INTRODUCTION
Non-Hodgkin lymphoma (NHL) is one of the most common cancers. Overall NHL incidence has been rising for several decades in industrialized countries and then started to level off in the mid-1990s.[1, 2] During 1973-2010, overall NHL incidence increased by 1.64% (from 3.43 to 5.05 per 100,000 person-years) in males and by 2.50% (from 1.77 to 3.65 per 100,000 person-years) in females in Shanghai, China.[3] However, age-standardized NHL mortality rate in China [4] has not declined over time as observed in other countries since the 1990s.[1, 5]
The causes of most NHL cases are poorly understood. There are significant differences in geographical distributions of NHL incidence,[6] probably due to differences in risk factors, genetic susceptibility, and diagnosis/classification. Immunosuppression plays an important role in lymphomagenesis [7] and certain immunodeficiency disorders including congenital immunodeficiency disorders, acquired immunodeficiency syndrome (AIDS), and autoimmune disorders (e.g., rheumatoid arthritis and Sjögren’s syndrome) increase the risk of developing NHL.[8] Early studies found that variants in genes involved in immunoregulatory pathways were associated with an increased or reduced risk of NHL.[9] A meta-analysis showed that TNF-α rs1800629 was associated with an increased risk of overall NHL and diffuse large B-cell lymphoma (DLBCL, a major NHL subtype).[10] After pooling three studies together, Hosgood III and colleagues found an association between IL-1RN rs2637988 and increased risk of overall NHL.[11] However, studies on these variants have been mainly conducted among subjects of European ancestry and have rarely been replicated in other populations. In this study, we examined the associations between immunoregulatory gene variants and NHL risk in a Chinese population.
RESULTS
Cases and controls were similar regarding the distributions of sex, age, marriage status, and alcohol consumption. But cases tended to be less educated, less likely to be overweight, and more likely to be a smoker (past or current) or have a family history of cancer (Table 1). All nine SNPs had a minor allele frequency larger than 5%. Eight of the nine SNPs had a Hardy-Weinberg Equilibrium (HWE) test P value in controls larger than 0.01. IL-4 rs2243267 had a HWE test P value in controls smaller than 1 x 10-4, which was removed from the association analysis. Table 2 shows that five SNPs were associated with an increased risk of overall NHL or B-cell lymphomas (particularly DLBCL). IL-10 rs1800893 was associated with an increased risk of overall NHL (per-minor-allele odds ratio [ORper-minor-allele]=2.64, 95% confidence interval [CI] 1.75-3.98), B-cell lymphomas (ORper-minor-allele=2.75, 95% CI 1.79-4.23), and DLBCL (ORper-minor-allele=3.03, 95% 1.85-5.01). Similar associations were found between IL-1RN rs4251961 and an increased risk of overall NHL (ORper-minor-allele =2.67, 95% CI 1.72-4.16), B-cell lymphomas (ORper-minor-allele=2.75, 95% CI 1.73-4.36), and DLBCL (ORper-minor-allele=3.39, 95% CI 1.99-5.97). Three TNF-α SNPs were associated with an increased NHL risk: rs1041981 was associated with an increased risk of B-cell lymphoma (ORper-minor-allele=1.45, 95% CI 1.02-2.05), particularly DLBCL (ORper-minor-allele=1.73, 95% CI 1.14-2.61); rs1800630 was associated with a higher risk of overall NHL (ORper-minor-allele for overall NHL=1.80, 95% CI 1.24-2.63), B-cell lymphomas (ORper-minor-allele =1.76, 95% CI 1.19-2.61), and DLBCL (ORper-minor-allele=1.88, 95% CI 1.17-3.03); rs2229094 was associated with an increased risk of overall NHL (ORper-minor-allele =1.55, 95% CI 1.02-2.37) and DLBCL (ORper-minor-allele=1.83, 95% CI 1.09-3.08). No statistically significant associations were found for the other three SNPs (IL-10 rs1518111, IL-10 rs3021094, and TNF-α rs1800629). We obtained linkage disequilibrium (LD) patterns of all 8 SNPs in the genes IL-10 and TNF-α using Haploview (Figure 1). As shown in Figure 1, the significant SNPs (TNF-α rs1800630 and TNF-α rs1041981) had a relatively high correlation (0.55) and were in fairly high LD with each other. The same trend was observed for the significant SNPs TNF-α rs1041981 and TNF-α rs2229094. We also found that IL-10 rs1518111, IL-10 rs3021094, and IL-10 rs1800893 had relatively high LD with each other.
Table 1: Characteristics of NHL Cases and Controls in Shanghai, China
Characteristics |
Controls (n=421) |
Cases (n=169) |
Total (n=590) |
|---|---|---|---|
Gender |
|||
Female |
258 (61.3) |
103 (60.9) |
361 (61.2) |
Male |
163 (38.7) |
66 (39.1) |
229 (38.8) |
Age (year) |
|||
<50 |
106 (25.2) |
50 (29.6) |
156 (26.4) |
50-65 |
186 (44.2) |
77 (45.6) |
263 (44.6) |
65+ |
129 (30.6) |
42 (24.9) |
171 (29.0) |
Mean (SD) |
57.9 (13.7) |
55.1 (12.9) |
57.1 (13.5) |
Education* |
|||
Less than secondary school |
73 (17.3) |
144 (85.2) |
217 (36.8) |
Secondary school |
268 (63.7) |
25 (14.8) |
293 (49.7) |
College or university |
80 (19.0) |
80 (14.5) |
|
Marriage |
|||
Never |
10 (2.4) |
7 (4.1) |
17 (2.9) |
Current |
361 (85.8) |
154 (91.1) |
515 (87.3) |
Divorced/Widowed |
33 (7.8) |
8 (4.7) |
18 (3.1) |
Missing |
17 (4.0) |
17 (2.9) |
|
BMI* (kg/m2) |
|||
<18.5 |
201 (47.7) |
104 (61.5) |
305 (51.7) |
18.5-24 |
11 (2.6) |
10 (5.9) |
21 (3.6) |
24-28 |
165 (39.2) |
46 (27.2) |
211 (35.7) |
28+ |
44 (10.5) |
9 (5.3) |
53 (8.9) |
Mean (SD) |
24.3 (6.7) |
22.9 (3.1) |
23.9 (5.9) |
Smoking* |
|||
Never |
267 (63.4) |
94 (55.6) |
361 (61.2) |
Past |
37 (8.8) |
35 (20.7) |
72 (12.2) |
Current |
116 (27.6) |
40 (23.7) |
156 (26.4) |
Missing |
1 (0.2) |
1 (0.2) |
|
Alcohol |
|||
No |
328 (77.9) |
128 (75.7) |
456 (77.3) |
Yes |
91 (21.6) |
41 (24.3) |
132 (22.4) |
Missing |
2 (0.5) |
2 (0.3) |
|
Family history of cancer* |
|||
No |
322 (76.5) |
51 (30.2) |
373 (63.2) |
Yes |
98 (23.3) |
118 (69.8) |
216 (36.6) |
Missing |
1 (0.2) |
1 (0.2) |
Note: The number in bracket is proportion (%) of the total number in controls; *, P<0.001 for the comparisons between cases and controls.
Table 2: Associations between immunity gene SNPs and NHL risk
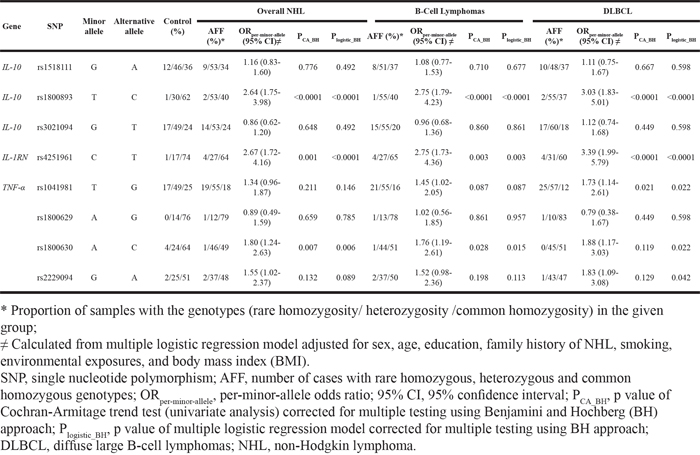
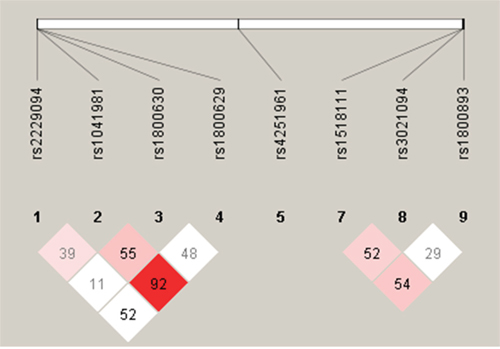
Figure 1: LD plot of the 8 SNPs in genes IL-10 and TNF-α.
Notes: LD plot showing LD patterns among the 8 SNPs in genes IL-10 and TNF-α genotyped in the 590 samples. The LD between the SNPs is measured as r2 and shown (×100) in the diamond at the intersection of the diagonals from each SNP. r2 = 0 is shown as white, 0< r2<1 is shown in pink and r2 =1 is shown in red. The analysis track at the top shows the SNPs according to chromosomal location.
DISCUSSION
In this study, we found associations between five immunoregulatory gene SNPs and a higher NHL risk: four SNPs (IL-10 rs1800893, IL-1RN rs4251961, TNF-α rs1800630, and TNF-α rs2229094) were associated with an increased risk of overall NHL and DLBCL, while TNF- α rs1041981was associated with an increased risk of DLBCL only.
Cytokines including tumor necrosis factor (TNF) play a role in lymphomagenesis and genetic variants that upregulate or downregulate the production of cytokines may relate to NHL risk.[12] Previous studies have shown an association between TNF-α rs1800629 and an increased risk of NHL (particularly subtypes DLBCL, marginal zone lymphoma, and T-cell lymphoma) when ethnicity was not considered.[9, 13] However, a pooled analysis showed that the association between the SNP and NHL risk varied by ethnicity: TNF-α rs1800629 minor allele A (genotypes GA/AA) was associated with a reduced risk of NHL in Asian participants living in Australia (OR=0.52, 95% CI 0.32-0.85, compared to genotype GG), but was associated with an increased NHL risk in African Americans and Hispanic Whites.[10] The ethnic difference was further verified by a recent meta-analysis.[14] The inverse association was confirmed in a recent study in Chinese and Korean populations,[15] where rs1800629 minor allele A was associated with 20% lower risk for NHL. The inverse association was not observed in the present study and this might relate to the small sample size and the difference in study methods (e.g., only Han Chinese living in China were included in the present study).
Another SNP (TNF- α rs1800630 minor allele A) was associated with an increased risk of NHL in the present study and this is consistent with the findings of the other two studies in Asia.[15, 16] The present study and the study by Gu [16] included Han Chinese only and found a 80% higher risk for NHL among TNF-α rs1800630 minor allele A carriers. Another study involving a mixed population of Han Chinese and Koreans showed a 19% increased NHL risk among minor allele A carriers.[15] In an earlier study in Australia, this SNP was also associated with a higher risk for NHL particularly for DLBCL.[17] Two TNF- α SNPs (rs1041981 and rs2229094) examined in the present study have been rarely investigated previously. TNF-α rs1041981 was associated with a higher risk of AIDS-related NHL without considering a CD4 count change, but the association did not hold after adjusting for the rate of the change in CD4 count.[18] In the present study, both TNF-α rs1041981 and TNF-α rs2229094 minor alleles are associated with a higher risk of NHL, particularly DLBCL, and the findings need to be tested in a larger study.
IL-10, also known as human cytokine synthesis inhibitory factor, is a cytokine with multiple effects in immunoregulation and anti-inflammation. We found that IL-10 rs1800893 was associated with an increased risk of NHL, but the association was not observed in a previous study in Germany.[19] Two other common IL-10 SNPs (rs1800890 and rs1800896) were associated with an increased risk of NHL and its major subtype DLBCL in Hispanic Whites, but not in Asians or African Americans in previous studies [10] (therefore they were not examined in the present study). A more recent study confirmed that there was no a significant association between IL-10 rs1800896 and NHL risk in Chinese.[20]
The protein encoded by IL-1RN gene inhibits the activities of interleukin 1 α and β and modulates a variety of interleukin 1 related immune and inflammatory responses. IL-1RN rs4251961 was found to be associated with an increased risk of cancers e.g. colorectal cancer,[21] but our study for the first time reports its association with an increased risk of NHL. Other IL-1RN SNPs (rs2637988 and rs454078) were also associated with an increased NHL risk in previous studies,[13, 15] but were not included in the present analysis.
Previous genome-wide association studies (GWAS) have identified a few SNPs that were associated with an increased risk of overall NHL,[22] chronic lymphocytic leukemia,[23, 24] and follicular lymphoma,[22, 23, 25, 26] in populations of European ancestry. Fewer studies have been conducted in Asian populations. In Japan, rs751837 was associated with a higher risk of DLBCL.[27] However, none of these GWAS associations have been replicated in other populations. Two recent Chinese studies have identified SNPs associated with an increased risk of B-cell NHL (rs6773854 located between BCL6 and LPP genes on chromosome 3q27, IRF4 rs872071, and rs2647012 in HLA-region)[28, 29] and T-cell NHL (ACOXL rs17483466 and IRF4 rs872071),[28] but none of these SNPs are in immunoregulatory genes.
NHL is a cancer of immune system and immune deficiency is one of a few well-established risk factors. There is also evidence on the role of other immune diseases including atopy and autoimmune diseases in NHL risk.[7] Single nucleotide variants may change immunoregulatory gene functions and therefore influence NHL risk through the immunoregulation pathways. This hypothesis is supported by the findings from the present study and previous studies despite the heterogeneity in findings.[10, 11, 30] GWAS studies have also identified a few immune gene SNPs associated with an increased risk of NHL or specific NHL subtypes, but overall the findings were inconsistent and suggestive.
There are several limitations in the present study. First, the observed associations may not be interpreted as causal relationships due to the nature of case-control design. It is likely that the findings have been biased as controls were more likely residents from other provinces. Information on covariates was collected retrospectively via an interview, so recall bias and misclassification could not be ruled out. The small sample size also limited the ability to analyze other NHL subtypes (e.g., follicular lymphoma, marginal zone lymphoma, T-cell lymphoma). The analysis was limited to a small number of candidate SNPs selected based on previous epidemiologic study findings and has not tested SNPs associated with NHL risk in more recent studies. Despite that, the present study supports that immunoregulatory gene variants play a role in NHL risk. Further research on the racial/ethnic difference in associations between genetic variants and NHL risk as well as the functional experiment analysis of the associations may advance the understanding of the etiology of NHL.
MATERIALS AND METHODS
Eligible cases were NHL patients diagnosed between 2003 and 2008 according to the WHO 2001 classification standard and aged 18 or older, as described in detail previously.[31] Patients with a provisional diagnosis of NHL from the participating hospitals were referred to a leukemia and lymphoma diagnosis and research laboratory located at Fudan University, China. Peripheral blood, bone marrow aspirates, tissue and core biopsies of patients were collected in conjunction with diagnostic procedures and were sent to the laboratory for morphological, immunological, cytogenetic, and molecular genetic analyses.[32, 33] Controls were randomly selected from patients in any other wards from the same hospitals matched on sex and age group frequency. Those with cancers or non-malignant lymphatic or hematopoietic diseases, or having family connections to the cases were excluded. One hundred and sixty-nine cases and 421 controls were included into the study. Each patient completed a questionnaire containing questions regarding demographics, family history, lifestyles, occupational and environmental exposures. This project has been approved by the Ethical Review Committee of Fudan University in accordance with the International Ethical Guidelines for Biomedical Research Involving Human Subjects (2002).[34] Informed consent was obtained from all individual participants in the study. Genes were selected based on two criteria: they are involved in immunoregulation and were associated with NHL risk in previous studies, using the method by Xu.[35] A total of nine SNPs in four genes were selected, including IL-10 (rs1518111, rs1800893, and rs3021094), IL-1RN (rs4251961), IL-4 (rs2243267), and TNF-α (rs1041981, rs2229094, rs1800629, and rs1800630). Genotyping was conducted using the real-time polymerase chain reaction method on an ABI 7900HT sequence detection system at the State Key Laboratory of Genetics at Fudan University.[36] For quality control, 10% of samples were randomly selected and re-genotyped. The concordance rates for replicated samples were 100%.
Chi-square test and t-test were used for comparisons of categorical and continuous variables between cases and controls. To test the associations between SNPs and NHL risk, we first performed the Cochran-Armitage trend test using only genotype data.[37] Multiple logistic regression models were then applied based on a genetic additive model. ORper-minor-allele and 95% CI were estimated using major allele as a reference. The models were fitted for overall NHL, B-cell lymphomas, and diffuse large B-cell lymphoma (DLBCL) separately, but not for T-cell lymphoma patients due to the small sample size. As a prior,[8] we included covariates sex, age, education, family history of NHL, smoking status, environmental exposures (i.e., exposed to one or more of substances including benzene, solvents, metals, agrichemicals, or pesticides), body mass index (BMI) and hair dye use in the initial model. The inclusion of a variable into the final model was determined using the stepwise approach (“stepAIC” function in MASS R package).[38] Covariates sex, age, education, family history of NHL, smoking, environmental exposures, and BMI were controlled in the final logistic regression model.[37] If a sample with missing data on any of the variables in the regression model, the sample was excluded from the model analysis. P values were adjusted using the Benjamini and Hochberg multiple testing procedure.[39] All statistical analyses were undertaken in statistical software R and PLINK. [37] Haplotype analysis was performed using Haploview.[40]
ACKNOWLEDGMENTS
We thank the Shanghai Health Study for sharing patient data and DNA specimens and the State Key Laboratory of Genetics of Fudan University for genotyping analysis.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
FUNDING
This project was supported by National Nature Science Foundation of China (project number 30671739).
Authors’ contributions
X.Y. and H.F. designed the study. C.W. performed laboratory and field works. K.Z. and P.H. analyzed the studies. X.Y. drafted the first version and all co-authors have reviewed and approved the manuscript.
REFERENCES
1. Bosetti C, Levi F, Ferlay J, Lucchini F, Negri E, Vecchia C La. Incidence and mortality from non-Hodgkin lymphoma in Europe: The end of an epidemic? Int J Cancer. 2008; 123: 1917–23.
2. Clarke CA, Glaser SL. Changing incidence of non-Hodgkin lymphomas in the United States. Cancer. 2002; 94: 2015–23.
3. Bao P-P, Zheng Y, Wu C-X, Huang Z-Z, Gao Y-T, Jin F, Xiang Y-B, Zhong W-J, Lu W, Wu F. Cancer incidence in urban Shanghai, 1973-2010: an updated trend and age-period-cohort effects. BMC Cancer; 2016; 16: 284. doi: 10.1186/s12885-016-2313-2.
4. Zhou M, Wang H, Zhu J, Chen W, Wang L, Liu S, Li Y, Wang L, Liu Y, Yin P, Liu J, Yu S, Tan F, et al. Cause-specific mortality for 240 causes in China during 1990-2013: A systematic subnational analysis for the Global Burden of Disease Study 2013. Lancet. 2016; 387: 251–72. doi: 10.1016/S0140-6736(15)00551-6.
5. Saika K, Zhang M. Comparison of time trends in Non-Hodgkin’s lymphoma mortality (1990-2006) between countries based on the WHO mortality database. Jpn J Clin Oncol. 2011; 41: 154–5.
6. Katanoda K, Yako-Suketomo H. Comparison of time trends in Hodgkin and non-Hodgkin lymphoma incidence (1973-97) in East Asia, Europe and USA, from cancer incidence in five continents Vol. IV-VIII. Jpn J Clin Oncol. 2008; 38: 391–3.
7. Grulich AE, Vajdic CM, Cozen W. Altered immunity as a risk factor for non-Hodgkin lymphoma. Cancer Epidemiol Biomarkers Prev. 2007; 16: 405–8. doi: 10.1158/1055-9965.EPI-06-1070.
8. Ekström-Smedby K. Epidemiology and etiology of non-Hodgkin lymphoma—a review. Acta Oncol. 2006; 45: 258–71. doi: 10.1080/02841860500531682.
9. Skibola CF, Curry JD, Nieters A. Genetic susceptibility to lymphoma. Haematologica. 2007; 92: 960–9. doi: 10.3324/haematol.11011.
10. Skibola CF, Bracci PM, Nieters A, Brooks-Wilson A, de Sanjosé S, Hughes AM, Cerhan JR, Skibola DR, Purdue M, Kane E, Lan Q, Foretova L, Schenk M, et al. Tumor necrosis factor (TNF) and lymphotoxin-alpha (LTA) polymorphisms and risk of non-Hodgkin lymphoma in the InterLymph Consortium. Am J Epidemiol. 2010; 171: 267–76. doi: 10.1093/aje/kwp383.
11. Hosgood HD, Purdue MP, Wang SS, Zheng T, Morton LM, Lan Q, Menashe I, Zhang Y, Cerhan JR, Grulich A, Cozen W, Yeager M, Holford TR, et al. A pooled analysis of three studies evaluating genetic variation in innate immunity genes and non-Hodgkin lymphoma risk. Br J Haematol. 2011; 152: 721–6. doi: 10.1111/j.1365-2141.2010.08518.x.
12. Purdue MP, Lan Q, Bagni R, Hocking WG, Baris D, Reding DJ, Rothman N. Prediagnostic serum levels of cytokines and other immune markers and risk of non-hodgkin lymphoma. Cancer Res. 2011; 71: 4898–907. doi: 10.1158/0008-5472.CAN-11-0165.
13. Rothman N, Skibola CF, Wang SS, Morgan G, Lan Q, Smith MT, Spinelli JJ, Willett E, De Sanjose S, Cocco P, Berndt SI, Brennan P, Brooks-Wilson A, et al. Genetic variation in TNF and IL10 and risk of non-Hodgkin lymphoma: a report from the InterLymph Consortium. Lancet Oncol. 2006; 7: 27–38. doi: 10.1016/S1470-2045(05)70434-4.
14. Zhai K, Ding J, Zhou Y. Different role of tumor necrosis factor-α polymorphism in non-Hodgkin lymphomas among Caucasian and Asian populations: a meta-analysis. Int J Mol Sci; 2014; 15: 7684–98. doi: 10.3390/ijms15057684.
15. Hosgood HD, Au W-YY, Kim HJHNH-JJHNH-J, Liu J, Hu W, Tse J, Song B, Wong K-FF, Lee J-JJ, Chanock SJ, Siu LP, Purdue MP, Shin M-HH, et al. IL10 and TNF variants and risk of non-Hodgkin lymphoma among three Asian populations. Int J Hematol. 2013; 97: 793–9. doi: 10.1007/s12185-013-1345-5.
16. Gu X, Shen Y, Fu L, Zuo HHY, Yasen H, He P, Guo XXH, Shi YY wei, Yusufu M. Polymorphic variation of inflammation-related genes and risk of non-hodgkin lymphoma for Uygur and Han Chinese in Xinjiang. Asian Pacific J Cancer Prev. Asian Pacific Organization for Cancer Prevention; 2014; 15: 9177–83. doi: 10.7314/APJCP.2014.15.21.9177.
17. Purdue MP, Lan Q, Kricker A, Grulich AE, Vajdic CM, Turner J, Whitby D, Chanock S, Rothman N, Armstrong BK. Polymorphisms in immune function genes and risk of non-Hodgkin lymphoma: findings from the New South Wales non-Hodgkin Lymphoma Study. Carcinogenesis. 2007; 28: 704–12. doi: 10.1093/carcin/bgl200.
18. Aissani B, Ogwaro KM, Shrestha S, Tang J, Breen EC, Wong H-L, Jacobson LP, Rabkin CS, Ambinder RF, Martinez-Maza O, Kaslow RA, Madeleine MM, Johnson LG, et al. The major histocompatibility complex conserved extended haplotype 8.1 in AIDS-related non-Hodgkin lymphoma. J Acquir Immune Defic Syndr. 2009; 129: 170–9. doi: 10.1007/s10549-011-1520-4.
19. Kube D, Hua T-D, Klöss M, Kulle B, Brockmöller J, Wojnowski L, Löffler M, Pfreundschuh M, Trümper L. The interleukin-10 gene promoter polymorphism -1087AG does not correlate with clinical outcome in non-Hodgkin’s lymphoma. Genes Immun. 2007; 8: 164–7. doi: 10.1038/sj.gene.6364364.
20. Lim Y-Y, Chin Y-M, Tai M-C, Fani S, Chang K-M, Ong T-C, Bee P-C, Gan G-G, Ng C-C. Analysis of interleukin-10 promoter single nucleotide polymorphisms and risk of non-Hodgkin lymphoma in a Malaysian population. Leuk Lymphoma. 2015; 56: 163–8. doi: 10.3109/10428194.2014.907895.
21. Bondurant KL, Lundgreen A, Herrick JS, Kadlubar S, Wolff RK, Slattery ML. Interleukin genes and associations with colon and rectal cancer risk and overall survival. Int J Cancer. 2013; 132: 905–15. doi: 10.1002/ijc.27660.
22. Vijai J, Kirchhoff T, Schrader KA, Brown J, Dutra-Clarke AV, Manschreck C, Hansen N, Rau-Murthy R, Sarrel K, Przybylo J, Shah S, Cheguri S, Stadler Z, et al. Susceptibility loci associated with specific and shared subtypes of lymphoid malignancies. PLoS Genet. 2013; 9: e1003220. doi: 10.1371/journal.pgen.1003220.
23. Conde L, Halperin E, Akers NK, Brown KM, Smedby KE, Rothman N, Nieters A, Slager SL, Brooks-Wilson A, Agana L, Riby J, Liu J, Adami H-O, et al. Genome-wide association study of follicular lymphoma identifies a risk locus at 6p21.32. Nat Genet. 2010; 42: 661–4. doi: 10.1038/ng.626.
24. Slager SL, Achenbach SJ, Asmann YW, Camp NJ, Rabe KG, Goldin LR, Call TG, Shanafelt TD, Kay NE, Cunningham JM, Wang AH, Weinberg JB, Norman AD, et al. Mapping of the IRF8 gene identifies a 3’UTR variant associated with risk of chronic lymphocytic leukemia but not other common non-Hodgkin lymphoma subtypes. Cancer Epidemiol Biomarkers Prev. 2013; 22: 461–6. doi: 10.1158/1055-9965.EPI-12-1217.
25. Skibola CF, Berndt SI, Vijai J, Conde L, Wang Z, Yeager M, de Bakker PIW, Birmann BM, Vajdic CM, Foo J-N, Bracci PM, Vermeulen RCH, Slager SL, et al. Genome-wide association study identifies five susceptibility loci for follicular lymphoma outside the HLA region. Am J Hum Genet. 2014; 95: 462–71. doi: 10.1016/j.ajhg.2014.09.004.
26. Smedby KE, Foo JN, Skibola CF, Darabi H, Conde L, Hjalgrim H, Kumar V, Chang ET, Rothman N, Cerhan JR, Brooks-Wilson AR, Rehnberg E, Irwan ID, et al. GWAS of follicular lymphoma reveals allelic heterogeneity at 6p21.32 and suggests shared genetic susceptibility with diffuse large B-cell lymphoma. PLoS Genet. 2011; 7: e1001378. doi: 10.1371/journal.pgen.1001378.
27. Kumar V, Matsuo K, Takahashi A, Hosono N, Tsunoda T, Kamatani N, Kong S-Y, Nakagawa H, Cui R, Tanikawa C, Seto M, Morishima Y, Kubo M, et al. Common variants on 14q32 and 13q12 are associated with DLBCL susceptibility. J Hum Genet. 2011; 56: 436–9. doi: 10.1038/jhg.2011.35.
28. Tan DEK, Foo JN, Bei J-X, Chang J, Peng R, Zheng X, Wei L, Huang Y, Lim WY, Li J, Cui Q, Chew SH, Ebstein RP, et al. Genome-wide association study of B cell non-Hodgkin lymphoma identifies 3q27 as a susceptibility locus in the Chinese population. Nat Genet. 2013; 45: 804–7. doi: 10.1038/ng.2666.
29. Qiao Y, Zhou Y, Wu C, Zhai K, Han X, Chen J, Tian X, Chang J, Lu Z, Zhang B, Yu D, Yao J, Shi Y, et al. Risk of genome-wide association study-identified genetic variants for non-Hodgkin lymphoma in a Chinese population. Carcinogenesis. 2013; 34: 1516–9. doi: 10.1093/carcin/bgt082.
30. Wang SS, Cerhan JR, Hartge P, Davis S, Cozen W, Severson RK, Chatterjee N, Yeager M, Chanock SJ, Rothman N. Common genetic variants in proinflammatory and other immunoregulatory genes and risk for non-Hodgkin lymphoma. Cancer Res. 2006; 66: 9771–80. doi: 10.1158/0008-5472.CAN-06-0324.
31. Wong O, Harris F, Wang Y, Fu H. A hospital-based case-control study of non-hodgkin lymphoid neoplasms in shanghai: Analysis of personal characteristics, lifestyle, and environmental risk factors by subtypes of the WHO classification. J Occup Environ Med. 2010; 52: 39–53. doi: 10.1097/JOM.0b013e3181c5c399.
32. Gross S, Zhu X, Bao L, Ryder J, Le A, Chen Y, Wang X, Irons R. A prospective study of 728 cases of non-hodgkin lymphoma from a single laboratory in shanghai, china. Int J Hematol. 2008; 88: 165–73.
33. Wong O, Harris F, Armstrong TW, Hua F. A hospital-based case-control study of non-Hodgkin lymphoid neoplasms in Shanghai: analysis of environmental and occupational risk factors by subtypes of the WHO classification. Chem Biol Interact. Elsevier Ireland Ltd; 2010; 184: 129–46. doi: 10.1016/j.cbi.2009.10.016.
34. The Council for International Organizations of Medical Sciences (CIOMS). International Ethical Guidelines for Biomedical Research Involving Human Subjects. Geneva, Switzerland.; 2002.
35. Xu Z, Taylor JA. SNPinfo: integrating GWAS and candidate gene information into functional SNP selection for genetic association studies. Nucleic Acids Res. 2009; 37: W600-5. doi: 10.1093/nar/gkp290.
36. Liu Y, Zhang H, Zhou K, Chen L, Xu Z, Zhong Y, Liu H, Li R, Shugart YY, Wei Q, Jin L, Huang F, Lu D, et al. Tagging SNPs in non-homologous end-joining pathway genes and risk of glioma. Carcinogenesis. 2007; 28: 1906–13. doi: 10.1093/carcin/bgm073.
37. Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, Maller J, Sklar P, de Bakker PPIWPPIW, Daly MMJMMJ, Sham PCP. PLINK: a tool set for wholegenome association and population-based linkage analyses. Am J Hum Genet. 2007; 81: 559–575. doi: 10.1086/519795.
38. Venables W, Ripley B. Modern Applied Statistics with S. 4th ed. Springer; 2002.
39. Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B. 1995; 57: 289–300.
40. Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005; 21: 263–5. doi: 10.1093/bioinformatics/bth457.

