INTRODUCTION
Follicular dendritic cell sarcoma (FDC-S) is the neoplastic proliferation of follicular dendritic cells (FDC) [1–2]. FDC belong to the stromal compartment of secondary lymphoid organs [3] and share crucial interaction capacities with the haematopoietic counterpart; both quality and effectiveness of immune responses depend from their integrity [4–6]. Therefore, FDC-S is included in the WHO classification of Haematopoietic tumors [1] and not in the WHO classification of Tumors of Soft Tissue and Bone [7] despite of its mesenchymal origin. The complex mechanisms mediating these functions are still largely unknown but new insights have been recently achieved in experimental models [4, 8–10].
FDC-S occurs in adults (median age 50 years), with no gender predilection, in both nodal and extranodal sites [11–15]. Its behavior is unpredictable; in a recent large review study, local recurrence and distant metastasis were reported to be equal to 28.1% and 27.2%, respectively [12].
On histology, FDC-S is characterized by extreme variability in cell composition and growth patterns, diagnosis is supported by the use of a panel of FDC-associated immunohistochemical markers combined with others, specific for epithelial neoplasms, soft tissue tumors and melanoma, to be used for exclusion [16]. Some differential diagnoses can be extremely challenging, particularly those entities typically prone to antigen loss (i.e. melanomas and undifferentiated carcinomas) the markers of which may be occasionally expressed by FDC-S (i.e. S100 and EMA) [16].
At present, the most widely used FDC-S markers are complement component receptor (CR) 1 (CD35), CR2 (CD21), Fc fragment of IgE receptor 2 (FCER2, CD23) and Clusterin. Recently, other proteins were identified including CXCL13 [17], D2-40 [18] and Claudin 4 [19]. Noteworthy, FDC-S frequently express a defective phenotype which may lead to misdiagnosis [12].
The molecular pathogenesis of FDC-S is largely unknown. Epidermal growth factor receptor signaling is known to support FDC-S proliferation [20–21]; mutations affecting the nuclear factor κ B pathway and cell cycle regulation were recently identified, by targeted next generation sequencing, in a subgroup of FDC-S [22]. Data on whole genome and whole transcriptome sequencing on this tumor are still lacking.
In order to explore the molecular and proteomic landscape of FDC-S we performed the first whole transcriptome sequencing analysis on two FDC-S cases. Pathway enrichment analysis was performed and new candidate immunohistochemical markers were identified and validated in a large cohort of FDC-S and controls.
RESULTS
Pathological features of FDC-S undergoing whole transcriptome sequencing
The clinical, morphological and phenotypical data of the two cases of FDC-S used for WTS are detailed in Table 1 and shown in Figure 1.
Table 1: Clinical, phenotypical and molecular data of FDC-S submitted for whole transcriptome sequencing
Case #1 |
Case #2 |
|||
|---|---|---|---|---|
Gender |
F |
M |
||
Age (years) |
71 |
76 |
||
Site |
Mediastinum |
Soft tissues of the thigh |
||
Maximum diameter |
5.9 cm |
NA |
||
Therapy and follow up |
Surgery and radiotherapy NED (15 months) |
NA |
||
FDC-S markers GENE_Protein |
protein expression by IHC |
gene transcript (RPKM) |
protein expression by IHC |
gene transcript (RPKM) |
CR2_CD21 |
– |
49.0621 |
+ |
940.95374 |
FCER2_CD23 |
+ |
3519.927 |
+ |
295.29196 |
CR1_CD35 |
– |
15.26415 |
– |
54.7856 |
CLU_Clusterin |
+ |
10482.2246 |
+ |
7388.229 |
PDPN_Podoplanin |
+ |
213.73975 |
+ |
103.46978 |
CXCL13_CXCL13 |
+ |
8698.03906 |
+ |
1487.44849 |
Expression values of the genes encoding for CD21, CD23, CXCL13, CD35, Clusterin and Podoplanin correlate with immunohistochemical expression of their protein.
F, female; FDC-S, follicular dendritic cell sarcoma; IHC, immunohistochemistry; M, male; NA, not available; NED, no evidence of disease; RPKM, kilobase per million mapped reads.
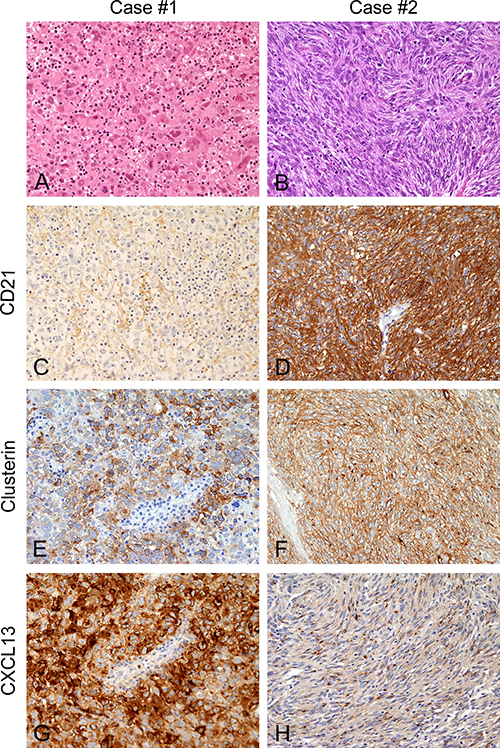
Figure 1: Morphology and phenotype of FDC-S cases submitted for transcriptome sequencing. In Case #1 (A, C, E and G) atypical cells were epithelioid, pleomorphic, intermingled by numerous, small lymphocytes (A, hematoxylin and eosin), they stained negative for CD21 (C) but intensely expressed Clusterin (E) and CXCL13 (G). Case #2 (B, D, F and H) showed a different morphology with spindle atypical cells, arranged in fascicles (B, hematoxylin and eosin), positive for CD21 (D), Clusterin (F) and CXCL13 (H), the latter with a punctuate Golgi dot pattern. Magnification: 200×.
The mediastinal biopsy of Case #1 showed highly pleomorphic cells, often multinucleated; with nuclear pseudo-inclusions and atypical mitoses. Reactive T and, less numerous, B- lymphocytes were intermingled within the tumor. Differently, in Case #2, tumor cells were bland looking, spindle, arranged in parallel fascicles; mitoses were few and never atypical, scattered mature lymphocytes were present. Tumor cell content was estimated on histological section to be above 40% in both cases. Their phenotype (reported in Table 1) showed expression of at least two FDC-specific markers.
Whole transcriptome of FDC-S
No fusion transcripts were detected by whole transcriptome sequencing excluding the presence of chromosome translocations with oncogenic significance.
Genes with reads per kilobase per million mapped reads (RPKM) higher than 1 were 17,956 in case #1 and 18,118 in case #2. IGK (RPKM = 32,171.7754) and FDCSP (RPKM = 11,593.3535) had the highest expression value, in case #1 and #2, respectively.
Gene ontology and functional annotation of both lists generated an equivalent output, despite the differences in tissue type and tumor morphology.
Annotation of gene list for biological processes suggested a metabolically active tumor given enrichment of “metabolic process” (GO:0008152) and “cellular process” (GO: 0009987). Among the cellular components actin-related cytoskeleton (GO: 0015629) resulted enriched, in line with the actin-dependent activity of FDC [23]. Overrepresentation analysis was performed with the Panther pathway annotation dataset separately, in the two cases. Among others, a statistically significant (p < 0.05) overrepresentation of “Ubiquitin Proteasome” (P00060) pathway was detected in both cases, in line with the recent evidence of NFκB pathway regulatory-factors loss of function in this tumor [22]. According to the histological evidence of a higher T-cell infiltrate, “T-cell activation” (P00053) was increased in the transcriptome of case #1 while “Integrin Signalling” (P00034) and “Epidermal growth factor receptor (EGFR) signalling” pathway (P00018) were overrepresented in case #2, consistent with previous observations of activated EGFR signalling in FDC-S. [20–21]. Fold changes and p-values are listed in Supplemental material, Supplementary Table 1.
Transcripts coding for established immunohistochemical FDC-S markers are among the top 10% of FDC-S transcriptome
Genes coding for FDC markers showed expression values higher than 55 RPKM and were among the “top 10%” genes of each transcriptome. Notably, CLU and CXCL13 were those with the highest RPKM; CLU had the second transcription value in both; CXCL13 was respectively at the 7th and 36th position (Supplementary Table 2). Accordingly, the transcriptome of the CD21-negative case #1 showed low expression values of the CD21-coding gene CR2 and CR1, gene coding for CD35, had low RPKM values in both transcriptomes, in line with the negativity of both cases for this protein by immunohistochemistry (Table 1, Figure 1).
FDCSP and SRGN have high expression values in FDC-S and are expressed by normal FDC
To identify additional markers for FDC-S we selected gene products with biological functions related to the germinal center reaction among the top 10% transcripts of both tumors.
Follicular dendritic cell secreted protein (FDCSP), also known as Chromosome 4 open reading frame 7(C4Orf7) and Serglycin (SRGN) were selected and antibodies against their proteins were tested on reactive tonsils. The cytoplasm of normal FDC, mostly located in the light zone of germinal centers of secondary lymphoid follicles, stained positive for both proteins with a perinuclear pattern (Figure 2A and 2B).
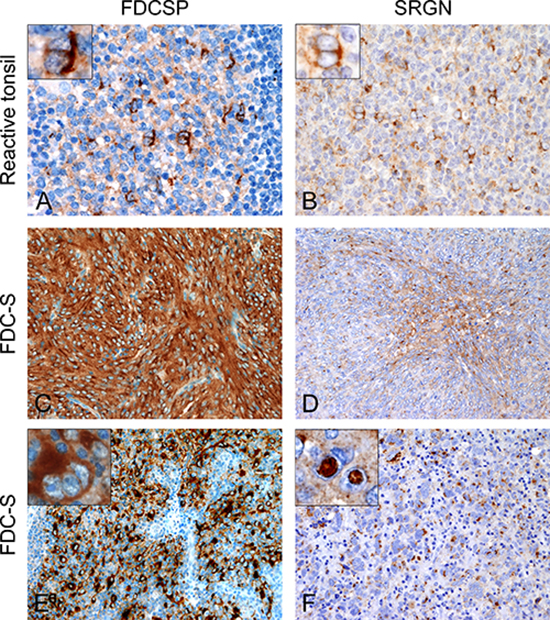
Figure 2: Expression patterns of FDCSP and SRGN on normal and neoplastic FDC. In the contest of reactive germinal centers normal FDC react for both FDCSP and SRGN protein with a perinuclear expression pattern (A, B). FDCSP (C, E) and SRGN (D, F) are axpressed also in the cytoplasm of neoplastic FDC either with diffuse and intense staining or with a more delicate punctuate, cytoplasmic dot. SRGN often stained nuclear pseudo-inclusions (F, inset). Magnification: (A and B) 400×. (C, D, E and G) 200×. Insets: 600×. Abbreviations: FDC-S, follicular dendritic cell sarcoma; FDCSP, FDC secreted protein; SRGN, Serglycin.
FDCSP and SRGN exhibit good sensitivity and specificity for FDC-S diagnosis
Immunohistochemistry on follicular dendritic cell sarcomas for FDCSP was positive in sixteen cases (16/22, 72.73%) with either an intense and diffuse cytoplasmic staining or a prevailing perinuclear positivity. Reaction for SRGN was detected in fifteen cases (15/22, 68.18%) and, similarly to CXCL13, displayed a strong Golgi staining pattern with or without diffuse cytoplasmic reaction; notably, nuclear pseudo inclusions were frequently SRGN-positive (Figure 2).
In order to test the specificity of FDCSP and SRGN in FDC-S diagnosis we stained 87 soft tissue tumors, 5 thymomas, 5 melanomas (Supplementary Table 3), 3 interdigitating dendritic cell sarcomas and 114 carcinomas [24]. FDCSP was expressed by two soft tissue tumors, one sinonasal haemangiopericytoma (SNHP-1) and one synovial sarcoma (SS-2); SRGN was negative in all (Supplementary Table 4). Notably, one case of interdigitating dendritic cell sarcoma expressed FDCSP in about 40% of neoplastic cells while it was negative for SRGN and other FDC markers. No reactivity for FDCSP and SRGN was detected on melanomas, thymomas or carcinomas.
CXCL13, CD21, CD35, FDCSP and SRGN are the best marker combination in FDC-S diagnosis
In order to compared the performance of FDCSP and SRGN with known FDC-S markers in the differential with soft tissue tumors, we tested CD21, CD23, CD35, CXCL13, Clusterin, Claudin 4 and Podoplanin in our groups of tumors and controls.
CXCL13, CD21 and Clusterin were the most retained markers in FDC-S with sensitivities of 90.91% for the first and 81.82% for the latter two (Table 2). CD35 and FDCSP were both expressed by 72.73% (16/22) of cases. The least sensitive were CD23 and Podoplanin, positive in 63.64% (14/22) of cases and Claudin 4, retained only by half of FDC-S (10/20). The highest specificity was observed for SRGN, CXCL13 and for complement and Fc receptors (CD21, CD35 and CD23): they were negative in all soft tissue tumors used as controls. Adversely, Clusterin showed a diffuse, undoubtful, expression in six cases of GIST (6/20, 30% not shown), all from extra-gastric sites. Podoplanin was expressed by two thirds of IMT cases (4/6, 66.67%, not shown), 25% (3/12) of SS, 15% (3/20) of GIST and 10.53% of SFT (2/19). Claudin 4 was expressed in one case of SS and one case of ALK-negative IMT.
Sensitivity and specificity of each marker were plotted on a ROC curve and the value of their area under the curve (AUC) was calculated (Figure 3, Table 2).
Table 2: Sensitivity, specificity, area under the curve (AUC) and significance p-value of FDC markers
Marker |
Sensitivity (+/FDC-S) |
Specificity (–/STT) |
AUC |
p-value |
|---|---|---|---|---|
CXCL13 |
90.91% (20/22) |
100.00% (72/72) |
0.955 |
< 0.0001 |
CD21 |
81.82% (18/22) |
100.00% (72/72) |
0.909 |
< 0.0001 |
CD35 |
72.73% (16/22) |
100.00% (72/72) |
0.864 |
< 0.0001 |
FDCSP |
72.73% (16/22) |
97.20% (70/72) |
0.850 |
< 0.0001 |
SRGN |
68.18% (15/22) |
100.00% (72/72) |
0.841 |
< 0.0001 |
Clusterin |
81.82% (18/22) |
81.94% (59/72) |
0.810 |
< 0.0001 |
CD23 |
63.64% (14/22) |
100.00% (72/72) |
0.800 |
< 0.0001 |
Podoplanin |
63.64% (14/22) |
83.33% (60/72) |
0.767 |
< 0.0001 |
Claudin4 |
50.00% (10/20) |
98.61% (71/72) |
0.736 |
< 0.0001 |
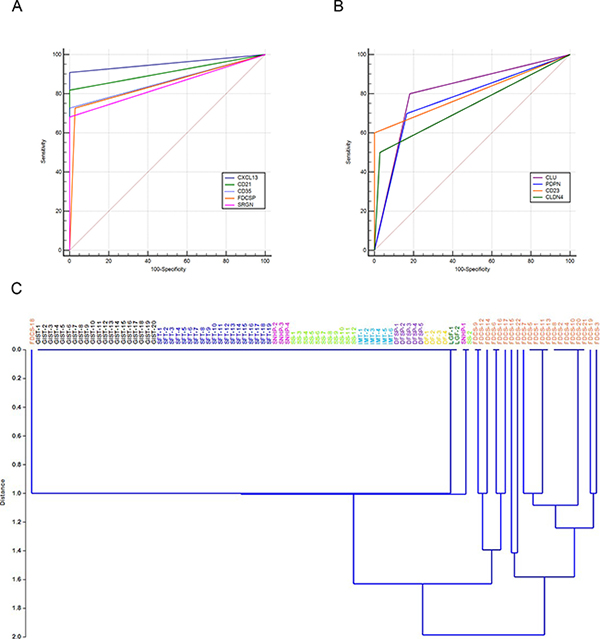
Figure 3: Representation of sensitivity and specificity of FDC markers by ROC curves and hierarchical clustering. The graphs represent the AUC of FDC markers: those with best sensitivity and specificity values, CXCL13, CD21, CD23, FDCSP and SRGN are plotted in (A) Clusterin, Podoplanin, CD23 and Claudin 4 are displayed in (B). Hierarchical clustering performed considering IHC expression scores of the five markers CXCL13, CD21, CD23, FDCSP and SRGN in 22 FDC-S and 72 soft tissue tumors show a neat separation of FDC-S from controls with the exception of a single FDC-S case (C). Abbreviations: DF, desmoid type fibromatosis; DFSP, dermatofibrosarcoma protuberans; FDC-S, follicular dendritic cell sarcoma; GIST gastrointestinal stromal tumor; IMT, inflammatory myofibroblastic tumor; LGF, low grade fibromyxoid sarcoma; SFT, solitary fibrous tumor; SNHP, sinonasal haemangiopericytoma; SS, synovial sarcoma.
Notably, CXCL13 showed the highest AUC, representing the marker with the best performance in the differential diagnosis of FDC-S with other tumors of mesenchymal origin. It was followed, in order, by CD21, CD35, FDCSP and SRGN (Figure 3A). Other widely used FDC-S markers such as Clusterin, CD23 and Podoplanin showed a lower AUC value (Figure 3B).
Evaluating the power of the combination of CXCL13, CD21, CD35, FDCSP and SRGN, by linear discriminant analysis (LDA), this marker combination showed a correct classification for 98.94% (93/94) of the cases tested, representing the best marker combination for the identification of FDC-S. When considering the same set of markers, FDC-S cases were nicely discriminated from the controls by principal component analysis (Supplementary Figure 1) with the exception of one case, FDC-S-18, by LDA (Figure 3C). The latter case developed as a retroperitoneal mass in a 73 years-old woman; neoplastic cells stained for Clusterin, Podoplanin and SRGN but were negative for all the other markers and may therefore represent an exceptional case.
DISCUSSION
Follicular dendritic cell sarcoma (FDC-S) is a mesenchymal tumor with morphological and phenotypical features of follicular dendritic cells [1, 4, 12, 16]. FDC-S have been reported in numerous sites of the body, both nodal and extranodal [12]. The morphological appearance of FDC-S includes different growth patterns: fascicular, storiform, whorled or diffuse; puzzle-like, nodular (i.e. folliculocentric) and angiomatoid architectures have also been described [1, 25]. Neoplastic FDC are spindle, oval or epithelioid and often show the same characteristics of normal FDC but nuclear pleomorphism may also occur [1]. Given its heterogeneous morphology, FDC-S diagnosis is supported by the use of immunohistochemistry, it demands the usage of a wide panel of antibodies, given its frequent loss of specific antigens and the possibility of focal reactivity for epithelial markers [16–17]. FDC-S in both nodal and extranodal sites may mimic a wide spectrum of neoplasms including metastatic melanomas, poorly differentiated carcinomas, both epithelioid and spindle cell sarcomas. The use of recently identified antibodies, such as STAT6 for solitary fibrous tumor and TLE1 for synovial sarcoma, are helpful in some settings. However, not all entities have highly specific and retained markers and a panel of antibodies must be applied, still.
We performed whole transcriptome sequencing (WTS) on two cases of FDC-S in order to explore the biology of this rare tumor in terms of gene fusions and gene expression.
Our study did not identify fusion transcripts in the two cases of FDC-S analyzed excluding the presence of chromosome translocations of oncogenic significance in these patients. However, we could use expression values obtained from WTS to identify two novel immunohistochemical markers for both normal and neoplastic FDC: Follicular dendritic cell secreted protein (FDCSP) and Serglycin (SRGN).
FDCSP, isolated in 2002 from human reactive tonsils, is specifically expressed by B-cell-activating-FDC [26]. The few studies performed to understand the functions of this protein suggest that it can modulate B-cell immune response [26–27] and may have a role in autoimmune diseases [28–29]. SRGN is an intracellular proteoglycan implicated in secretory granules formation and immune response modulation as demonstrated in knock-out mouse models [30–31]. SRGN expression was reported in the transcriptome of different cell types, including stromal and endothelial cells [30]. Notably, both markers were previously associated with metastasis and dismal prognosis in a wide range of carcinomas [31–36] but we could not detect their expression, by immunohistochemistry, in our cohort of controls.
In FDC-S diagnosis, FDCSP and SRGN showed both a good sensitivity, higher than CD23 and Claudin 4, and an extremely good specificity, evaluated on 214 controls including carcinomas, soft tissue tumors, melanomas, thymomas and interdigitating dendritic cell sarcomas. SRGN, together with CD21 and CD35, was negative in all non-FDC-S tumors; FDCSP, expressed by one case of SNHP, one case of SS and one case of interdigitating dendritic cell sarcoma, still showed a better performance than other widely used FDC markers. Clusterin, highly expressed at transcriptome level, is a highly sensitive marker for FDC-S [37–38]. However, in our study, it stained also 60% of GISTs occurring in the GI tract, outside stomach (e.g. ileum wall), suggesting to use it with caution (eventually together with CD117 and DOG1). Meanwhile, Podoplanin, a sensitive marker for FDC-S [18], was expressed in two thirds of IMT, in a fraction of GIST, SFT and SS. Pathologists should be aware of these potential pitfalls in the differential diagnosis of FDC-S occurring in extranodal sites and should consider applying also other newly, highly specific markers, such as STAT6 [39] and TLE1 [40], in support.
In our study the combination of CXCL13, CD21, CD35, FDCSP and SRGN reached a very good discriminatory power in distinguishing FDC-S from other soft tissue tumors. In particular, CXCL13, a chemokine ligand recently reported in FDC-S [17], was the marker with best sensitivity and specificity combination (Table 2). The usefulness of our five marker combination was confirmed by the good results obtained by linear discriminant analysis and the almost perfect separation in the hierarchical clustering profile where all FDC-S cases, but one, grouped together. Notably, all the correctly classified FDC-S cases were positive for at least two markers among CXCL13, CD21, CD35, FDCSP and SRGN; this information could be easily applicable in the routine diagnosis of this tumor.
FDC-S with a highly defective phenotype do exist and were present also in our set of cases, this could be related to a different grade of differentiation or “activation” of neoplastic FDCs and suggests to further explore the biology of these tumors in correlation with other mesenchymal tumors with the same, putative, cell of origin [9].
In conclusion, by a combined approach of next generation sequencing and high throughput immunohistochemistry we identified two novel markers for FDC-S, FDCSP and SRGN, and suggested a highly effective and ready-to-use approach for its differential diagnosis with other mesenchymal tumors.
MATERIALS AND METHODS
Case selection
Twenty-two cases of follicular dendritic cell sarcoma cases (FDC-S) were retrieved from the archives of pathology of the University Hospitals of Brescia, Frankfurt am Main, Bologna, Lisbon, Barcelona, Vienna and Mumbay. Diagnosis were confirmed at a multi head microscope by the authors (C.A., F.F, M.L.H., S.H. and S.A.P.).
Eighty-seven cases of soft tissue tumors were retrieved from the archives of the department of pathology of Spedali Civili di Brescia. They included 19 Solitary fibrous tumors (SFT, all STAT6-positive) and 4 Sinonasal haemangiopericytoma (SNHP, STAT6- negative [41]), 20 gastrointestinal stromal tumors (GIST, DOG1 and/or CD117-positive), 12 synovial sarcomas (SS, all TLE1-positive, 9 with split of 18q11.2, evaluated by fluorescence in situ hybridization with SS18 probe), 6 inflammatory myofibroblastic tumors (IMT, 4 ALK-positive), 5 dermatofibrosarcomas protuberans (DFSP), 5 leiomyosarcomas (LMS), 5 angiosarcomas (AS), 4 desmoid-type fibromatosis (DTF), 3 undifferentiated pleomorphic sarcomas (UPS), 2 dedifferentiated liposarcomas (DLS) and 2 low-grade fibromyxoid sarcomas (LGFMS, all MUC4-positive). Cases were revised and diagnosis confirmed by expert soft tissue pathologists (Demographical data are detailed in Supplementary Table 3). Informed consent was obtained according to the Declaration of Helsinki and the local ethics committees agreed on the study.
RNA extraction from frozen material and whole transcriptome sequencing
Total ribonucleic acid (RNA) was extracted from the frozen tissue of four FDC-S (RNeasy Mini kit, Qiagen, Hilden, Germany). RNA integrity was evaluated by a Bioanalyzer (Agilent, Waldbronn, Germany) and two of the four cases showed sufficient quality for whole transcriptome sequencing (WTS). The latter was performed on an Illumina HiSeq platform (EMBL, Heidelberg, Germany) after library preparation with the SMARTer™ Ultra Low RNA Kit (Clontech, Mountain View, CA USA). The hg19 Genome Reference Consortium GRCh37 was used as reference genome applying the Genomatix Software (Genomatix GmbH. Munich, Germany) using default parameters.
Biostatistical analysis
Transcriptome data were analyzed for fusion transcripts by Genomatix Mining Station and Genome Analyzer.
Gene expression values were normalized and ordered per kilobase per million mapped reads (RPKM). Genes with RPKM higher or equal to 1 underwent gene annotation analysis by Gene Ontology Consortium-annotations-based Panther software (www.pantherdb.org) [42]; overrepresentation test (by bimodal test with Bonferroni correction) and enrichment test (by Mann-Whitney test) were performed on both Panther and Genomatix softwares.
After immunohistochemical analysis, sensitivity, specificity and corresponding p-values were calculated for candidate and known FDC-S markers. Receiver Operating Characteristic (ROC) curve analysis was performed with the ©1993–2016 MedCalc Software using the methodology detailed in DeLong et al. [43] for calculation of the Standard Error and the exact Binomial Confidence Interval for the Area Under the Curve. Hierarchical clustering, principal component analysis (PCA) and linear discriminant analysis were performed with the PAST software [44].
Tissue microarray preparation and immunohistochemistry
Tissue microarray (TMA) blocks were created from FDC-S and 72 controls (control group #1, Supplementary Table 3), using an automated tissue microarray (TMA Master, 3DHistech, Budapest, Hungary). Three representative tumor cores (1 or 1.5 mm diameter) were identified on hematoxylin and eosin (H&E) stained sections and punched from the original tissue blocks.
Immunohistochemistry for known and novel FDC markers was performed on these TMAs. In order to further confirm their specificity, FDCSP and SRGN were additionally applied on TMAs including 114 carcinomas of different origin, previously described [24], and on whole tissue sections of 28 additional control cases (control group #2, Supplementary Table 3). Antibodies applied are detailed in Supplementary Table 5.
Immunohistochemistry for FDCSP and SRGN was performed with automatic immunostainer (Dako, Glostrup, Denmark) or manually, showing equivalent results; immunohistochemistry on controls was performed automatically (Leica Biosystems, Wetzlar, Germany).
Stained sections were digitalized using the Aperio Scanscope System (Leica Biosystems) and evaluated by two pathologists (S.H. and L.L.). Cases were scored positive when reaction occurred on at least 30% of tumor cells.
ACKNOWLEDGEMENTS
The authors are indebted to the colleagues pathologists who sent cases in consultation and to the pathologists of Spedali Civili of Brescia who made the diagnosis of tumors used as controls. The authors thank Prof William Vermi, Dr Antonella Laginestra and Dr Andrea Lorenzi for helpful comments, Yvonne Michel and Katharina Sandkühler for excellent technical assistance. Preliminary data from the study were presented at the EAHP Meeting 2016, held in Basel, Switzerland.
CONFLICTS OF INTEREST
The authors have no conflicts of interest to declare.
FUNDING
M.B. is supported by Fondazione Beretta (Brescia, Italy). Heinrich- und Erna-Schaufler-Stiftung to SH.
REFERENCES
1. Swerdlow SH, Campo E, Harris NL, Jaffe ES, Pileri SA, Stein H, Thiele J, JW V. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues. IARC. 2008.
2. Facchetti F, Lorenzi L. Follicular dendritic cells and related sarcoma. Semin Diagn Pathol. 2016; 33:159–66.
3. Mabbott NA, Kenneth Baillie J, Kobayashi A, Donaldson DS, Ohmori H, Yoon SO, Freedman AS, Freeman TC, Summers KM. Expression of mesenchyme-specific gene signatures by follicular dendritic cells: insights from the meta-analysis of microarray data from multiple mouse cell populations. Immunology. 2011; 133:482–98.
4. Aguzzi A, Kranich J, Krautler NJ. Follicular dendritic cells: origin, phenotype, and function in health and disease. Trends Immunol. 2014; 35:105–13.
5. Malhotra D, Fletcher AL, Turley SJ. Stromal and hematopoietic cells in secondary lymphoid organs: partners in immunity. Immunol Rev. 2013; 251:160–76.
6. Suzuki K, Maruya M, Kawamoto S, Sitnik K, Kitamura H, Agace WW, Fagarasan S. The sensing of environmental stimuli by follicular dendritic cells promotes immunoglobulin A generation in the gut. Immunity. 2010; 33:71–83.
7. Fletcher C.D.M. BJA, Hogendoorn P.C.W., Mertens F. WHO Cassification of Tumors of Soft tissue and Bone. IARC. 2013.
8. Kranich J, Krautler NJ, Heinen E, Polymenidou M, Bridel C, Schildknecht A, Huber C, Kosco-Vilbois MH, Zinkernagel R, Miele G, Aguzzi A. Follicular dendritic cells control engulfment of apoptotic bodies by secreting Mfge8. J Exp Med. 2008; 205:1293–302.
9. Krautler NJ, Kana V, Kranich J, Tian Y, Perera D, Lemm D, Schwarz P, Armulik A, Browning JL, Tallquist M, Buch T, Oliveira-Martins JB, Zhu C, et al. Follicular dendritic cells emerge from ubiquitous perivascular precursors. Cell. 2012; 150:194–206.
10. Kranich J, Krautler NJ. How Follicular Dendritic Cells Shape the B-Cell Antigenome. Front Immunol. 2016; 7:225.
11. Gounder M, Desai V, Kuk D, Agaram N, Arcila M, Durham B, Keohan ML, Dickson MA, D’Angelo SP, Shukla N, Moskowitz C, Noy A, Maki RG, et al. Impact of surgery, radiation and systemic therapy on the outcomes of patients with dendritic cell and histiocytic sarcomas. Eur J Cancer. 2015; 51:2413–22.
12. Saygin C, Uzunaslan D, Ozguroglu M, Senocak M, Tuzuner N. Dendritic cell sarcoma: a pooled analysis including 462 cases with presentation of our case series. Crit Rev Oncol Hematol. 2013; 88:253–71.
13. Jabbour MN, Fedda FA, Tawil AN, Shabb NS, Boulos FI. Follicular dendritic cell sarcoma of the head and neck expressing thyroid transcription factor-1: a case report with clinicopathologic and immunohistochemical literature review. Appl Immunohistochem Mol Morphol. 2014; 22:705–12.
14. Wang RF, Han W, Qi L, Shan LH, Wang ZC, Wang LF. Extranodal follicular dendritic cell sarcoma: A clinicopathological report of four cases and a literature review. Oncol Lett. 2015; 9:391–8.
15. Gong S, Auer I, Duggal R, Pittaluga S, Raffeld M, Jaffe ES. Epstein-Barr virus-associated inflammatory pseudotumor presenting as a colonic mass. Hum Pathol. 2015; 46:1956–61.
16. Chan JK, Fletcher CD, Nayler SJ, Cooper K. Follicular dendritic cell sarcoma. Clinicopathologic analysis of 17 cases suggesting a malignant potential higher than currently recognized. Cancer. 1997; 79:294–313.
17. Vermi W, Lonardi S, Bosisio D, Uguccioni M, Danelon G, Pileri S, Fletcher C, Sozzani S, Zorzi F, Arrigoni G, Doglioni C, Ponzoni M, Facchetti F. Identification of CXCL13 as a new marker for follicular dendritic cell sarcoma. J Pathol. 2008; 216:356–64.
18. Yu H, Gibson JA, Pinkus GS, Hornick JL. Podoplanin (D2–40) is a novel marker for follicular dendritic cell tumors. Am J Clin Pathol. 2007; 128:776–82.
19. Facchetti F, Lonardi S, Gentili F, Bercich L, Falchetti M, Tardanico R, Baronchelli C, Lucini L, Santin A, Murer B. Claudin 4 identifies a wide spectrum of epithelial neoplasms and represents a very useful marker for carcinoma versus mesothelioma diagnosis in pleural and peritoneal biopsies and effusions. Virchows Arch. 2007; 451:669–80.
20. Vermi W, Giurisato E, Lonardi S, Balzarini P, Rossi E, Medicina D, Bosisio D, Sozzani S, Pellegrini W, Doglioni C, Marchetti A, Rossi G, Pileri S, et al. Ligand-dependent activation of EGFR in follicular dendritic cells sarcoma is sustained by local production of cognate ligands. Clin Cancer Res. 2013; 19:5027–38.
21. Sun X, Chang KC, Abruzzo LV, Lai R, Younes A, Jones D. Epidermal growth factor receptor expression in follicular dendritic cells: a shared feature of follicular dendritic cell sarcoma and Castleman’s disease. Hum Pathol. 2003; 34:835–40.
22. Griffin GK, Sholl LM, Lindeman NI, Fletcher CD, Hornick JL. Targeted genomic sequencing of follicular dendritic cell sarcoma reveals recurrent alterations in NF-kappaB regulatory genes. Mod Pathol. 2016; 29:67–74.
23. Heesters BA, Myers RC, Carroll MC. Follicular dendritic cells: dynamic antigen libraries. Nat Rev Immunol. 2014; 14:495–504.
24. Vermi W, Micheletti A, Lonardi S, Costantini C, Calzetti F, Nascimbeni R, Bugatti M, Codazzi M, Pinter PC, Schakel K, Tamassia N, Cassatella MA. slanDCs selectively accumulate in carcinoma-draining lymph nodes and marginate metastatic cells. Nat Commun. 2014; 5:3029.
25. Lorenzi L, Lonardi S, Petrilli G, Tanda F, Bella M, Laurino L, Rossi G, Facchetti F. Folliculocentric B-cell-rich follicular dendritic cells sarcoma: a hitherto unreported morphological variant mimicking lymphoproliferative disorders. Hum Pathol. 2012; 43:209–15.
26. Marshall AJ, Du Q, Draves KE, Shikishima Y, HayGlass KT, Clark EA. FDC-SP, a novel secreted protein expressed by follicular dendritic cells. J Immunol. 2002; 169:2381–9.
27. Hou S, Landego I, Jayachandran N, Miller A, Gibson IW, Ambrose C, Marshall AJ. Follicular dendritic cell secreted protein FDC-SP controls IgA production. Mucosal Immunol. 2014; 7:948–57.
28. Meng HX, Li HN, Geng JS, Ohe R, Yu XY, E XQ, Ye F, Yang SR, Kato T, Zhang L, Ishida A, Ohta N, Jin XM, et al. Decreased expression of follicular dendritic cell-secreted protein correlates with increased immunoglobulin A production in the tonsils of individuals with immunoglobulin A nephropathy. Transl Res. 2015; 166:281–91.
29. Hamm CM, Reimers MA, McCullough CK, Gorbe EB, Lu J, Gu CC, Li E, Dieckgraefe BK, Gong Q, Stappenbeck TS, Stone CD, Dietz DW, Hunt SR. NOD2 status and human ileal gene expression. Inflamm Bowel Dis. 2010; 16:1649–57.
30. Kolset SO, Pejler G. Serglycin: a structural and functional chameleon with wide impact on immune cells. J Immunol. 2011; 187:4927–33.
31. Korpetinou A, Skandalis SS, Labropoulou VT, Smirlaki G, Noulas A, Karamanos NK, Theocharis AD. Serglycin: at the crossroad of inflammation and malignancy. Front Oncol. 2014; 3:327.
32. Wang C, Zhou L, Li S, Wei J, Wang W, Zhou T, Liao S, Weng D, Deng D, Weng Y, Wang S, Ma D. C4orf7 contributes to ovarian cancer metastasis by promoting cancer cell migration and invasion. Oncol Rep. 2010; 24:933–9.
33. Chia CS, Ong WS, Li XJ, Soong YL, Chong FT, Tan HK, Soo KC, Qian CN, Teh BT, Iyer NG. Serglycin expression: An independent marker of distant metastases in nasopharyngeal carcinoma. Head Neck. 2016; 38:21–8.
34. He L, Zhou X, Qu C, Tang Y, Zhang Q, Hong J. Serglycin (SRGN) overexpression predicts poor prognosis in hepatocellular carcinoma patients. Med Oncol. 2013; 30:707.
35. Li XJ, Ong CK, Cao Y, Xiang YQ, Shao JY, Ooi A, Peng LX, Lu WH, Zhang Z, Petillo D, Qin L, Bao YN, Zheng FJ, et al. Serglycin is a theranostic target in nasopharyngeal carcinoma that promotes metastasis. Cancer Res. 2011; 71:3162–72.
36. Li XJ, Qian CN. Serglycin in human cancers. Chin J Cancer. 2011; 30:585–9.
37. Grogg KL, Lae ME, Kurtin PJ, Macon WR. Clusterin expression distinguishes follicular dendritic cell tumors from other dendritic cell neoplasms: report of a novel follicular dendritic cell marker and clinicopathologic data on 12 additional follicular dendritic cell tumors and 6 additional interdigitating dendritic cell tumors. Am J Surg Pathol. 2004; 28:988–98.
38. Grogg KL, Macon WR, Kurtin PJ, Nascimento AG. A survey of clusterin and fascin expression in sarcomas and spindle cell neoplasms: strong clusterin immunostaining is highly specific for follicular dendritic cell tumor. Mod Pathol. 2005; 18:260–6.
39. Thway K, Ng W, Noujaim J, Jones R, Fisher C. The Current Status of Solitary Fibrous Tumor: Diagnostic Features, Variants, and Genetics. Int J Surg Pathol. 2016; 24:281–92.
40. Thway K, Fisher C. Synovial sarcoma: defining features and diagnostic evolution. Ann Diagn Pathol 2014; 18:369–80.
41. Agaimy A, Barthelmess S, Geddert H, Boltze C, Moskalev EA, Koch M, Wiemann S, Hartmann A, Haller F. Phenotypical and molecular distinctness of sinonasal haemangiopericytoma compared to solitary fibrous tumour of the sinonasal tract. Histopathology. 2014; 65:667–73.
42. Mi H, Muruganujan A, Casagrande JT, Thomas PD. Large-scale gene function analysis with the PANTHER classification system. Nat Protoc. 2013; 8:1551–66.
43. DeLong ER, DeLong DM, Clarke-Pearson DL. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics. 1988; 44:837–45.
44. Hammer Øyvind HDAT, Ryan Paul D. PAST: Paleontological statistics software package for education and data analysis. Palaeontologia Electronica 2001; 4:4:9.

