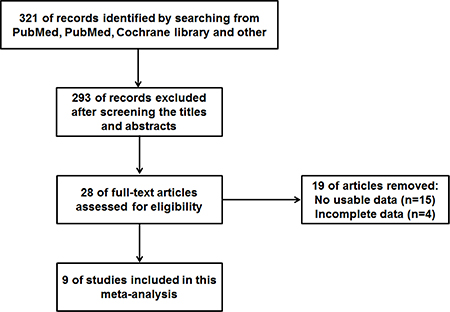
Figure 1: The flow diagram of this meta-analysis.
Table 1 summarized the main characteristics of the included 9 studies ranging 2013 to 2016. 9 studies enrolling 536 participants, with a maximum sample size of 80 and a minimum sample size of 21 patients. Because the cut-off definitions were various, the cut-off values were different in these studies. Not all studies examined both OS and RFS, because most of the studies were retrospective cohort studies; 6 studies investigated the association between MEG3 and OS [22–26, 30], while 2 studies assessed the association between CCAT2 and RFS [25, 29]. Meanwhile, in the including 9 studies, 5 articles performed the relationship between the expression of MEG3 and gender [23, 25–27, 30], 3 articles demonstrated that MEG3 were correlated with lymph node metastasis (LNM) [23, 28, 30], and 3 were on distant metastasis DM [23, 26, 30].
Table 1: Characteristics of the included studies
First author |
Year |
Country |
Tumor type |
Sample |
Reference |
Detection method |
Sample size |
Outcome |
Cut-off value |
|---|---|---|---|---|---|---|---|---|---|
Kaihua Lu [22] |
2013 |
China |
NSCLC |
tissue |
GAPDH |
q-PCR |
44 |
OS |
0.27 |
Ming Sun [23] |
2014 |
China |
GC |
tissue |
GAPDH |
q-PCR |
72 |
OS |
0.377-fold |
Lingfei Jia [24] |
2014 |
China |
TSCC |
tissue |
β-actin |
q-PCR |
76 |
OS |
mean |
Zhenye Li [27] |
2014 |
China |
NFPAs |
tissue |
GAPDH |
q-PCR |
52 |
— |
median |
Han Zhuo [25] |
2015 |
China |
HCC |
tissue |
GAPDH |
q-PCR |
72 |
OS, RFS |
median |
Zhizhong Tian [26] |
2015 |
China |
osteosarcoma |
tissue |
GAPDH |
q-PCR |
64 |
OS |
median |
Gang Luo [28] |
2015 |
China |
PC |
tissue |
GAPDH |
q-PCR |
21 |
— |
NA |
Feifei Meng [30] |
2016 |
China |
GC |
tissue |
GAPDH |
q-PCR |
55 |
OS |
NA |
Weili Duan [29] |
2016 |
China |
BC |
tissue |
GAPDH |
q-PCR |
80 |
RFS |
median |
NSCLC: non-small cell lung cancer, GC: gastric cancer (GC), TSCC: tongue squamous cell carcinoma, NFPAs: non-functioning pituitary adenomas, HCC: hepatocellular carcinoma, PC: prostate cancer (PC), BC: bladder cancer; OS: overall survival; RFS: recurrence-free survival.
Association between lncRNA MEG3 and clinicopathological characteristics
As shown in Figure 2, we performed a meta-analysis to evaluate the relationship between the transcription levels of MEG3 and clinicopathological characteristics of patients with cancer. Our results demonstrated that the expression levels of MEG3 were not associated with the gender of patients (OR = 1.41, 95% CI = 0.80–2.47, P = 0.23, fixed-effect) (Figure 2A). Three studies reported the relation between MEG3 and DM. The fixed-effects model was adopted as the significant heterogeneity (I2 = 36%, P = 0.21). Compared with high MEG3 expression group, low MEG3 expression group had a statistic significant elevated DM rate (OR = 2.16, 95% CI = 0.99–4.71, P = 0.05, fixed-effect) (Figure 2B). Unfortunately, there was no correlation in LNM (OR = 2.00, 95% CI = 0.50–8.02, P = 0.33, random-effect) (Figure 2C).
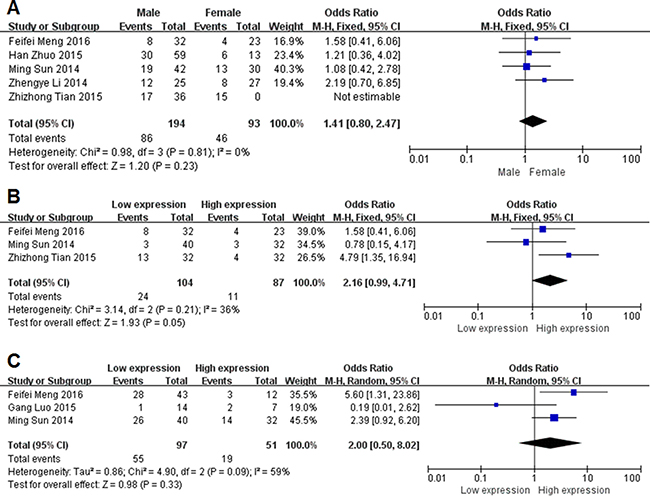
Figure 2: Forest plot for the association between lncRNA MEG3 and Clinicopathological characteristics. (A) gender; (B) distant metastasis; (C) lymph node metastasis.
Association between lncRNA MEG3 and prognosis
We analysis pooled HRs of two group. Six studies with 384 patients were included in this meta-analysis of OS (Figure 3A). Because of no significant heterogeneity (I2 = 11%, P = 0.34), the fixed-effects model was chosen to estimate the pooled HRs with corresponding 95% CIs. Compared with high MEG3 expression group, low MEG3 expression group had a statistic significant reduced OS (HR = 0.43, 95% CI = 0.15–1.24, P = 0.006, fixed-effect) (Figure 3B). Low MEG3 expression correlated with a worse survival. Due to significant heterogeneity (I2 = 0%, P = 0.48), fixed-effects model was used. Analysis showed a pooled HR = 0.43 (95% CI = 0.29–0.92, P = 0.02). Compared with high MEG3 expression group, low MEG3 had a statistic significant reduced RFS.
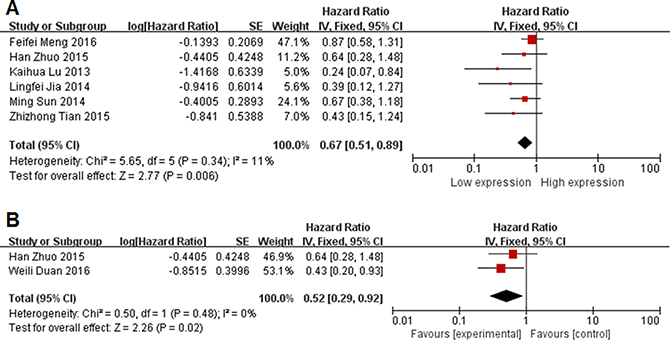
Figure 3: Forest plot for the association between lncRNA MEG3 and prognosis. (A) overall survival; (B) recurrence-free survival.
Publication bias and sensitivity analysis
As shown in Figure 4, Begg’s test was used to perform the publication bias, respectively. In our meta-analysis, Begg’s test indicated there were no publication bias in all groups, due to all the values of P > 0.05. Meanwhile, we used Stata11.0 software to evaluate sensitivity analysis to assess whether the individual studies affected the overall results. The results suggested that individual study had little influence on our final results (Figure 5), and demonstrated that our analysis was relatively stable and credible.
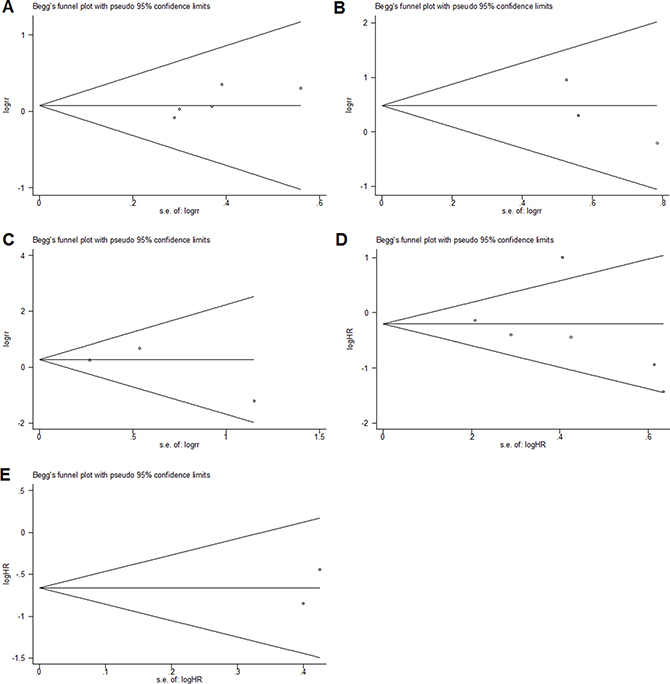
Figure 4: Begg’s test for publication bias. (A) gender; (B) distant metastasis; (C) lymph node metastasis; (D) overall survival; (E) recurrence-free survival.
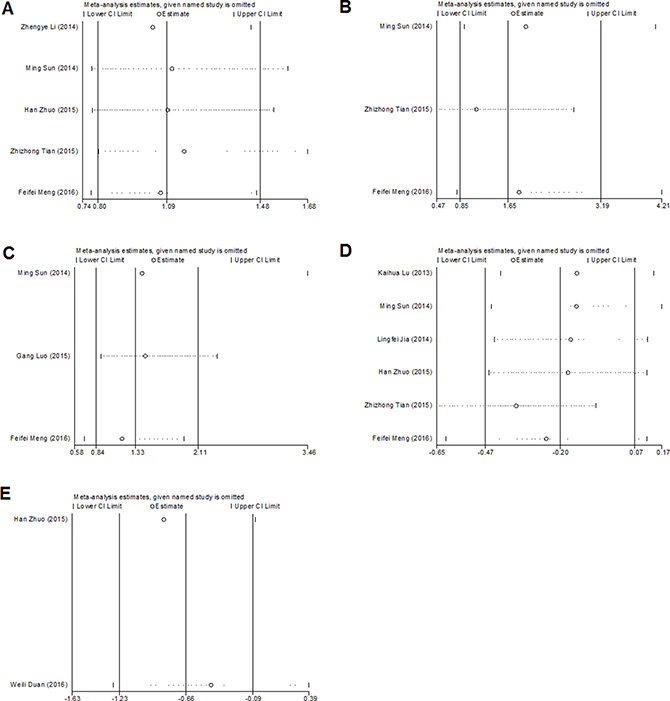
Figure 5: Sensitivity analyses of the studies. (A) gender; (B) distant metastasis; (C) lymph node metastasis; (D) overall survival; (E) recurrence-free survival.
DISCUSSION
Along with the research of human genomics, it is found that only 2% of the genomic sequences are translated into proteins, and most of which are transcribed into non-coding RNA [31]. LncRNAs were once considered to be the noise of genome transcription without biological function. However, recent studies have indicated that lncRNAs are closely related to many diseases, such as neurodegenerative diseases [32], cardiovascular diseases [33], rheumatoid diseases [34] and so on. Meanwhile, abnormal expressions of lncRNAs have been found in tumor tissues, which play important role in the carcinogenesis and aggressive progression of human malignancies [35]. These finding suggested that the potential role of lncRNAs should be further investigated.
MEG3 is an imprinted gene belonging to the DLK1-MEG3 locus located on chromosome 14q32.3 which is the first lncRNA to be found with tumor suppressor function [36]. Previous studies have showed that MEG3 was expressed in brain [36], pituitary [37], ovary [38] and other normal tissues, while the expression was reduced or even lost in a variety of tumor cell lines [39]. Meanwhile, overexpression of MEG3 can inhibit the proliferation of tumor cell lines, which indicates that it plays a role of tumor suppressor genes. In non-small cell lung cancer, Lu et al. have showed that lower expression of MEG3 had advanced clinical features and poor prognosis [22]. In 2015, Yin et al. [40] demonstrated that MEG3 was remarkably decreased in colorectal cancer tissues, comparing adjacent normal control tissues, and the lower expression of MEG3 could promote cells proliferation in vitro. Sun et al. [41] reported that down-regulated MEG3 enhanced the cell proliferation and migration in vitro and increased tumor growth and metastasis in gastric cancer. Thus, MEG3 could be considered as a potential prognostic factor for various cancers. In our meta-analysis, we assessed the prognostic role of MEG3 in cancers. Our results indicated that lower expressions of MEG3 represented a risk factor for OS in cancers (HR = 0.43, 95% CI = 0.15–1.24, P = 0.006, fixed-effect). Furthermore, we found that there was significantly relationship between MEG3 and RFS (95% CI = 0.29–0.92, P = 0.02). From these results, lncRNA MEG3 could be as biomarker for the prognosis of cancers. However, further large-scale studies should be conducted.
It has been reported that MEG3 can play important role in inhibiting cancer through a variety of ways, which related to DNA methylation, P53 pathway, Rb pathway and so on [42]. Sun L et al. suggested that lncRNA EWSAT1 can enhance osteosarcoma cell growth and metastasis through suppression of MEG3 expression [43]. Besides, MEG3 plays important role in the epigenetic regulation of epithelial-mesenchymal transition promotes in lung cancer. In this meta-analysis, we evaluate the association between expression levels of MEG3 and cancer. We found that lower levels of MEG3 were more prone to lead to DM (OR = 2.16, 95% CI = 0.99-4.71, P = 0.05). Unfortunately, there was no correlation in LNM (OR = 2.00, 95% CI = 0.50–8.02, P = 0.33); insufficient sample size was the possible reason for the different results in this meta-analysis. Therefore, further studies should be done with larger sample sizes.
It should be stressed the limitations in our analysis. Most studies reported positive results, but those with negative results were generally less likely to be published. In addition, studies included in the meta-analysis most came from People’s Republic of China, which might affect the results. Finally, there were insufficient data to fully confirm the association between MEG3 and clinicopathological characteristics, which needs more studies. Therefore, the results of this meta-analysis should be confirmed in future studies.
In conclusion, despite the limitations described above, our meta-analysis reveals that the depressed expression of lncRNA MEG3 is significantly associated with DM, OS and RFS in patients with diverse cancers and could be a potential prognostic marker for cancers. However, large-scale and comprehensive researches were needed to illuminate our results. Well-designed studies related to specific cancer types and large sample sizes are needed to confirm the prognostic value of decreased lncRNA MEG3 in various cancers.
MATERIALS AND METHODS
Literature search strategies
We searched the databases PubMed, Cochrane library, Chinese National Knowledge Infrastructure, and Chinese Wan Fang database for studies published up to November 2016 to obtain relevant articles for the meta-analysis. The search strategy used both medical subject heading terms and free-text words to increase the sensitivity of the search. The keywords for the search were as follows: “MEG3 and cancer”, “long non-coding RNA MEG3”, “lncRNA MEG3”, “MEG3”. There was no language restriction. Meanwhile, reference lists of relevant articles were also reviewed to identify potential eligible papers.
Inclusion and exclusion criteria
In this meta-analysis, eligible studies had to meet the following standards: 1)cohort design, 2) articles investigating the relation of MEG3 and cancer patients; 3) the expression levels of MEG3 in primary tumor tissues were measured, 4) sufficient original data for calculating odds ratios (ORs), hazard ratios (HRs) and their 95% confidence interval (95% CI). If the articles only provided survival curves without offering HR and 95% CI directly, appropriate data were extracted from the survival curves using Engauge Digitizer 4.1 software and the logHR and selogHR calculated according to Tierney et al. [44]. If there were duplicated data, we chose the most complete data or the most recent one. Exclusion criteria were as follow: 1) studies without usable or insufficient data, 2) case reports, 3) letters and conference abstracts.
Data extraction
Two investigators extracted and reviewed relevant data from the eligible studies independently, including first author, year of publication, country, site of cancer, method, case number, cut-off value. If there were disagreements, a consensus was reached by the third investigator.
Statistical analysis
Odds ratios (ORs) and 95% CI were used to evaluate the relationship between MEG3 and in these inclusive articles. The features included gender, distant metastasis (DM), and lymph node metastasis (LNM). Meanwhile, HRs and 95% CIs were used to assess the association between MEG3 and cancer prognosis (Relapse-free survival and overall survival). We used Revman5.3 Software (Revman, the Cochrane Collaboration) to perform the meta-analysis and evaluate heterogeneity between studies by Cochrane Q-test and P-values. If heterogeneity was present (I2 ≥ 50% or P ≤ 0.05), random-effect model was used to calculate pooled HRs or ORs. If not, the fixed-effect model was more appropriate [45, 46]. The Stata11.0 Software (Stata, College Station) was performed to evaluate the sensitivity and publication bias of the studies. Publication bias was evaluated by Begg’s test, P < 0.05 was considered statistically significant.
ACKNOWLEDGMENTS AND FUNDING
We are indebted to the donors whose names were not included in the author list, but who participated in this program. This work was supported by the National Natural Science Foundation of China (81670270).
CONFLICTS OF INTEREST
All authors declare that there is no conflicts of interests.
REFERENCES
1. Hsieh HJ, Peng G. Cellular responses to replication stress: Implications in cancer biology and therapy. DNA repair. 2016.
2. Dai L, Ma C, Zhang Z, Zeng S, Liu A, Tang S, Ren Q, Sun Y, Xu C. DAPK Promoter Methylation and Bladder Cancer Risk: A Systematic Review and Meta-Analysis. PloS one. 2016; 11:e0167228.
3. Schuler PJ, Brandau S. Letter to the Editor: Adenosine Producing Mesenchymal Stem Cells. Stem cells. 2016.
4. Sequeiros T, Rigau M, Chiva C, Montes M, Garcia-Grau I, Garcia M, Diaz S, Celma A, Bijnsdorp I, Campos A, Di Mauro P, Borros S, Reventos J, et al. Targeted proteomics in urinary extracellular vesicles identifies biomarkers for diagnosis and prognosis of prostate cancer. Oncotarget. 2017; 8:4960–4976. doi: 10.18632/oncotarget.13634.
5. Stoffel EM, Yurgelun MB. Genetic predisposition to colorectal cancer: Implications for treatment and prevention. Semin Oncol. 2016; 43:536–542.
6. Kojima T, Kawai K, Miyazaki J, Nishiyama H. Biomarkers for precision medicine in bladder cancer. Int J Clin Oncol. 2016.
7. Ribeiro IP, Barroso L, Marques F, Melo JB, Carreira IM. Early detection and personalized treatment in oral cancer: the impact of omics approaches. Mol Cytogenet. 2016; 9:85.
8. Gorin MA, Verdone JE, van der Toom E, Bivalacqua TJ, Allaf ME, Pienta KJ. Circulating tumour cells as biomarkers of prostate, bladder, and kidney cancer. Nat Rev Urol. 2016.
9. Meng Q, Ren M, Li Y, Song X. LncRNA-RMRP Acts as an Oncogene in Lung Cancer. PloS one. 2016; 11:e0164845.
10. Zhuo C, Jiang R, Lin X, Shao M. LncRNA H19 inhibits autophagy by epigenetically silencing of DIRAS3 in diabetic cardiomyopathy. Oncotarget. 2017; 8:1429–1437. doi: 10.18632/oncotarget.13637.
11. Fotouhi Ghiam A, Taeb S, Huang X, Huang V, Ray J, Scarcello S, Hoey C, Jahangiri S, Fokas E, Loblaw A, Bristow RG, Vesprini D, Boutros P, et al. Long non-coding RNA urothelial carcinoma associated 1 (UCA1) mediates radiation response in prostate cancer. Oncotarget. 2017; 8:4668–4689. doi: 10.18632/oncotarget.13576.
12. Han D, Zhang X, Zhang J, Guo X, Zheng Y, Sui S, Zheng J. Oleanolic acid suppresses vascular smooth muscle cell proliferation by increasing lincRNA-p21 expression. Oncol Lett. 2016; 12:3519–3522.
13. Wang D, Wang D, Wang N, Long Z, Ren X. Long Non-Coding RNA BANCR Promotes Endometrial Cancer Cell Proliferation and Invasion by Regulating MMP2 and MMP1 via ERK/MAPK Signaling Pathway. Cell Physiol Biochem. 2016; 40:644–656.
14. Liu S, Zhang M, Qu P. Expression level and clinical significance of HOX transcript antisense intergenic RNA in cervical cancer: a meta-analysis. Sci Rep. 2016; 6:38047.
15. Chang L, Wang G, Jia T, Zhang L, Li Y, Han Y, Zhang K, Lin G, Zhang R, Li J, Wang L. Armored long non-coding RNA MEG3 targeting EGFR based on recombinant MS2 bacteriophage virus-like particles against hepatocellular carcinoma. Oncotarget. 2016; 7:23988–24004. doi: 10.18632/oncotarget.8115.
16. Ying L, Huang Y, Chen H, Wang Y, Xia L, Chen Y, Liu Y, Qiu F. Downregulated MEG3 activates autophagy and increases cell proliferation in bladder cancer. Mol Biosyst. 2013; 9:407–411.
17. Qin R, Chen Z, Ding Y, Hao J, Hu J, Guo F. Long non-coding RNA MEG3 inhibits the proliferation of cervical carcinoma cells through the induction of cell cycle arrest and apoptosis. Neoplasma. 2013; 60:486–492.
18. Cao X, Zhuang S, Hu Y, Xi L, Deng L, Sheng H, Shen W. Associations between polymorphisms of long non-coding RNA MEG3 and risk of colorectal cancer in Chinese. Oncotarget. 2016; 7:19054–19059. doi: 10.18632/oncotarget.7764.
19. Modali SD, Parekh VI, Kebebew E, Agarwal SK. Epigenetic regulation of the lncRNA MEG3 and its target c-MET in pancreatic neuroendocrine tumors. Mol Endocrinol. 2015; 29:224–237.
20. Tang W, Dong K, Li K, Dong R, Zheng S. MEG3, HCN3 and linc01105 influence the proliferation and apoptosis of neuroblastoma cells via the HIF-1alpha and p53 pathways. Sci Rep. 2016; 6:36268.
21. Yan H, Yuan J, Gao L, Rao J, Hu J. Long noncoding RNA MEG3 activation of p53 mediates ischemic neuronal death in stroke. Neuroscience. 2016; 337:191–199.
22. Lu KH, Li W, Liu XH, Sun M, Zhang ML, Wu WQ, Xie WP, Hou YY. Long non-coding RNA MEG3 inhibits NSCLC cells proliferation and induces apoptosis by affecting p53 expression. BMC cancer. 2013; 13:461.
23. Sun M, Xia R, Jin F, Xu T, Liu Z, De W, Liu X. Downregulated long noncoding RNA MEG3 is associated with poor prognosis and promotes cell proliferation in gastric cancer. Tumour Biol. 2014; 35:1065–1073.
24. Jia LF, Wei SB, Gan YH, Guo Y, Gong K, Mitchelson K, Cheng J, Yu GY. Expression, regulation and roles of miR-26a and MEG3 in tongue squamous cell carcinoma. Int J Cancer. 2014; 135:2282–2293.
25. Zhuo H, Tang J, Lin Z, Jiang R, Zhang X, Ji J, Wang P, Sun B. The aberrant expression of MEG3 regulated by UHRF1 predicts the prognosis of hepatocellular carcinoma. Mol Carcinog. 2016; 55:209–219.
26. Tian ZZ, Guo XJ, Zhao YM, Fang Y. Decreased expression of long non-coding RNA MEG3 acts as a potential predictor biomarker in progression and poor prognosis of osteosarcoma. Int J Clin Exp Pathol. 2015; 8:15138–15142.
27. Li Z, Li C, Liu C, Yu S, Zhang Y. Expression of the long non-coding RNAs MEG3, HOTAIR, and MALAT-1 in non-functioning pituitary adenomas and their relationship to tumor behavior. Pituitary. 2015; 18:42–47.
28. Luo G, Wang M, Wu X, Tao D, Xiao X, Wang L, Min F, Zeng F, Jiang G. Long Non-Coding RNA MEG3 Inhibits Cell Proliferation and Induces Apoptosis in Prostate Cancer. Cell Physiol Biochem. 2015; 37:2209–2220.
29. Duan W, Du L, Jiang X, Wang R, Yan S, Xie Y, Yan K, Wang Q, Wang L, Zhang X, Pan H, Yang Y, Wang C. Identification of a serum circulating lncRNA panel for the diagnosis and recurrence prediction of bladder cancer. Oncotarget. 2016; 7:78850–78858. doi: 10.18632/oncotarget.12880.
30. Meng FF, Si Jl, Liu L, Cui JY, Qi YQ, Lv M. Expression of long non-coding RNA MEG3 and its relationship with the prognosis of human gastric cancer. Chin J Clin Oncol. 2016; :659–662.
31. Carninci P, Kasukawa T, Katayama S, Gough J, Frith MC, Maeda N, Oyama R, Ravasi T, Lenhard B, Wells C, Kodzius R, Shimokawa K, Bajic VB, et al. The transcriptional landscape of the mammalian genome. Science. 2005; 309:1559–1563.
32. Kumar L, Shamsuzzama, Haque R, Baghel T, Nazir A. Circular RNAs: the Emerging Class of Non-coding RNAs and Their Potential Role in Human Neurodegenerative Diseases. Mol Neurobiol. 2016.
33. Bar C, Chatterjee S, Thum T. Long Noncoding RNAs in Cardiovascular Pathology, Diagnosis, and Therapy. Circulation. 2016; 134:1484–1499.
34. Zhang Y, Xu YZ, Sun N, Liu JH, Chen FF, Guan XL, Li A, Wang F, Zhao QF, Wang HY, Song SS, Yu W, Zhao JN, et al. Long noncoding RNA expression profile in fibroblast-like synoviocytes from patients with rheumatoid arthritis. Arthritis Res Ther. 2016; 18:227.
35. He Q, Tian L, Jiang H, Zhang J, Li Q, Sun Y, Zhao J, Li H, Liu M. Identification of laryngeal cancer prognostic biomarkers using an inflammatory gene-related, competitive endogenous RNA network. Oncotarget. 2017; 8:9525–9534. doi: 10.18632/oncotarget.13627.
36. Liu J, Li Q, Zhang KS, Hu B, Niu X, Zhou SM, Li SG, Luo YP, Wang Y, Deng ZF. Downregulation of the Long Non-Coding RNA Meg3 Promotes Angiogenesis After Ischemic Brain Injury by Activating Notch Signaling. Mol Neurobiol. 2016.
37. Chunharojrith P, Nakayama Y, Jiang X, Kery RE, Ma J, De La Hoz Ulloa CS, Zhang X, Zhou Y, Klibanski A. Tumor suppression by MEG3 lncRNA in a human pituitary tumor derived cell line. Mol Cell Endocrinol. 2015; 416:27–35.
38. Sheng X, Li J, Yang L, Chen Z, Zhao Q, Tan L, Zhou Y, Li J. Promoter hypermethylation influences the suppressive role of maternally expressed 3, a long non-coding RNA, in the development of epithelial ovarian cancer. Oncol Rep. 2014; 32:277–285.
39. Terashima M, Tange S, Ishimura A, Suzuki T. MEG3 long noncoding RNA contributes to the epigenetic regulation of epithelial-mesenchymal transition in lung cancer cell lines. J Biol Chem. 2016.
40. Yin DD, Liu ZJ, Zhang E, Kong R, Zhang ZH, Guo RH. Decreased expression of long noncoding RNA MEG3 affects cell proliferation and predicts a poor prognosis in patients with colorectal cancer. Tumour Biol. 2015; 36:4851–4859.
41. Sun XH, Yang LB, Geng XL, Wang R, Zhang ZC. Increased expression of lncRNA HULC indicates a poor prognosis and promotes cell metastasis in osteosarcoma. Int J Clin Exp Pathol. 2015; 8:2994–3000.
42. Balik V, Srovnal J, Sulla I, Kalita O, Foltanova T, Vaverka M, Hrabalek L, Hajduch M. MEG3: a novel long noncoding potentially tumour-suppressing RNA in meningiomas. J Neurooncol. 2013; 112:1–8.
43. Sun L, Yang C, Xu J, Feng Y, Wang L, Cui T. Long Noncoding RNA EWSAT1 Promotes Osteosarcoma Cell Growth and Metastasis Through Suppression of MEG3 Expression. DNA Cell Biol. 2016.
44. Tierney JF, Stewart LA, Ghersi D, Burdett S, Sydes MR. Practical methods for incorporating summary time-to-event data into meta-analysis. Trials. 2007; 8:16.
45. DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986; 7:177–188.
46. Mantel N, Haenszel W. Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst. 1959; 22:719–748.

