INTRODUCTION
Sertoli cells are essential for normal spermatogenesis which comprises the mitosis of spermatogonial stem cells, the meiosis of spermatocytes, and the spermiogenesis of haploid spermatids. As the only type of somatic cells within the seminiferous tubule of the testis, Sertoli cells provide critical structural and nutritional supports for male germ cells, including spermatogonia, spermatocytes and spermatids [1]. The tight junction, constituted by Sertoli cells, is the key structure of the blood-testis barrier (BTB) to ensure the stabilization of the microenvironment or niche of the testis. Tight junction proteins, e.g., zona occludens 1 (ZO1), claudin 11 (CLDN11) [2], and occluding (OCLN), are produced by Sertoli cells, and they play vital roles in controlling the function of BTB [3]. In addition to structural support, Sertoli cells play significant roles in promoting the self-renewal, differentiation, and apoptosis of spermatogonial stem cells via secreting a number of growth factors, including bone morphogenetic protein 4 (BMP4), stem cell factor (SCF), leukemia inhibitory factor (LIF), glial cell derived neurotrophic factor (GDNF), fibroblast growth factor 2 (FGF2), and epidermal growth factor (EGF) [1]. Consequently, any abnormality in the number and/or function of Sertoli cells can lead to spermatogenic arrest and eventual spermatogenesis failure [4–6]. Therefore, Sertoli cells have significant applications in reproductive medicine.
A number of lines have implicated that Sertoli cells could have important applications in regenerative medicine because of their great plasticity. Firstly, Sertoli cells could secrete insulin persistently after being genetically transformed [7], which may provide new therapeutic approach for diabetes. Secondly, Sertoli cells could be reprogrammed to the pluripotent stem cells and to morphologic, phenotypic and functional neural stem cells by transferring certain genes or transcription factors [8–10]. These studies illustrate that Sertoli cells might be utilized for treating neural disorders and other human diseases. Finally, Sertoli cells could be converted into Leydig cells after deletion of gene Wt1 (Wilms tumor 1) [11], suggesting that Sertoli cells could be used in the therapy for testosterone- or other hormones-deficiency disorders.
On the other hand, the application of human Sertoli cells has been handicapped due to the following factors: i) it is hard to obtain human testicular tissues to separate human Sertoli cells; ii) the isolation and purification of human Sertoli cells from testis tissues seem to be tedious and time-consuming; iii) the number of primary human Sertoli cells is rare; and iv) primary human Sertoli cells proliferate limitedly in vitro. Therefore, it is necessary to establish a human Sertoli cell line to obtain adequate human Sertoli cells for basic research to uncover the mechanisms underlying human spermatogenesis as well as for their applications in both reproductive and regenerative medicine. Currently, several germ cell lines in human or rodents and Sertoli cell lines in rodents have been established with either simian virus large T antigen (SV40-LTAg) or telomerase (TERT) [12–15]. However, human Sertoli cell line has not yet been available. In this study, we have for the first time reported a stable human Sertoli cell line, namely hS1 cells, via the overexpression of human telomerase (hTERT). This cell line possessed the phenotypic characteristics of primary human Sertoli cells and it had an unlimited proliferation potential and high safety, which could offer a sufficient source of human Sertoli cells for their use from the bench to the bed side.
RESUTLS
Immortalization of human Sertoli cells
Human Sertoli cells were separated from testicular tissues of obstructive azoospermic (OA) patients with normal spermatogenesis using a two-step enzymatic digestion and followed by differential plating (Figure 1A, 1C), and they were infected with the lentivirus carrying hTRET by polybrene. The expression of hTRET was driven by the promoter of EF1A (Figure 1B), and EGFP was utilized as a reporter gene (Figure 1B). GFP-positive cells (the immortalized Sertoli cells), namely hS1 cells, were isolated and purified by FACS, and the EGFP was stably detected under a fluorescence microscope (Figure 1D). Western blots showed that the protein of hTERT was stably expressed in human Sertoli cell line at passage 10 (P10), P15, and P20 (Figure 1E). In morphology, the immortalized human Sertoli cells extended their cytoplasm with irregular nuclei under a phase-contrast microscopy.
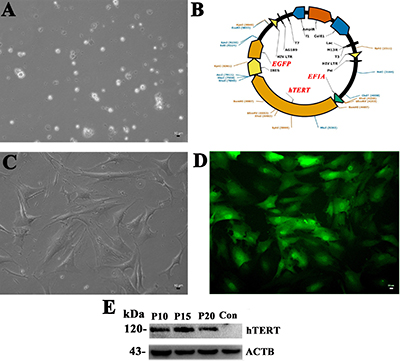
Figure 1: Immortalization of human Sertoli cells. (A) Human Sertoli cells were freshly isolated from human testis tissues derived from OA patients by a two-step enzymatic digestion and differential plating. (B) The diagram showed the structure of lentivirus vector namely Lv-EF1A-hTERT-IRES-EGFP. (C) The isolated human Sertoli cells adhered to the culture dish. (D) EGFP-positive Sertoli cells infected with Lv-EF1A-hTERT-IRES-EGFP were sorted by FACS. Scale bars in A, C, D = 10 μm. (E) The expression of hTERT in the immortalized Sertoli cells at passage 10, 15, 20 and primary human Sertoli cells (the control). ACTB was used as a loading control of the proteins.
Phenotypic identification of human Sertoli cell line
To check whether the EGFP-positive cells were human Sertoli cell in phenotype, we detected a number of markers for primary human Sertoli cells using RT-PCR, Western blots and immunocytochemistry. RT-PCR revealed that the transcripts of AR, BMP4, BMP6, GATA4, GDNF, SCF, SOX9 and WT1 were expressed in the immortalized human cells (Figure 2A), which was comparable to the expression level of these genes in primary human Sertoli cells (Figure 2B). In contrast, the mRNA of 3β-HSD, SMA and VASA, markers for Leydig cells, myoid cells and germ cells, respectively, was undetectable in these cells (Figure 2A). High level of hTERT was observed in the immortalized human cells (Figure 2A). Western blots showed that the proteins of SCF, GDNF, BMP4, WT1 and SOX9 were expressed in human hS1 cells (Figure 2C), whereas 3β-HSD, SMA and VASA were undetected in this cell line (Figure 2C). Notably, the levels of FSHR and AR were much higher in hS1 cells compared to primary human Sertoli cells (Figure 2C).
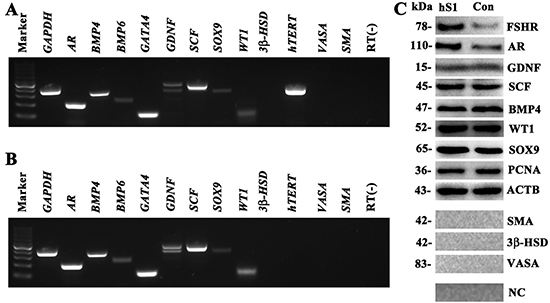
Figure 2: Phenotypic feature of the immortalized human Sertoli cells. (A–B) RT-PCR showed the expression of AR, BMP4, BMP6, GATA4, GDNF, SCF, SOX9, WT1, hTERT, 3β-HSD, SMA, and VASA in the immortalized human Sertoli cells (A) and primary human Sertoli cell (B). GAPDH was used as a loading control of total RNA, and RNA sample without RT (RT-) but with PCR of GAPDH primers served as a negative control. (C) Western blot revealed the proteins of FSHR, AR, GDNF, SCF, BMP4, WT1, SOX9, PCNA, 3β-HSD, VASA, and SMA in the immortalized human Sertoli cells (hS1) and primary Sertoli cells (Con). ACTB was used as a loading control of proteins, while replacement of primary antibodies with PBS served as negative controls (NC).
Furthermore, immunocytochemistry displayed that the immortalized cells were positive for SOX9 (Figure 3A), WT1 (Figure 3B), OCLN (Figure 3C), VIM (Figure 3D), SCF (Figure 3E), BMP4 (Figure 4A), GDNF (Figure 4B), but not for 3β-HSD (Figure 4C), SMA (Figure 4D), and VASA (Figure 4E). Replacement of primary antibodies with isotype IgGs was used as a negative control, and no immunnostaining was seen in these cells (Figure 4F), thus verifying specific expression of these proteins mentioned above in the immortalized cells.
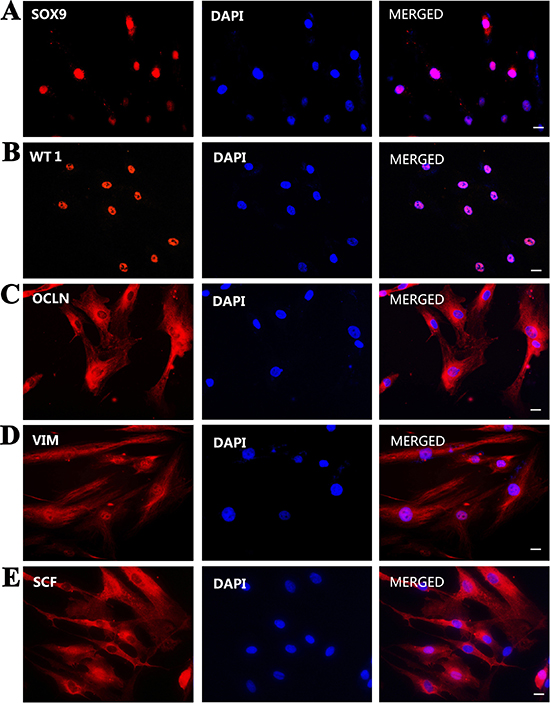
Figure 3: Phenotypic characteristics of the immortalized human Sertoli cells. (A–E) Immunocytochemistry demonstrated the expression of SOX9 (A), WT1 (B), OCLN (C), VIM (D), and SCF (E) in the immortalized human Sertoli cells. Scale bars in A–E = 10 μm.
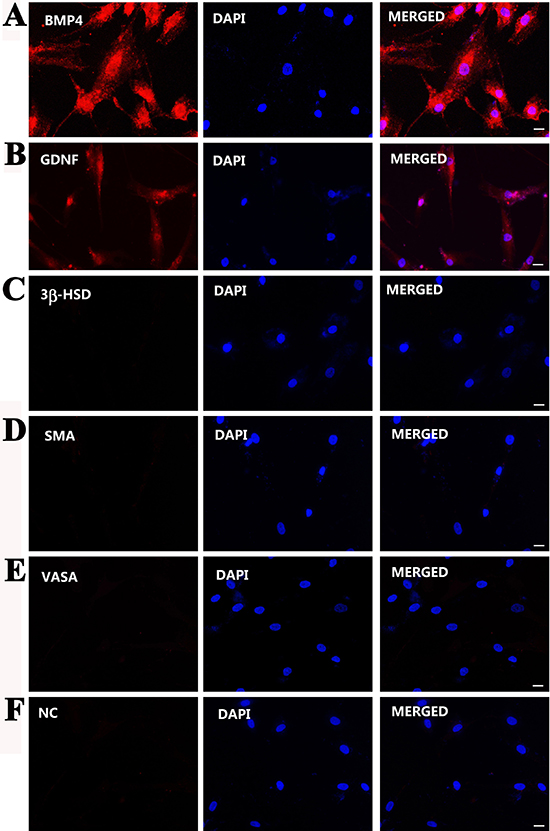
Figure 4: Phenotypic characterization of the immortalized human Sertoli cells. (A–F) Immunocytochemistry displayed the expression of BMP4 (A), GDNF (B), 3β-HSD (C), SMA (D), and VASA (E) in the immortalized human Sertoli cells. Replacement of primary antibodies with isotype IgGs (F) served as negative controls (NC). Scale bars in A–F = 10 μm.
Similar global gene expression profiles between human Sertoli cell line and primary human Sertoli cells
We next compared global gene expression profiling between the immortalized human cell line and primary human Sertoli cells using microarray analysis. Total RNA was quantified by the NanoDrop ND-2000 and RNA integrity was assessed using Agilent Bioanalyzer 2100. The total RNA from the immortalized human Sertoli cells and primary human Sertoli cells was of great quality, as evidenced by the RNA integrity number (RIN) values over 8.0 (Figure 5C, 5E, Table 1). In total, 24,118 genes were detected in both cell types by microarray analysis. There were 3,641 (15.0%) and 3,496 (14.5%) differentially expressed genes (up-regulated or down-regulated with 2.0 folds or more, respectively) between the immortalized human Sertoli cells and primary human Sertoli cells (Table 2). Thus, human Sertoli cell line had a 70.4% similarity of global gene profiles with primary human Sertoli cells. Microarray analysis showed that there was no significant change in the transcripts of WT1, GATA4, GDNF, GATA1 and SOX9 between the immortalized human Sertoli cells and primary human Sertoli cells (Table 3). The expression level of BMP4, BMP6 and FGF2 was higher in the immortalized human Sertoli cells than primary human Sertoli cells, while LIF mRNA was lower in the immortalized human Sertoli cells than primary Sertoli cells (Table 3).
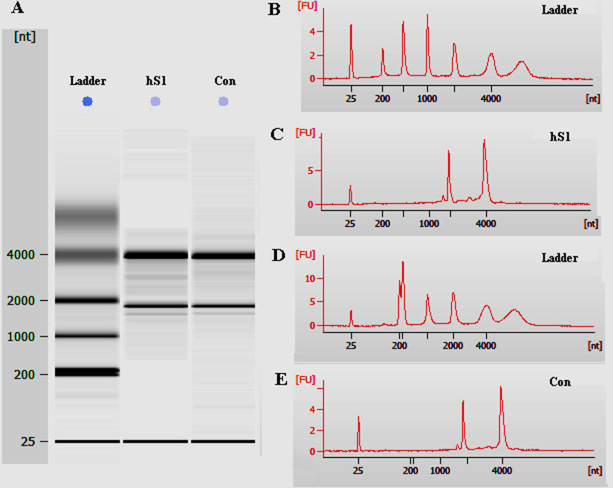
Figure 5: The quality assessment of total RNA of the immortalized human Sertoli cells and primary human Sertoli cells. (A) Total RNA was quantified by the NanoDrop ND-2000 and the RNA integrity was assessed using Agilent Bioanalyzer 2100. (B-E) Electropherogram by Agilent bioanalyzer indicated the concentrations and nucleotides (nt) of RNA isolated from the immortalized human Sertoli cells (C) and primary human Sertoli cells (E). B and D were RNA ladders.
Table 1: The quality of total RNA of the immortalized Sertoli cells (hS1) and primary human Sertoli cells (Control) by the NanoDrop ND-2000
Samples |
Concentration |
A260/280 |
A260/230 |
Volume(μl) |
Quality |
|
|---|---|---|---|---|---|---|
(μg/μl) |
28S/18S |
RIN |
||||
Control |
0.0967 |
1.73 |
0.18 |
10 |
2 |
9 |
hS1 cells |
0.1123 |
2.06 |
0.17 |
10 |
1.9 |
8.8 |
Table 2: The differently expressed gene profiles between the immortalized Sertoli cells and primary human Sertoli cells (Control)
Control |
Human Sertoli cell line |
Up-regulated |
Down-regulated |
Total genes |
Similarity |
|---|---|---|---|---|---|
Primary Sertoli cells |
hS1 cells |
3,641(15.0%) |
3,496(14.5%) |
24,118 |
70.4% |
Table 3: The expression of representative genes between the immortalized Sertoli cells and primary human Sertoli cells (Control)
Gene Symbol |
Primary Sertoli cell signal (normalized) |
hS1 cell signal |
Log2 FC (hS1 vs. Control) |
|---|---|---|---|
GDNF |
2.7378905 |
4.001061 |
1.2631705 |
BMP4 |
7.070616 |
10.179245 |
3.108629 |
BMP6 |
2.913889 |
6.345799 |
3.43191 |
LIF |
8.199608 |
9.237209 |
1.037601 |
FGF2 |
7.0719824 |
10.704937 |
3.6329546 |
SOX9 |
1.4255941 |
1.4340813 |
0.0084872 |
WT1 |
2.236927 |
2.0595062 |
–0.1774208 |
GATA4 |
10.017227 |
9.562365 |
–0.4548 |
GATA1 |
2.7784665 |
2.5375004 |
–0.2409661 |
To confirm the results of microarray analysis, real-time PCR was performed to check the expression of numerous genes. We found that there was no significant difference in the expression of GATA4, GDNF, SOX9, GATA1 and WT1 between the immortalized human Sertoli cells and primary human Sertoli cells (Figure 6). The transcripts of BMP4, BMP6 and FGF2 were higher in the immortalized human Sertoli cells than primary human Sertoli cells, whereas LIF was expressed at a lower level in the immortalized human Sertoli cells compared to primary Sertoli cells (Figure 6). These results of real-time PCR were consistent with the data of our mRNA microarray.
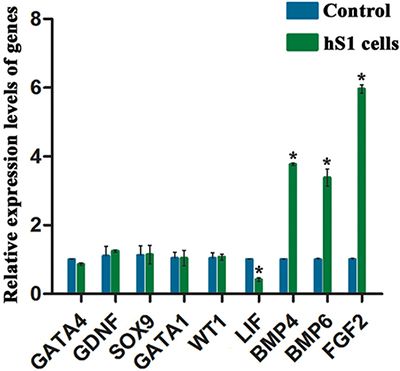
Figure 6: Expression levels of numerous genes between the immortalized human Sertoli cells and primary human Sertoli cells. Real-time PCR demonstrated that the expression of GATA4, GDNF, SOX9, GATA1, WT1, LIF, BMP4, BMP6, and FGF2 in the immortalized human Sertoli cells and primary human Sertoli cells. * indicated statistically significant differences (p < 0.05) between the immortalized human Sertoli cells and primary human Sertoli cells.
Proliferation potential of human Sertoli cell line
We utilized Western blots, immunocytochemistry, and CCK-8 assay to evaluate the proliferative potentials of human Sertoli cell line. Western blots showed similar expression level of PCNA protein between human Sertoli cell line and primary human Sertoli cells (Figure 2C). Immunocytochemistry revealed that more than 70% of human Sertoli cell line was positive for ki-67 (Figure 7A), suggesting this cell line had a high proliferation capacity.
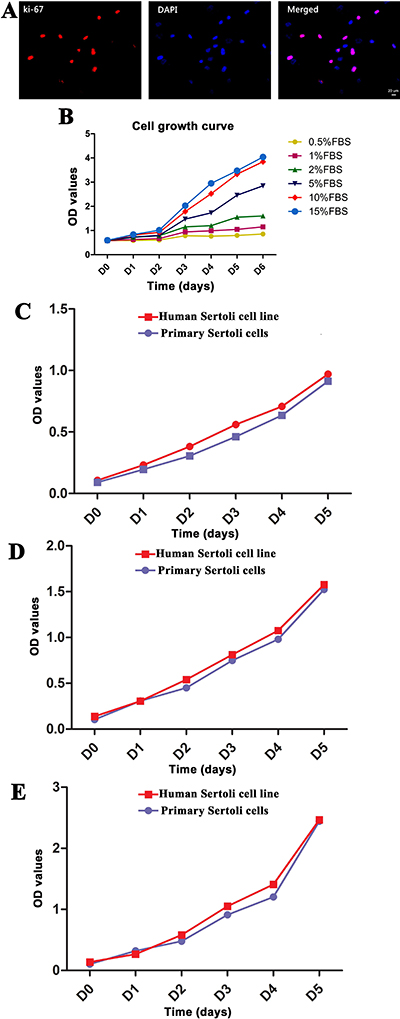
Figure 7: Proliferation capacity of the immortalized human Sertoli cells in vitro. (A) Immunocytochemistry revealed the expression of Ki-67 in human Sertoli cell line. Scale bar in A = 20 μm. (B) CCK-8 assays were utilized to compare the effect of various concentrations of FBS ranging from 0.5% to 15% on the growth of human Sertoli cell line. (C–E) CCK-8 assays were employed to compare the proliferation of human Sertoli cell line and primary human Sertoli cells when they were cultured with 2% FBS (C), 5% FBS (D), and 10% FBS (E).
We next checked the proliferative potentials of the immortalized human Sertoli cells under different concentrations of fetal bovine serum (FBS). CCK-8 assays showed that the proliferation of the immortalized human Sertoli cells was dependent on the concentrations of FBS. There was no obvious difference in the proliferation of the cell line when cultured in the medium supplemented with 0.5%–2% FBS, and 10% and 15% FBS had the optimal effect for promoting cell proliferation (Figure 7B). The doubling time of human Sertoli cell line was 1.3 days for culturing with 10% FBS and 1.03 days with 15% FBS (Figure 7B). Notably, human Sertoli cell line proliferate more rapidly than primary human Sertoli cells cultured with 2% FBS (Figure 7C), 5% FBS (Figure 7D), and 10% FBS (Figure 7E). Until now, the immortalized human Sertoli cells had been cultured for over 6 months in DMEM/F-12 with 10% FBS for more than 30 passages. During this long period of culture and different passages, this cell line didn’t show morphological change or contact inhibition.
Human Sertoli cell line assumed 90% of normal karyotype and excluded Y chromosome microdeletions
Abnormal karyotype may occur in cell line due to transferring genes. We asked whether this alteration existed in human Sertoli cell line after overexpressing hTERT. Conventional cytogenetic analysis displayed that 90% of the immortalized human Sertoli cells had normal karyotype with 23 pairs of chromosomes (Figure 8A–Figure 8D), while about 10% of these cells showed abnormal karyotype with unbalanced translocation (Figure 8E–8F).
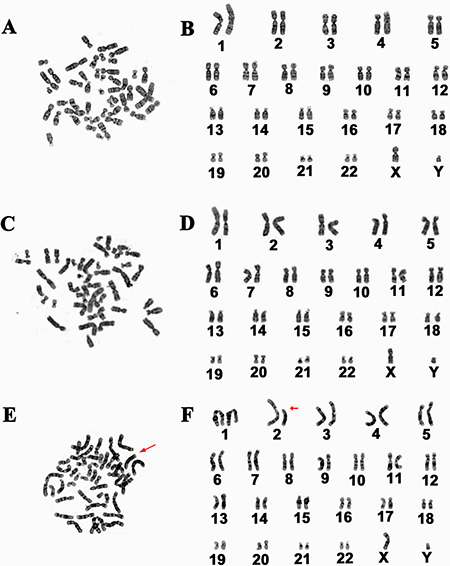
Figure 8: Karyotype analysis of the immortalized human Sertoli cells. (A–F) Cytogenetic assay revealed normal karyotype (A–D) and abnormal karyotype (E–F) in the immortalized human Sertoli cells. The data were calculated from more than 100 cells of human Sertoli cell line.
Multiplex real-time PCR was used to assess whether there were Y microdeletions in the immortalized human Sertoli cells. As illustrated in Figure 9A, eight specific STS markers from AZFa, AZFb and AZFc regions, including ZFX/ZFY, SRY, sY254, sY127, sY86, sY134, sY84 and sY255, were all detected in the immortalized human Sertoli cells, implicating that this cell line excludes Y chromosome microdeletions.
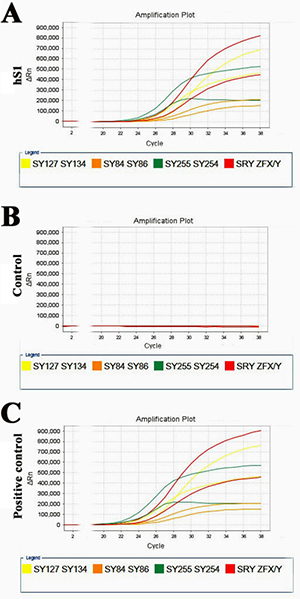
Figure 9: Y chromosome microdeletion analysis of the immortalized human Sertoli cells. (A–C) Multiplex real-time PCR displayed the expression of all eight specific STS markers from AZFa, AZFb and AZFc regions, containing ZFX/ZFY, SRY, sY254, sY127, sY86, sY134, sY84 and sY255, in the immortalized human Sertoli cells (A). DNA from normal human blood served as a positive control (C) and water substituted for DNA as a negative control (B).
Human Sertoli cell line didn’t form tumors in xenografting mice
We further asked whether the immortalized human Sertoli cells had the ability to form tumors. Thirty nude mice were utilized for xenotransplantation assay in vivo. After 8 weeks of cell transplantation, no tumor formation was observed at each skin site of recipient mice (Figure 10A). H&E staining showed that there was regular and normal structure in the skin sites of transplantation (Figure 10B). In parallel, no tumor formation was seen in recipient mice without cell transplantation (Figure 10C–10D). In contrast, the tissue sections from skin of mice transplanted with PC3 cells displayed aberrant structure, and plenty of cancer cells occurred under the epidermis (Figure 10E–10F).
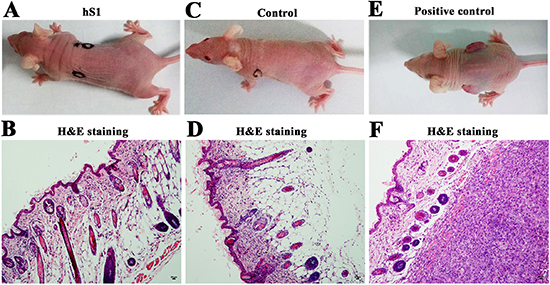
Figure 10: Tumorgenesis of the immortalized human Sertoli cells by nude mouse xenografting assays. (A-F) Digital camera and H&E staining revealed no tumor formation in recipient mice with transplantation of the immortalized human Sertoli cells (A, B) or without cell transplantation (C, D). Tumor formation was observed in the recipient mice with transplantation of PC3 cells (E, F). Scale bars in B, D, F= 20 μm.
DISCUSSION
Sertoli cells have great applications in basic research on uncovering mechanisms of human spermatogenesis and clinical usage. Although much progress has been made on genetic and epigenetic regulation of Sertoli cells in rodent spermatogenesis, very little is known about the roles and molecular mechanisms of human Sertoli cells in controlling the process of human spermatogenesis. This might be due to the facts that it is rather difficult to obtain human testis tissues and that primary human Sertoli cells have a limited proliferation capacity in vitro. Therefore, it is essential to establish a human Sertoli cell line to obtain sufficient cells for their usage from the bench to the bed side. There were several Sertoli cell lines that have been established in rodents [13, 16, 17]. Nevertheless, no human Sertoli cell line has yet been available. Here we have for the first time established a stable human Sertoli cell line by overexpressing hTERT. Immortalization of cells using TERT has certain advantages over the methods using the SV40 large T antigen or other approaches: i) TERT can immortalize cells without transformation or tumor formation [18]; and ii) cells immortalized by TERT not only proliferate but also differentiate or undergo maturation, whereas immortalized cells using SV40 large T antigen might lead to only proliferation without differentiation or maturation. We demonstrated that hTERT was stably expressed at both transcriptional and translational levels in human Sertoli cell line.
Significantly, human Sertoli cell line, namely hS1 cells, assumed the characteristics of primary human Sertoli cells. Firstly, this cell line had morphological features similar to primary human Sertoli cells when they were freshly separated by FACS and in culture. Secondly, human Sertoli cell line shared phenotypic characteristics with primary human Sertoli cells, since a number of markers for Sertoli cells, including GDNF, FGF2, BMP4, SCF, SOX9, GATA4, WT1, BMP6, and VIM, but not 3β-HSD, SMA and VASA (markers for Leydig cells, peritubular myoid cell and germ cells, respectively), were expressed in these cells. As a key component of the niche, Sertoli cells facilitate spermatogenesis by secreting a variety of growth factors and cytokines, e.g., GDNF, FGF2, BMP4, and SCF. All of these growth factors were detected in the human Sertoli cell line. It has been reported that GDNF and FGF2, produced by Sertoli cells, can promote the proliferation of spermatogonial stem cells from both humans and rodents [19–23]. GDNF, a member of transforming growth factor (TGF-β) super-family, is indispensable for the self-renewal of spermatogonial stem cells both in vitro and in vivo [22, 24]. BMP4, another member of the TGF-β superfamily, is secreted by Sertoli cells, and it stimulates the differentiation of spermatogonial stem cells by means of inducing KIT expression in the undifferentiated spermatogonia [25]. We have recently demonstrated that BMP4 regulates the proliferation of human Sertoli cells via Smad1/5 and ID2/3 pathway through an autocrine manner [26]. SCF, also produced by Sertoli cells, stimulates the proliferation and differentiation of type A spermatogonia through SCF/KIT pathway [14, 27]. SOX9 (sex-determining region Y box9 gene), GATA4 [28, 29] and WT1 [30] are essential for testis development. SOX9 can promote the maturity of Sertoli cells and the maintenance of spermatogenesis [31–33]. It has been reported that abortion of WT1 leads to gonadal agenesis owing to the failure of genital ridge development [34]. Further studies unveil that Sertoli cell could be reprogramed into Leydig cell by deleting WT1. On the contrary, Leydig cell could express Sertoli cell markers by means of overexpressing WT1 [35]. Similar to WT1, deletion of Gata4 in Sertoli cells results in complete loss of germ cells [36]. In addition, the connection between Sertoli cells and germ cells depend on VIM to a large extent, and exposure to dibutyl phthalate could result in aberrant vimentin cytoskeleton and abnormal spermatogenesis [37, 38]. Notably, the expression of GDNF, SCF, BMP4, SOX9, WT1 and VIM proteins in human Sertoli cell line was comparable to primary human Sertoli cells, suggesting that human Sertoli cell line might have similar function with primary human Sertoli cells in regards to the regulation of human male germ cells.
AR and FSHR, mature markers for adult Sertoli cells, are the two vital receptors expressed exclusively in Sertoli cells within seminiferous tubules, by which follicular stimulating hormone (FSH) and testosterone control the spermatogenesis [13, 39, 40]. AR-knockout mice are infertile and display abdominally located testes and developmental arrest of spermatocytes, round and elongated spermatids [6, 41, 42], implicating that AR-deficient Sertoli cells block spermatogenesis by prohibiting Sertoli cell maturation. Significantly, we found that the levels of FSHR and AR were much higher in human Sertoli cell line than primary human Sertoli cells, reflecting that human Sertoli cell line has some advantages over primary human Sertoli cells due to its more maturation for the usage in basic studies and clinical applications. In addition, the transcripts of BMP4, BMP6, and FGF2 were higher in human Sertoli cell line than primary human Sertoli cells due to the overexpression of hTERT, which could be helpful for regulation of human spermatogenesis.
Human primary Sertoli cells are rare owing to the limited availability of human testis and the phenotype changes of Sertoli cells occur in culture with passages. It is worth noting that our human Sertoli cell line immortalized by hTERT has been cultivated in vitro for over 30 passages with no morphological or phenotypic changes. Furthermore, this human Sertoli cell line has great safety. It has been reported that immortalized cells exhibit ether structural or numerical chromosomal malformation by transducing SV40 or papillomaviruses E6 and E7 [43, 44]. We found that 90% of this human Sertoli cell line had normal karyotype. Additionally, neither Y chromosome microdeletions nor tumor formation was seen in human Sertoli cell line, implying vital applications of these cells in translational medicine.
MATERIALS AND METHODS
Acquisition of testicular biopsies from OA patients with normal spermatogenesis
Testicular biopsies were obtained from OA patients who underwent micro-dissection and testicular sperm extraction at Ren Ji Hospital, affiliated to Shanghai Jiao Tong University School of Medicine. Spermatogenesis was normal in all OA patients as evaluated by histology analysis of testis sections using hematoxylin and eosin (H&E) staining. The azoospermia of these patients were caused by inflammation or vasoligation rather than congenital absence of the vas deferens or other diseases. All experiments were conducted in accordance with relevant guidelines and regulations of the Institutional Ethical Review Committee of Ren Ji Hospital (license number of ethics statement: 2012–01), affiliated to Shanghai Jiao Tong University School of Medicine, and the written informed consents for testicular biopsies for research only were obtained by the patients.
Isolation of human Sertoli cells from OA patients
Testicular tissues from OA patients were washed three times aseptically in Dulbecco modified Eagle medium Nutrient Mixture F-12 (DMEM/F-12) (Gibco) containing antibiotics with penicillin and streptomycin (Gibco). Seminiferous tubules were separated from testis biopsies by the first enzymatic digestion containing 2 mg/ml collagenase IV (Gibco) and 1 μg/ml DNase I (Roche) in 34°C water bath for 15 min. Sertoli cells and human male germ cells were obtained from seminiferous tubules using a second enzymatic digestion comprising 4 mg/ml collagenase IV, 2.5 mg/ml hyaluronidase (Sigma), 2 mg/ml trypsin (Sigma) and μg/ml DNase I and followed by differential plating pursuant to the protocol as previously described [45]. Cell suspension was seeded into polystyrene-treated dishes (Corning) in DMEM/F-12 supplemented with 10% FBS (Gibco) and incubated at 34°C in 5% CO2 for 1 day. In this case, suspending male germ cells and other types of cells were removed, and the adherent cells were Sertoli cells.
Immortalization of human Sertoli cells
The expression vector, namely Lv-EF1A-hTERT-IRES-EGFP (Figure 1B), was purchased from Sidansai Biotechnology CO. LTD (Shanghai, China). Lenti-virus was packed by co-transfecting three plasmids, including VSVG, Δ8.9 and plasmid carrying gene hTERT, into 293T cells with lipofectamine® 3000 (Invitrogen) according to the manufacturer’s instruction. The titer was about 106 TU/ml by infecting 293T cells with the packed lenti-virus. Human Sertoli cells were seeded in 6-well plates at 60% confluence, and the medium was removed and replenished with fresh DMEM/F-12 supplemented with 10% FBS containing 100 μl lenti-virus and 8 μg/ml polybrene (Sigma), and the cells were incubated at 37°C in 5% CO2 overnight. Twenty-four hours later, medium was changed with fresh DMEM/F-12 and 10% FBS, and the expression of EGFP was detected under a fluorescence microscope (Nikon) at 48 hours after infection. The immortalized human Sertoli cells, namely the EGFP-positive cells, were sorted by fluorescence-activated cell sorting (FACS) and cultured with DMEM/F-12 supplemented with 10% FBS in 5% CO2.
RNA extraction, reverse transcription-polymerase chain reaction (RT-PCR) and real-time PCR
Total RNA was extracted from the immortalized human Sertoli cells at passage 2 (P2) using Trizol (Invitrogen) followed by the treatment of DNase I to remove potential contamination of genomic DNA. The cDNAs were synthesized using the RevertAid First Strand cDNA Synthesis Kit (Thermo Scientific) pursuant to the manufacturer’s instruction. The primers of the chosen genes, including GATA4 (GATA binding protein 4), WT1, SOX9 (Sex Determining Region Y-Box9), GDNF, BMP4, SCF, FSHR (follicle-stimulating hormone receptor), AR (androgen receptor), 3β-HSD (hydroxy-delta-5-steroid dehydrogenase, 3 beta), SMA (alpha-smooth muscle actin), VASA (DEAD-box helicase 4, also called DDX4) and GAPDH (glyceraldehyde-3-phosphate dehydrogenase), were designed and listed in Table 4. The PCR reactions started at 94 °C for 5 min and were performed as follows: denaturation at 95°C for 30 sec, annealing at 55–60°C for 30 sec as listed in Table 4, and elongation at 72°C for 30 sec, for 35cycles. After amplification, the samples were incubated for another 5 min at 72°C. PCR products were separated by electrophoresis in 1.5% agarose gels and visualized with ethidium bromide. Images were recorded and band intensities were analyzed using chemiluminescence (ChemiDocTM XRS+, Bio-Rad). Total RNA without RT (RT-) but with PCR using GAPDH primers served as negative controls.
Table 4: Primer sequences of genes used for RT-PCR
Genes |
Primer sequences |
Products size (bp) |
Tm (°C) |
|
|---|---|---|---|---|
GATA4 |
Forward |
GCCTCCTCTGCCTGGTAAT |
120 |
60 |
Reverse |
CAGTCCCATCAGCGTGTAAA |
|||
WT1 |
Forward |
TGACTCTCCACTCCTCCTCAC |
115 |
60 |
Reverse |
ACCAACTCTTCCAGGCACAC |
|||
SOX9 |
Forward |
AGGTGCTCAAAGGCTACGACTG |
322 |
60 |
Reverse |
TGCCCGTTCTTCACCGACT |
|||
GDNF |
Forward |
GAAGTTATGGGATGTCGTG |
419 |
58 |
Reverse |
TCAGTTCCTCCTTGGTTTC |
|||
SCF |
Forward |
GTCATTGTTGGATAAGCGAGAT |
457 |
60 |
Reverse |
ATGGCTGCCCAGTGTAGG |
|||
BMP4 |
Forward |
TTTGTTCAAGATTGGCTGTC |
324 |
60 |
Reverse |
AGATCCCGCATGTAGTCC |
|||
AR |
Forward |
CCTTCACCAATGTCAACTCC |
198 |
60 |
Reverse |
CCACTGGAATAATGCTGAAGAG |
|||
BMP6 |
Forward |
AGCAATCTGTGGGTTGTGACT |
228 |
60 |
Reverse |
GGTAGAGCGATTACGACTCTGTT |
|||
3β-HSD |
Forward |
GCCGATTCCTTTCTGCTAGTAT |
378 |
58 |
Reverse |
TGACTATGTGGCGGTTGAAG |
|||
SMA |
Forward |
GATCTGGCACCACTCTTTCTAC |
479 |
58 |
Reverse |
CAGGCAACTCGTAACTCTTCTC |
|||
VASA |
Forward |
GCAGAAGGAGGAGAAAGTAGTG |
289 |
60 |
Reverse |
CTCGTCCTGCAAGTATGATAGG |
|||
hTERT |
Forward |
GGTGAACTTCCCTGTAGAAGAC |
374 |
60 |
Reverse |
GGTTCTTCCAAACTTGCTGATG |
|||
GAPDH |
Forward |
AATCCCATCACCATCTTCC |
382 |
58 |
Reverse |
CATCACGCCACAGTTTCC |
Real-time PCR were performed using Power SYBR® Green PCR Master Mix (Applied Biosystems) and a StepOnePlus Real-Time PCR System (Applied Biosystems) according to the manufacture’s instruction. To quantify the PCR products, the 2-ΔΔCt method was used in terms of the protocol as described previously [46]. The threshold of cycle values was normalized against threshold values of human housekeeping gene GAPDH. The primer pairs of chosen genes were listed in Table 5.
Table 5: The primer sequences of genes used for real-time PCR
Genes |
Primer sequences |
Tm (°C) |
|
|---|---|---|---|
BMP6 |
Forward |
AGCAATCTGTGGGTTGTGACT |
60 |
Reverse |
GGTAGAGCGATTACGACTCTGTT |
||
BMP4 |
Forward |
TAGCAAGAGTGCCGTCATTCC |
60 |
Reverse |
GCGCTCAGGATACTCAAGACC |
||
SOX9 |
Forward |
AGCGAACGCACATCAAGAC |
60 |
Reverse |
CTGTAGGCGATCTGTTGGGG |
||
GATA1 |
Forward |
TTGTCAGTAAACGGGCAGGTA |
60 |
Reverse |
CTTGCGGTTTCGAGTCTGAAT |
||
FGF2 |
Forward |
AGTGTGTGCTAACCGTTACCT |
60 |
Reverse |
ACTGCCCAGTTCGTTTCAGTG |
||
GATA4 |
Forward |
GCCTCCTCTGCCTGGTAAT |
60 |
Reverse |
CAGTCCCATCAGCGTGTAAA |
||
WT1 |
Forward |
TGACTCTCCACTCCTCCTCAC |
60 |
Reverse |
ACCAACTCTTCCAGGCACAC |
||
LIF |
Forward |
CCAACGTGACGGACTTCCC |
60 |
Reverse |
TACACGACTATGCGGTACAGC |
||
GDNF |
Forward |
GGCAGTGCTTCCTAGAAGAGA |
60 |
Reverse |
AAGACACAACCCCGGTTTTTG |
||
GAPDH |
Forward |
CAGGAGGCATTGCTGATGAT |
60 |
Reverse |
GAAGGCTGGGGCTCATTT |
Immunocytochemistry of the immortalized human Sertoli cells
For immunocytochemical staining, the immortalized human Sertoli cells were fixed with 4% paraformaldehyde (PFA) for 15 min, washed three times with phosphate-buffered saline (PBS) for 5 min each and permeabilized in 0.5% triton-X 100 (Sigma) for 20 min. After washing with PBS, the cells were blocked in 3% serum or BSA for 60 min and followed by incubation with primary antibodies, including anti-SOX9 (Millipore), anti-WT1 (Santa Cruz), anti-OCLN (Abcam), anti-VIM (vimentin, Santa Cruz), anti-SCF (Sigma), anti-BMP4 (Abcam), anti-GDNF (Abcam), anti-3β-HSD (Santa Cruz), anti-VASA(Santa Cruz), and anti-Ki-67 (BD Biosciences), overnight at 4°C. Replacement of primary antibodies with isotype IgGs (Santa Cruz) served as negative controls. After washing three times in PBS for 10 min each, the cells were incubated with the secondary antibody (Sigma) conjugated with rhodamine at a 1:200 dilution for 1 hour at room temperature. DAPI (4, 6-diamidino-2-phenylindole, Sigma) was used to counterstain the nuclei, and images were captured with a fluorescence microscope (Nikon).
Western blots
The immortalized human Sertoli cells and human primary Sertoli were lysed with RIPA buffer (Santa Cruz) for 30 min on ice. Cell lysates were cleared by centrifugation at 12,000 g for 10 min at 4°C, and the concentrations of total proteins were measured by BCA kit (Dingguo Company, China). Thirty micrograms of cell lysates from each sample were used for SDS-PAGE, and Western blots were performed according to the protocol as described previously [24]. The chosen antibody included anti-hTERT (Santa Cruz), anti-FSHR (Abcam), anti-AR (Santa Cruz), anti-GDNF, anti-SCF, anti-BMP4, anti-WT1, anti-SOX9, anti-PCNA (proliferating cell nuclear antigen, Santa Cruz), anti-3β-HSD, anti-VASA, anti-SMA, and anti-ACTB (ProteintechTM). Replacement of primary antibodies with PBS served as negative controls (NC). After extensive washes in TBST, the images were detected by chemiluminescence (ChemiDocTM XRS+, Bio-Rad).
Microarray analysis
Total RNA was extracted from immortalized human Sertoli cells at passage 15 (P15) and primary Sertoli cells using Trizol reagent (Invitrogen). Total RNA was quantified by the NanoDrop ND-2000 (Thermo Scientific) and the RNA integrity was assessed using Agilent Bioanalyzer 2100 (Agilent Technologies). MRNA microarrays were applied on Agient Human microarray 8*60 array (Ouyi, Shanghai, China), and the results were confirmed by real-time PCR. Briefly, total RNA were transcribed to double strand cDNA, synthesized into cRNA and labeled with Cyanine-3-CTP. The labeled cRNAs were hybridized onto the microarrays. After extensive washes, the arrays were scanned by the Agilent Scanner G2505C (Agilent Technologies). Feature Extraction software (version10.7.1.1, Agilent Technologies) was used to analyze array images to get raw data. Genespring software (version13.1, Agilent Technologies) was employed to perform the basic analysis with the raw data. The significantly differentially expressed genes were selected according to the criteria: P value < 0.05, Fold change ≥ 2.
CCK-8 assays
Human immortalized Sertoli cells at passage 6 (P6) and primary human Sertoli cells were seeded in 96-well plates at a concentration of 2,000 or 1,000 cells per well in 100 μl DMEM/F-12 medium containing 0.5%, 1%, 2%, 5%, 10% and 15% of FBS respectively, and incubated at 37 °C in 5% CO2 for 7 days. Cellular proliferation was measured every 24 hours according to the protocol of the Cell Counting Kit-8 (CCK-8) Kit (Dojin Laboratories).
Karyotyping assays
Chromosomal karyotype analysis of the exponentially growing immortalized human Sertoli cells at passage 15 (P15) was conducted pursuant to the procedure described previously [12]. In brief, cells were inhibited with 5 μg/ml colchicine for 3 hours, followed by hypotonic treatment with 0.075 M KCl solution, fixed with 3:1 methanol-glacial acetic acid, and dropped onto chilled slides. Cells were stained with Giemsa and counted under a microscope. The recommendation of the International System for Human Cytogenetic Nomenclature was applied to analyze the karyotypes.
Multiplex real-time PCR analysis
Multiplex real-time PCR (Tellgen Corporation, Shanghai) was utilized to assess whether Y microdeletions existed in the immortalized human Sertoli cells at passage 20 (P20) according to the manufacturer’s instructions. Three different regions, AZFa, AZFb and AZFc, were analyzed with six specific sequence-tagged site (STS) markers, including sY254, sY127, sY86, sY134, sY84 and sY255. Zinc finger protein, X-linked (ZFX)/zinc finger protein, Y-linked (ZFY), and sex determining region Y (SRY), were used as internal controls. DNA from normal human blood served as positive controls and water without DNA with all primers was used as negative controls.
Tumor formation of the immortalized human Sertoli cells by xenotransplantation assays
Thirty male nude mice (nu/nu BALB-c) at 6-week-old were used for in vivo tumor-formation potential of the immortalized human Sertoli cells at passage 20 (P20). The mice were housed in constant laboratory conditions of a 12-hour light, 12-hour dark cycle and pathogen-free conditions and fed with water and food ad libitum. All mice were treated in accordance with the animal care and use guidelines of Ren Ji Hospital animal care and ethics review committee. For xenograft study, the mice were divided into three groups. In the first group, each nu/nu nude mouse was injected subcutaneously into the left and right posterior axillary fossa without cells in a total volume of 100 μl DMEM/F-12 and Matrigel (1:1). In the latter two groups, each nu/nu nude mouse was injected subcutaneously with 2 × 106 immortalized human Sertoli cells or 2 × 106 human prostate cancer cells PC3 suspended with 100 μl DMEM/F-12 and Matrigel (1:1). Two weeks later, the tissues from mouse transplanted sites were fixed in paraformaldehyde overnight, embedded in paraffin, and sectioned at 5 μm thickness. The sections were stained with H&E and observed under a microscope (Nikon).
Statistical analysis
All data were presented as mean ± SEM from at least three independent experiments and analyzed by Student’s t-test or one-way ANOVA with the appropriate post-hoc tests (Dunnet’s test or Turkey’s multiple comparison) using Prism (version 5, GraphPad Software), and p < 0.05 was considered statistically significant.
SUMMARY
In conclusion, we have established the first human Sertoli cell line with the morphological and phenotypic attributes of primary human Sertoli cells. This human cell line can facilitate our understanding the biology of human Sertoli cells. Notably, this human Sertoli cell line has almost normal karyotype, excludes Y chromosome microdeletions and doesn’t form tumor. Moreover, the unlimited proliferative capacity of this cell line ensures us to obtain adequate source of human Sertoli cells for uncovering their roles in regulating the complex human spermatogenesis and the treatments for male infertility as well as other human diseases.
CONFLICTS OF INTEREST
The authors declare that there was no conflicts of interest that could be perceived as prejudicing the impartiality of the review.
FUNDING
This work was supported by grants from National Nature Science Foundation of China (31230048, 31671550, 31171422, 31401250) and Chinese Ministry of Science and Technology (2016YFC1000606, 2014CB943101), The Program for Professor of Special Appointment (Eastern Scholar) at Shanghai Institutions of Higher Learning (2012.53), Shanghai Municipal Education Commission-Gaofeng Clinical Medicine Grant Support (20152511), Shanghai Hospital Development Center (SHDC12015122), and a key grant from the Science and Technology Commission of Shanghai Municipality (12JC1405900).
REFERENCES
1. Hai Y, Hou J, Liu Y, Yang H, Li Z, He Z. The roles and regulation of Sertoli cells in fate determinations of spermatogonial stem cells and spermatogenesis. Semin Cell Dev Biol. 2014; 29:66–75.
2. Nah WH, Lee JE, Park HJ, Park NC, Gye MC. Claudin-11 expression increased in spermatogenic defect in human testes. Fertil Steril. 2011; 95:385–388.
3. Lui WY, Mruk D, Lee WM, Cheng CY. Sertoli cell tight junction dynamics: their regulation during spermatogenesis. Biol Reprod. 2003; 68:1087–1097.
4. Sharpe RM, McKinnell C, Kivlin C, Fisher JS. Proliferation and functional maturation of Sertoli cells, and their relevance to disorders of testis function in adulthood. Reproduction. 2003; 125:769–784.
5. Brehm R, Zeiler M, Ruttinger C, Herde K, Kibschull M, Winterhager E, Willecke K, Guillou F, Lecureuil C, Steger K, Konrad L, Biermann K, Failing K, et al. A sertoli cell-specific knockout of connexin43 prevents initiation of spermatogenesis. Am J Pathol. 2007; 171:19–31.
6. Chang C, Chen YT, Yeh SD, Xu Q, Wang RS, Guillou F, Lardy H, Yeh S. Infertility with defective spermatogenesis and hypotestosteronemia in male mice lacking the androgen receptor in Sertoli cells. P Natl Acad Sci USA. 2004; 101:6876–6881.
7. Kaur G, Thompson LA, Pasham M, Tessanne K, Long CR, Dufour JM. Sustained expression of insulin by a genetically engineered sertoli cell line after allotransplantation in diabetic BALB/c mice. Biol Reprod. 2014; 90:109.
8. Sun H, Zhang G, Dong F, Wang F, Cao W. Reprogramming sertoli cells into pluripotent stem cells. Cell Reprogram. 2014; 16:196–205.
9. Wang X, Qin J, Zhao RC, Zenke M. Reduced immunogenicity of induced pluripotent stem cells derived from Sertoli cells. Plos One. 2014; 9:e106110.
10. Sheng C, Zheng Q, Wu J, Xu Z, Wang L, Li W, Zhang H, Zhao XY, Liu L, Wang Z, Guo C, Wu HJ, Liu Z, et al. Direct reprogramming of Sertoli cells into multipotent neural stem cells by defined factors. Cell Res. 2012; 22:208–218.
11. Zhang L, Chen M, Wen Q, Li Y, Wang Y, Qin Y, Cui X, Yang L, Huff V, Gao F. Reprogramming of Sertoli cells to fetal-like Leydig cells by Wt1 ablation. P Natl Acad Sci USA. 2015; 112:4003–4008.
12. Hou J, Niu M, Liu L, Zhu Z, Wang X, Sun M, Yuan Q, Yang S, Zeng W, Liu Y, Li Z, He Z. Establishment and Characterization of Human Germline Stem Cell Line with Unlimited Proliferation Potentials and no Tumor Formation. Sci Rep. 2015; 5:16922.
13. Chuang CK, Lee KH, Fan CT, Su YS. FSH-sensitive murine sertoli cell lines immortalized by human telomerase gene hTERT express the androgen receptor in response to TNF-alpha stimulation. Bioscience Rep. 2007; 27:403–411.
14. Schlegel PG, Wolfl M, Schick J, Winkler B, Eyrich M. Transient loss of consciousness in pediatric recipients of dimethylsulfoxide (DMSO)-cryopreserved peripheral blood stem cells independent of morphine co-medication. Haematologica. 2009; 94:1473–1475.
15. Nakagawa H, Nishijo M, Morikawa Y, Miura K, Tawara K, Kuriwaki J, Kido T, Ikawa A, Kobayashi E, Nogawa K. Urinary cadmium and mortality among inhabitants of a cadmium-polluted area in Japan. Environ Res. 2006; 100:323–329.
16. Tabuchi Y, Takahashi R, Ueda M, Obinata M. Development of a conditionally immortalized testicular Sertoli cell line RTS3–3 from adult transgenic rats harboring temperature-sensitive simian virus 40 large T-antigen gene. Cell Struct Funct. 2003; 28:87–95.
17. Moroi S, Saitou M, Fujimoto K, Sakakibara A, Furuse M, Yoshida O, Tsukita S. Occludin is concentrated at tight junctions of mouse/rat but not human/guinea pig Sertoli cells in testes. Am J Physiol. 1998; 274:C1708–1717.
18. Morales CP, Holt SE, Ouellette M, Kaur KJ, Yan Y, Wilson KS, White MA, Wright WE, Shay JW. Absence of cancer-associated changes in human fibroblasts immortalized with telomerase. Nat Genet. 1999; 21:115–118.
19. Sadri-Ardekani H, Mizrak SC, van Daalen SK, Korver CM, Roepers-Gajadien HL, Koruji M, Hovingh S, de Reijke TM, de la Rosette JJ, van der Veen F, de Rooij DG, Repping S and van Pelt AM. Propagation of human spermatogonial stem cells in vitro. Jama. 2009; 302:2127–2134.
20. Kanatsu-Shinohara M, Miki H, Inoue K, Ogonuki N, Toyokuni S, Ogura A, Shinohara T. Long-term culture of mouse male germline stem cells under serum-or feeder-free conditions. Biol Reprod. 2005; 72:985–991.
21. Kubota H, Avarbock MR, Brinster RL. Growth factors essential for self-renewal and expansion of mouse spermatogonial stem cells. P Natl Acad Sci USA. 2004; 101:16489–16494.
22. Krill D, Shuman M, Thompson MT, Becich MJ, Strom SC. A simple method for the isolation and culture of epithelial and stromal cells from benign and neoplastic prostates. Urology. 1997; 49:981–988.
23. Fink C, Weigel R, Hembes T, Lauke-Wettwer H, Kliesch S, Bergmann M, Brehm RH. Altered expression of ZO-1 and ZO-2 in Sertoli cells and loss of blood-testis barrier integrity in testicular carcinoma in situ. Neoplasia. 2006; 8:1019–1027.
24. He Z, Jiang J, Kokkinaki M, Golestaneh N, Hofmann MC, Dym M. Gdnf upregulates c-Fos transcription via the Ras/Erk1/2 pathway to promote mouse spermatogonial stem cell proliferation. Stem Cells. 2008; 26:266–278.
25. Pellegrini M, Grimaldi P, Rossi P, Geremia R, Dolci S. Developmental expression of BMP4/ALK3/SMAD5 signaling pathway in the mouse testis: a potential role of BMP4 in spermatogonia differentiation. J Cell Sci. 2003; 116:3363–3372.
26. Hai Y, Sun M, Niu M, Yuan Q, Guo Y, Li Z, He Z. BMP4 promotes human Sertoli cell proliferation via Smad1/5 and ID2/3 pathway and its abnormality is associated with azoospermia. Discov Med. 2015; 19:311–325.
27. Ohta H, Yomogida K, Dohmae K, Nishimune Y. Regulation of proliferation and differentiation in spermatogonial stem cells: the role of c-kit and its ligand SCF. Development. 2000; 127:2125–2131.
28. Hu YC, Okumura LM, Page DC. Gata4 is required for formation of the genital ridge in mice. Plos Genet. 2013; 9:e1003629.
29. Viger RS, Guittot SM, Anttonen M, Wilson DB, Heikinheimo M. Role of the GATA family of transcription factors in endocrine development, function, and disease. Mol Endocrinol. 2008; 22:781–798.
30. Pelletier J, Bruening W, Kashtan CE, Mauer SM, Manivel JC, Striegel JE, Houghton DC, Junien C, Habib R, Fouser L and et al. Germline mutations in the Wilms’ tumor suppressor gene are associated with abnormal urogenital development in Denys-Drash syndrome. Cell. 1991; 67:437–447.
31. Morais da Silva S, Hacker A, Harley V, Goodfellow P, Swain A, Lovell-Badge R. Sox9 expression during gonadal development implies a conserved role for the gene in testis differentiation in mammals and birds. Nat Genet. 1996; 14:62–68.
32. de Santa Barbara P, Moniot B, Poulat F, Berta P. Expression and subcellular localization of SF-1, SOX9, WT1, and AMH proteins during early human testicular development. Dev Dynam. 2000; 217:293–298.
33. Banco B, Palmieri C, Sironi G, Fantinato E, Veronesi MC, Groppetti D, Giudice C, Martignoni B, Grieco V. Immunohistochemical expression of SOX9 protein in immature, mature, and neoplastic canine Sertoli cells. Theriogenology. 2016; 85:1408–1414.e1401.
34. Kreidberg JA, Sariola H, Loring JM, Maeda M, Pelletier J, Housman D, Jaenisch R. WT-1 is required for early kidney development. Cell. 1993; 74:679–691.
35. Zhang L, Chen M, Wen Q, Li Y, Wang Y, Wang Y, Qin Y, Cui X, Yang L, Huff V, Gao F. Reprogramming of Sertoli cells to fetal-like Leydig cells by Wt1 ablation. P Natl Acad Sci USA. 2015; 112:4003–4008.
36. Chen SR, Tang JX, Cheng JM, Li J, Jin C, Li XY, Deng SL, Zhang Y, Wang XX, Liu YX. Loss of Gata4 in Sertoli cells impairs the spermatogonial stem cell niche and causes germ cell exhaustion by attenuating chemokine signaling. Oncotarget. 2015; 6:37012–37027. doi: 10.18632/oncotarget.6115.
37. Kleymenova E, Swanson C, Boekelheide K, Gaido KW. Exposure in utero to di(n-butyl) phthalate alters the vimentin cytoskeleton of fetal rat Sertoli cells and disrupts Sertoli cell-gonocyte contact. Biol Reprod. 2005; 73:482–490.
38. Zhang X, Wang X, Liu T, Mo M, Ao L, Liu J, Cao J, Cui Z. Smad2/3 Upregulates the Expression of Vimentin and Affects Its Distribution in DBP-Exposed Sertoli Cells. Ppar Res. 2015; 2015:489314.
39. Wang RS, Yeh S, Chen LM, Lin HY, Zhang C, Ni J, Wu CC, di Sant’Agnese PA, deMesy-Bentley KL, Tzeng CR, Chang C. Androgen receptor in sertoli cell is essential for germ cell nursery and junctional complex formation in mouse testes. Endocrinology. 2006; 147:5624–5633.
40. Ohta T, Miyake H, Miura C, Kamei H, Aida K, Miura T. Follicle-stimulating hormone induces spermatogenesis mediated by androgen production in Japanese eel, Anguilla japonica. Biol Reprod. 2007; 77:970–977.
41. Abel MH, Baker PJ, Charlton HM, Monteiro A, Verhoeven G, De Gendt K, Guillou F, O’Shaughnessy PJ. Spermatogenesis and sertoli cell activity in mice lacking sertoli cell receptors for follicle-stimulating hormone and androgen. Endocrinology. 2008; 149:3279–3285.
42. De Gendt K, Swinnen JV, Saunders PT, Schoonjans L, Dewerchin M, Devos A, Tan K, Atanassova N, Claessens F, Lecureuil C, Heyns W, Carmeliet P, Guillou F, et al. A Sertoli cell-selective knockout of the androgen receptor causes spermatogenic arrest in meiosis. Proc Natl Acad Sci USA. 2004; 101:1327–1332.
43. Balducci L, Blasi A, Saldarelli M, Soleti A, Pessina A, Bonomi A, Cocce V, Dossena M, Tosetti V, Ceserani V, Navone SE, Falchetti ML, Parati EA, et al. Immortalization of human adipose-derived stromal cells: production of cell lines with high growth rate, mesenchymal marker expression and capability to secrete high levels of angiogenic factors. Stem Cell Res Ther. 2014; 5:63.
44. Akimov SS, Ramezani A, Hawley TS, Hawley RG. Bypass of senescence, immortalization, and transformation of human hematopoietic progenitor cells. Stem Cells. 2005; 23:1423–1433.
45. He Z, Kokkinaki M, Jiang J, Dobrinski I, Dym M. Isolation, characterization, and culture of human spermatogonia. Biol Reprod. 2010; 82:363–372.
46. He Z, Jiang J, Hofmann MC, Dym M. Gfra1 silencing in mouse spermatogonial stem cells results in their differentiation via the inactivation of RET tyrosine kinase. Biol Reprod. 2007; 77:723–733.

