INTRODUCTION
Head and neck cancer is the sixth most common malignant tumors all around the world, accounting for approximately 6% of all cancers mortality [1–3]. Epithelial tumors arising in the oral cavity is the most high-frequently occurred cancer in head and neck region, and tongue squamous cell carcinoma (TSCC) is the most common histological malignancy of in this region [4]. TSCC comprises 25% to 40% of oral cancers and is the main cause of this type cancers mortality [5–7]. In spite of improvements in diagnostic tools and treatment prescription such as operation, chemotherapy and radiotherapy over the past few decades, TSCC cells extraordinary local invasion and lymphnode metastasis still are enormous challenges of the disease course of treatment, and it still has a high risk of developing secondary or recurrent tumors in the surrounding area and overall survival rates of advanced (stage III-IV) of it is less than 65% [8–10].
Long non-coding RNA (lncRNA) is a newly-identified class of RNAs that are more than 200 nucleotides in length and without protein-encoding ability. Accumulating evidence has shown that they play crucial roles in many physiological processes, such as carcinogenesis, by modulating gene expression at the epigenetic, transcriptional and posttranscriptional levels [11–15]. Furthermore, recent studies have identified multiple functional effects of lncRNAs involving in various cellular processes, such as tumor initiation, progression and metastasis, and the dysregulation of lncRNAs are a primary feature of some human cancers [16–19]. At the same time, a great quantity identified results revealed that lncRNAs play an important role in pathogenesis of head and neck cancers [20–22]. However, the TSCC associated lncRNAs and their influence on TSCC cellular progress need to be elucidated further.
In this study, we aim to profile expression patterns and dysregulation of lncRNAs in TSCC by analyzing two previously published TSCC data sets (GSE9844 [23] and GSE30784 [24]), based on Affymetrix gene expression microarray platform. One lncRNA, long intergenic non-coding RNA 673 (LINC00673) was significantly up-regulated in TSCC tissues than that in corresponding non-tumor tissues. Then, we examined its expression in fresh TSCC biopsies and paraffin-embedded tissues by quantitative real time polymerase chain reaction (qRT-PCR) and in situ hybridization, respectively. Furthermore, we performed in vitro knockdown experiments inhibiting LINC00673 expression in TSCC cells to assess the changes in tumor cell behavior when LINC00673 expression is lost.
This study represents a significant step forward in understanding the importance of lncRNAs in TSCC, and provided a novel insight concerning the role of LINC00673 in the progression of TSCC. Future studies based on these findings may lead to discover novel TSCC biomarkers or targeted therapies.
RESULTS
The expression profile of lncRNAs in TSCC tissues
In an attempt to identify novel long non-coding RNA in TSCC, two GEO datasets GSE9844 and GSE30784 were used to explore the lncRNA’s differential expression between TSCC tissues and normal lingual mucous membrane (Figure 1A). Through aggregation of differentially expressed lncRNAs signatures from these two GEO datasets, 12 overlapping probesets, revealing 6 upregulated and 6 downregulated lncRNAs (Figure 1B). Most were unknown and not well identified, such as LINC00673, LINC00152, LINC00520, LIN00094, LINC00511, EPB41L4A-AS1, and LINC00341. H19 [25–31], a famous oncogenic lncRNA in many cancers, also listed and overexpressed in TSCC samples among these overlapping probe sets. To investigate the role of lncRNAs in TSCC tumorigenesis, we focused on LINC00673, for which the expression was most significantly different and had never been reported in human cancer.
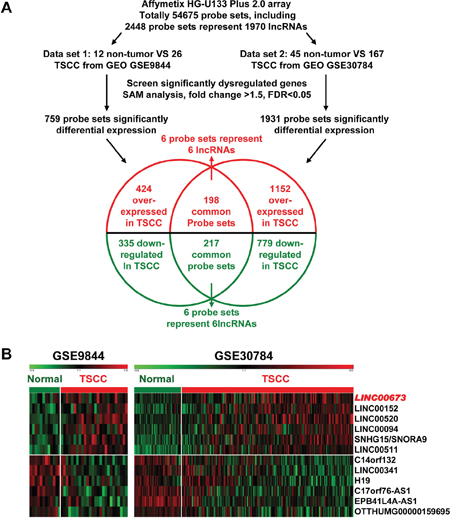
Figure 1: Dysregulated lncRNAs expression analysis using two independent TSCC cohorts and cDNA microarray analysis. A. Schematic overview of the workflow used to identify dysregulated lncRNAs in two TSCC microarray data cohorts (GSE9844 and GSE30784). B. Heatmap of 12 dysregulated lncRNAs mined from the GEO data set.
LINC00673 is high expressed in TSCC
LINC00673 expression was one of the most significantly upregulated in the TSCC tissues compared to non-tumor tissues according to the GSE9844 and GS30784 datasets (Figure 2A & 2B, p < 0.001 in both datasets). To assess the role of LINC00673 in TSCC progression, the expression levels of LINC00673 in 15 paired TSCC tissues and adjacent non-tumor tissues were tested by qRT-PCR. Results showed that the transcript levels of LINC00673 in TSCC tissues were significantly up-regulated compared to that in adjacent normal tissues (p = 0.020, Figure 2C), which was consistent with the GEO datasets.
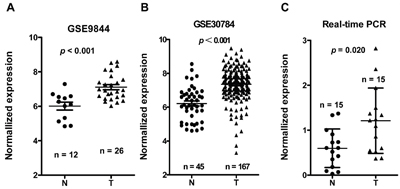
Figure 2: LINC00673 expression was upregulated in three independent cohorts of TSCC biopsies. LINC00673 expression, as measured by Affymetrix microarray, was upregulated in TSCC biopsies when compared with non-tumor lingual mucous membrane tissues in GSE30784 A. and GSE9844 B. C. LINC00673 was upregulated in TSCC through measured in 15 paired TSCC tissues and adjacent non-tumor tissues using quantitative PCR (qRT-PCR). T, tumor; N, non-tumor tissues.
Correlations between aberrant expression of LINC00673 and TSCC clinical pathological parameters
In situ hybridization was used to evaluate correlation of LINC00673 expression level such as with patients’ clinicopathological parameters in TSCC, such as tumor size (T stage), lymph-vascular invasion (N stage), invasion muscles of tongue, pathological stage, relapse and so on. Representative images of LINC00673 signals are shown in Figure 3A. High LINC00673 expression was positively correlated with Tumor size (p = 0.034, Figure 3B), invasion muscles of tongue (p = 0.045, Figure 3C), higher TNM stage (p = 0.018, Figure 3D), and relapse (p = 0.002, Figure 3E). However, other clinical parameters, such as gender, age, smoking and chewing areca, were found not to be significantly correlated with LINC00673 expression in this study. The detailed results of clinical parameters and expressions were shown in Supplemental Table 1.
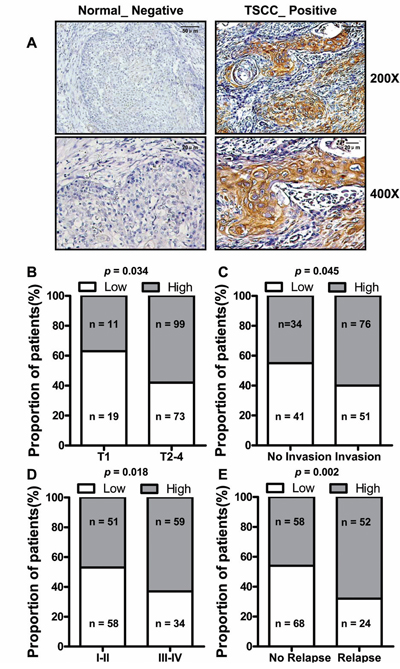
Figure 3: The relationship between LINC00673 and pathological clinical feature. A. LINC00673 expression was measured by in situ hybridization in paraffin embedded TSCC biopsies (T, n = 202). Representative cases of non-tumor lingual mucous membrane (N) and TSCC biopsies are shown. Proportion of TSCC patients with LINC00673 expression shown that upregulated LINC00673 was positively correlated with tumor size B., tongue muscle invasion C., TNM stage D. and relapse E.
High expression of LINC00673 predicts poor prognosis in TSCC
Additionally exploring of the correlation between LINC00673 expression and clinical outcomes demonstrated that, the median overall survival (OS) time was 39.5 and 29 months in TSCC patients with low and high expression of LINC00673, respectively (Figure 4A, p = 0.009), while the median relapse-free survival (RFS) time of TSCC patients with low and high expression of LINC00673 was 39 and 25 months, respectively (Figure 4B, p = 0.008). High expression of LINC00673 was associated with poor overall survival and poor relapse-free survival, and could be regarded as an independent predictor for overall survival in TSCC
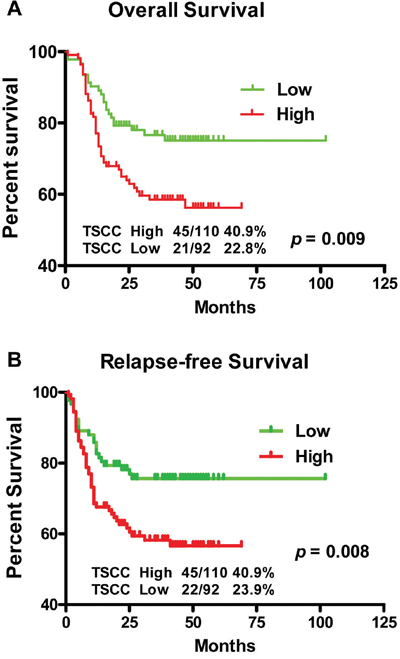
Figure 4: Kaplan-Meier survival curves in TSCC patients according to LINC00673 expression levels. The highly expressed LINC00673 was correlated with shorter overall survival A. or relapse free survival B. of TSCC patients.
Knockdown LINC00673 inhibits TSCC cell migration and invasion
To explored the function of LINC00673 in TSCC, we knockdown LINC00673 expression by using two LINC00673 targeting short interfering RNAs (siRNAs) in TSCC cell line, Tca8113 and Cal27. qRT-PCR results indicated that LINC00673 expression were significantly inhibited in both Tca8113 and Cal27 cells after siRNAs transfection (Figure 5). Wound healing assays demonstrated that the migratory potential of LINC00673-silenced cells were significantly reduced when compared with scrambled control siRNA treated (Figure 6). The transwell assay results showed that knockdown of LINC00673 inhibited Tca8113 and Cal27 cells invasion and compared to the control group (Figure 7).
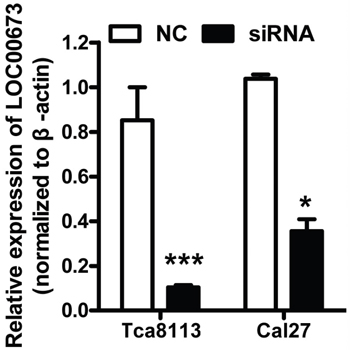
Figure 5: siRNAs successfully knockdowned LINC00673 expression. Tca8113 and Cal27 cells were transfected with siRNAs targeting LINC00673. LINC00673 expression was significantly decreased when measured by qRT-PCR.
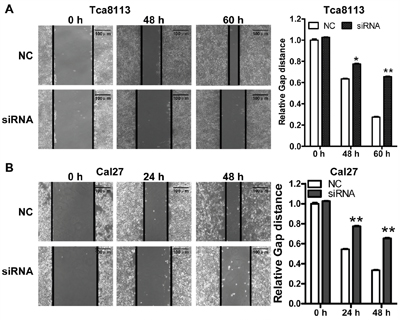
Figure 6: Knockdown LINC00673 suppressed TSCC tumor cell migration. Wound healing assays demonstrated that the migratory potential of Tca8113 A. and Cal27 B. cells were significantly reduced by siRNAs specifically targeting LINC00673 (siRNAs) when compared with scrambled control siRNA (NC) treated cells.
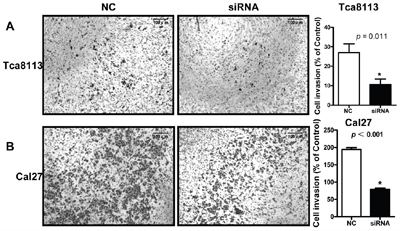
Figure 7: Knockdown LINC00673 suppressed TSCC tumor cell invasion. Transwell assay results showed that the invasion potential of LINC00673-silenced Tca8113 A. and Cal27 B. cells (siRNAs) were significantly reduced when compared with the control group (NC).
DISCUSSION
Recent reports indicate that lncRNAs play a critical role in a wide range of tumor progression such as differentiation, proliferation, and metastasis [32–34]. The pathogenesis and carcinogenesis of TSCC are multi-step and heterogeneous processes, and involve different genetic and epigenetic changes [35]. A number of lncRNAs have been discovered to play a vital role in carcinogenesis and progression of TSCC [36], including, HOTTIP [37], MALAT1 [38], TUC338 [39], UCA1 [40], etc. In the current study, we combined two previously published GEO dataset to identify differentially expressed lncRNAs in TSCC. We found 12 differentially expressed lncRNAs in TSCC tissues when compared with the noncancerous tissue samples. Among these 12 lncRNAs, LINC00152 had been reported that differentially expressed and participating in tumorigenesis in many tumors, was also included and upregulated in TSCC samples compared with the adjacent non-tumor tissues [41–43]; H19, another famous lncRNA, was also listed [25–31]. While other ten lncRNAs have not been well characterized. These dysregulated lncRNAs might have function in the carcinogensis of TSCC and more functional study is needed to identify valuable and effective lncRNAs-based diagnosis, prognosis and therapeutic biomarkers in TSCC.
In this study, we chose and focused on LINC00673 since it was significantly high express among lncRNAs dysregulated in TSCC but never been reported in TSCC previously. We performed qRT-PCR and in situ hybridization to verify our results using existing data in the GEO database. We found LINC00673 was upregulated in TSCC and it’s upregulation was associated with tumor progression, such as tumor size, invasion of muscles, clinical stages, recurrence and poor survival. After siRNA-mediated silencing of LINC00673, we found that could significantly inhibit TSCC cancer cell invasive and migratory abilities. In recent findings, some lncRNAs, such as actin filament associated protein 1 antisense RNA1 (AFAP1-AS1) [44, 45], HOX transcript antisense RNA (HOTAIR) [46], growth-arrest-specific transcript 5 (GAS5) [47], have been reported to regulate cancer metastasis process by multiple ways. The potential role of LINC00673 abnormalities in TSCC is worth for further investigated.
In conclusion, our findings indicate that LINC00673 might be involved in TSCC development, which may play a vital role in the tumorigenesis of TSCC, and may serve a useful biomarker for the prediction of TSCC prognosis and as potential target for therapy in the future.
MATERIALS AND METHODS
Data mining and analysis
Two independent cohorts of primary TSCC gene expression profiling (GEP) data based on the Affymetrix Human Genome U133 Plus 2.0 platform, GSE9844 [23] and GSE30784 [24], and their correlated clinic data were obtained from the Gene Expression Omnibus (GEO) database (http://www.ncbi.nlm.nih.gov/geo/). GSE9844 contain 26 cancer tissues and 12 normal adjacent tissues. GSE30784 has 167 cancer tissues and 45 normal adjacent tissues. Significant Analysis of Microarray (SAM) [48–51] software was used to analyze the different expression of lncRNAs between normal lingual mucous membrane and TSCC tissue samples in these two published TSCC datasets. The cut-off value for differentially expressed lncRNA was set at ≥ 1.5-fold change and the false discovery ratio (FDR) was < 0.05. The data analysis procedures are shown in Figure 1A.
Clinical samples
Two sets of clinical samples were collected for this study: Set 1 for qRT-PCR, containing 15 paired primary TSCC tissues and adjacent non-tumor lingual mucous membrane biopsies, collected from TSCC patients when they underwent surgical resection. None of the patients had received radiotherapy or chemotherapy prior to surgery. Set 2 for in situ hybridization, paraffin-embedded TSCC tissue samples from 202 cases of patients who underwent surgical operation from January 2009 to December 2013 at the Affiliated Cancer Hospital of Central South University. Clinical data were reported in Supplemental Table 1. This study was approved by the Ethical Committee at the Affiliated Cancer Hospital of Central South University and every patient provided the written informed consent.
RNA isolation and qRT-PCR
Total RNAs were extracted using TRIzol reagent (Invitrogen, Carlbad, CA, USA). One μg of total RNA from the samples was reverse transcribed using a Reverse Transcription Kit (Biorad, Hercules, CA, USA). Real-time PCR was performed using SYBRGreen (Biorad) in the CFX96 Real-Time PCR Detection System (Bio-Rad) [52–54]. The primers used were LINC00673: 5’-TTCTCC TGTAACGTGTGGCC-3’ and 5’-CTGGTGGGAATGTGGATCA GT-3’; ACTB (β-actin): 5’-TCACCAACTGGGACGACATG-3’ and 5’-GTCACCGGAGTC CATCACGAT -3’. ACTB was used as the reference and normalization control. LINC00673 expression was normalized to the respective ACTB expression level. Relative expression was calculated using the equation: ΔCt = Ct (target gene) – Ct (ACTB), fold expression = 2–(ΔCt(tumor) – ΔCt(normal)).
In situ hybridization
In situ hybridization was performed to detect LINC00673 expression as previously described [55–57]. Three probes from different LINC00673 regions (5’-GAAAAACCTCTTG CACCACCTTAGTCTCCAAAGA-3’, 5’-CTTTCCTGTTCTTTCTCCTACCCTTCCTGAC TAG -3’, and 5’-CATGAAGTAATAATAAAGGTTCCGCTTATCAACC-3’) were synthesized and labeled with DIG-dUTP at the 3’ end (Invitrogen, Shanghai, China). Three GAPDH probes used as positive controls were 5’-CCACTTTACCAGAGTTAAAAGCAGCCCTGG-3’, 5’-CAGTAGAGGCAGGGATGATGTTCTGGAGAG-3’, and 5’-GTCAGAGGAGACC ACCTGGTGCTCAGTGTA-3’. A semi-quantitative scoring criterion for in situ hybridization was used in which both the staining intensity and the number of positive cells were recorded. The scoring was graded as 0 (negative), 1 (< 10% positive), 2 (10% - 50% positive), or 3 (> 50% positive) in accordance with the staining proportion and intensity. The final scores were regarded as low expression (0-1) and high expression (2-3). All sections were scored in randomly selected five representative fields of vision at medium magnification by two pathologists who were blinded to the clinicopathological features and the clinical data.
Cell line and gene silencing
The TSCC cell lines Tca8113 and Cal27 were maintained in an atmosphere of 5 % CO2 at 37°C and cultured in RPMI 1640 medium supplemented with 1 % antibiotics (100U/ml penicillin and100 μg/ml streptomycin sulfates) and 10 % fetal bovine serum (FBS, GIBCO).
For gene knockdown, cells were seed in six-well plates to confluency and transfected siRNAs by using Lipofectamine RNAiMAX Reagent (Invitrogen) in OptiMEM medium (Invitrogen) [58–61]. Two sequences of LINC00673 targeting siRNAs were: 5’-CCAGTTGTC CTTGACTGCATGGTTT-3’; and 5’-AGGGAACCACAGGATTCCATGTGAT-3’. Sequences of non-target scramble controls were provided by Invitrogen.
Cell migration and invasion assay
Wound healing assay was used to examine TSCC cells invasion capacity. The cells were seeded in six-well culture plates and grown to 90 % confluence. Vertical wounds in the cell monolayer were created by a 10 ul tip and washed three times with PBS to remove cell debris. Wound width was measured by microscopy at the designated time periods [62–65].
Transwell assay was used to assess tumor cell migration capacity. A total of 1×105 cells in 100 μl of serum-free medium were added to the top of transwell Cell Culture chambers (8 μm pore size, BD Biosciences, New Jersey, USA), and 600 μl of 10% FBS containing medium was added to the lower chamber. Cells were incubated for 36 h at 37 °C, and then, the migrated cells were fixed with methanol, stained with 0.5% crystal violet. Cells on the upper surface were wiped by a cotton bud. Numbers of invasive cells were counted from six randomly selected 200× fields in under a microscope and shown as the average per field [66–68].
Statistical analysis
All experiments were independently repeated at least triplicate. All the Statistical analysis was carried out by using SPSS software, version 19.0 (SPSS, Chicago, IL, USA) and presented with Graph-pad prism software. Differences between two independent groups were evaluated by Student’s t-tests and differences for multiple comparisons were evaluate by one way ANOVA. Overall survival (OS) or relapse-free survival (RFS) were calculated using the Kaplan-Meier method, and the results of the analysis were considered significant in a log-rank test if p < 0.05. All data are represented as means ± standard deviation. A two-tailed p value of 0.05 or less was considered statistically significant.
Abbreviations
lncRNA, long non-coding RNA; TSCC, tongue squamous cell carcinoma; LINC00673, long intergenic non-coding RNA 673; qRT-PCR, quantitative real time polymerase chain reaction; GEP, gene expression profiling; GEO, Gene Expression Omnibus; SAM, Significant Analysis of Microarray; FDR, false discovery ratio; OS, Overall survival; RFS, relapse-free survival; siRNA, short interfering RNA; AFAP1-AS1, actin filament associated protein 1 antisense RNA1; HOTAIR, HOX transcript antisense RNA; GAS5, growth-arrest-specific transcript 5.
CONFLICTS OF INTEREST
The authors have no conflicts of interest to declare.
FUNDING
This study was supported in part by grants from the National Natural Science Foundation of China (81372907, 81301757, 81472531, 81402009, 81572787, and 81672683) and the Natural Science Foundation of Hunan Province (14JJ1010 and 2015JJ1022).
Author contributions
W.X. and Z.Z. designed the research, analyzed and interpreted the data, and drafted the manuscript; J.Y., and Y.L. performed the research, contributed clinical samples, analyzed and interpreted the data, and drafted the manuscript; Z.G., S.Z., G.C., X.L., Y.T., L.Y., Y.H., F.W., Y.W., Q.L., W.Z., X.L., Y.L. and G.L. performed the research, analyzed and interpreted the data.
REFERENCES
1. Parkin DM, Bray F, Ferlay J, Pisani P. Global cancer statistics, 2002. CA Cancer J Clin. 2005; 55: 74-108. doi: 10.3322/canjclin.55.2.74.
2. Zeng ZY, Zhou YH, Zhang WL, Li XL, Xiong W, Liu HY, Fan SQ, Qian J, Wang LL, Li Z, Shen SR, Li GY. Family-based association analysis validates chromosome 3p21 as a putative nasopharyngeal carcinoma susceptibility locus. Genetics in Medicine. 2006; 8: 156-60. doi: 10.1097/01.gim.0000196821.87655.d0.
3. Xiong W, Zeng ZY, Xia JH, Shen SR, Li XL, Hu DX, Tan C, Xiang JJ, Zhou J, Deng H, Fan SQ, Li WF, Wang R, et al. A susceptibility locus at chromosome 3p21 linked to familial nasopharyngeal carcinoma. Cancer Research. 2004; 64: 1972-4. doi: 10.1158/0008-5472.Can-03-3253.
4. Argiris A, Eng C. Epidemiology, staging, and screening of head and neck cancer. Cancer Treat Res. 2003; 114: 15-60. doi: 10.1007/0-306-48060-3_2.
5. Mignogna MD, Fedele S, Lo Russo L. The World Cancer Report and the burden of oral cancer. Eur J Cancer Prev. 2004; 13: 139-42. doi: 10.1097/01.cej.0000124501.37552.29.
6. Kimple AJ, Welch CM, Zevallos JP, Patel SN. Oral cavity squamous cell carcinoma—an overview. Oral Health Dent Manag. 2014; 13: 877-82.
7. Warnakulasuriya S. Global epidemiology of oral and oropharyngeal cancer. Oral Oncol. 2009; 45: 309-16. doi: 10.1016/j.oraloncology.2008.06.002.
8. Liu X, Chen Z, Yu J, Xia J, Zhou X. MicroRNA profiling and head and neck cancer. Comp Funct Genomics. 2009: 837514. doi: 10.1155/2009/837514.
9. Pulte D, Brenner H. Changes in survival in head and neck cancers in the late 20th and early 21st century: a period analysis. Oncologist. 2010; 15: 994-1001. doi: 10.1634/theoncologist.2009-0289.
10. Tu CF, Qi P, Li XY, Mo YZ, Li XL, Xiong W, Zeng ZY, Li GY. Tumor Heterogeneity: The Challenge of Precision Medicine. Progress in Biochemistry and Biophysics. 2015; 42: 881-90. doi: 10.16476/j.pibb.2015.0214.
11. Gong Z, Zhang S, Zhang W, Huang H, Li Q, Deng H, Ma J, Zhou M, Xiang J, Wu M, Li X, Xiong W, Li X, et al. Long non-coding RNAs in cancer. Sci China Life Sci. 2012; 55: 1120-4. doi: 10.1007/s11427-012-4413-9.
12. Lian Y, Li XY, Tang YY, Yang LT, Li XL, Xiong W, Li GY, Zeng ZY. Long Non-coding RNAs Function as Competing Endogenous RNAs to Regulate Cancer Progression. Progress in Biochemistry and Biophysics. 2016; 43: 219-25. doi: 10.16476/j.pibb.2015.0371.
13. Wang Y, Xue D, Li Y, Pan X, Zhang X, Kuang B, Zhou M, Li X, Xiong W, Li G, Zeng Z, Yang T. The Long Noncoding RNA MALAT-1 is A Novel Biomarker in Various Cancers: A Meta-analysis Based on the GEO Database and Literature. J Cancer. 2016; 7: 991-1001. doi: 10.7150/jca.14663.
14. Gong Z, Yang Q, Zeng Z, Zhang W, Li X, Zu X, Deng H, Chen P, Liao Q, Xiang B, Zhou M, Li X, Li Y, et al. An integrative transcriptomic analysis reveals p53 regulated miRNA, mRNA, and lncRNA networks in nasopharyngeal carcinoma. Tumour Biol. 2016; 37: 3683-95. doi: 10.1007/s13277-015-4156-x.
15. Li YW, Wang YM, Zhang XY, Xue D, Kuang B, Pan XY, Jing YZ, Li XL, Zhou M, Xiong W, Zeng ZY, Li GY. Progress of Long Noncoding RNA HOTAIR in Human Cancer. Progress in Biochemistry and Biophysics. 2015; 42: 228-35. doi: 10.16476/j.pibb.2014.0230.
16. Zeng Z, Huang H, Huang L, Sun M, Yan Q, Song Y, Wei F, Bo H, Gong Z, Zeng Y, Li Q, Zhang W, Li X, et al. Regulation network and expression profiles of Epstein-Barr virus-encoded microRNAs and their potential target host genes in nasopharyngeal carcinomas. Sci China Life Sci. 2014; 57: 315-26. doi: 10.1007/s11427-013-4577-y.
17. Tang K, Wei F, Bo H, Huang HB, Zhang WL, Gong ZJ, Li XY, Song YL, Liao QJ, Peng SP, Xiang JJ, Zhou M, Ma J, et al. Cloning and Functional Characterization of a Novel Long Non-coding RNA Gene Associated With Hepatocellular Carcinoma. Progress in Biochemistry and Biophysics. 2014; 41: 153-62. doi: 10.3724/Sp.J.1206.2012.00613.
18. Gong Z, Zhang S, Zeng Z, Wu H, Yang Q, Xiong F, Shi L, Yang J, Zhang W, Zhou Y, Zeng Y, Li X, Xiang B, et al. LOC401317, a p53-regulated long non-coding RNA, inhibits cell proliferation and induces apoptosis in the nasopharyngeal carcinoma cell line HNE2. PLoS One. 2014; 9: e110674. doi: 10.1371/journal.pone.0110674.
19. Zhang W, Huang C, Gong Z, Zhao Y, Tang K, Li X, Fan S, Shi L, Li X, Zhang P, Zhou Y, Huang D, Liang F, et al. Expression of LINC00312, a long intergenic non-coding RNA, is negatively correlated with tumor size but positively correlated with lymph node metastasis in nasopharyngeal carcinoma. J Mol Histol. 2013; 44: 545-54. doi: 10.1007/s10735-013-9503-x.
20. Bo H, Gong Z, Zhang W, Li X, Zeng Y, Liao Q, Chen P, Shi L, Lian Y, Jing Y, Tang K, Li Z, Zhou Y, et al. Upregulated long non-coding RNA AFAP1-AS1 expression is associated with progression and poor prognosis of nasopharyngeal carcinoma. Oncotarget. 2015; 6: 20404-18. doi: 10.18632/oncotarget.4057.
21. Sun Q, Liu H, Li L, Zhang S, Liu K, Liu Y, Yang C. Long noncoding RNA-LET, which is repressed by EZH2, inhibits cell proliferation and induces apoptosis of nasopharyngeal carcinoma cell. Med Oncol. 2015; 32: 226. doi: 10.1007/s12032-015-0673-0.
22. Zou AE, Ku J, Honda TK, Yu V, Kuo SZ, Zheng H, Xuan Y, Saad MA, Hinton A, Brumund KT, Lin JH, Wang-Rodriguez J, Ongkeko WM. Transcriptome sequencing uncovers novel long noncoding and small nucleolar RNAs dysregulated in head and neck squamous cell carcinoma. RNA. 2015; 21: 1122-34. doi: 10.1261/rna.049262.114.
23. Ye H, Yu T, Temam S, Ziober BL, Wang J, Schwartz JL, Mao L, Wong DT, Zhou X. Transcriptomic dissection of tongue squamous cell carcinoma. BMC Genomics. 2008; 9: 69. doi: 10.1186/1471-2164-9-69.
24. Chen C, Mendez E, Houck J, Fan W, Lohavanichbutr P, Doody D, Yueh B, Futran ND, Upton M, Farwell DG, Schwartz SM, Zhao LP. Gene expression profiling identifies genes predictive of oral squamous cell carcinoma. Cancer Epidemiol Biomarkers Prev. 2008; 17: 2152-62. doi: 10.1158/1055-9965.EPI-07-2893.
25. Li H, Li J, Jia S, Wu M, An J, Zheng Q, Zhang W, Lu D. miR675 upregulates long noncoding RNA H19 through activating EGR1 in human liver cancer. Oncotarget. 2015; 6: 31958-84. doi: 10.18632/oncotarget.5579.
26. Liang WC, Fu WM, Wong CW, Wang Y, Wang WM, Hu GX, Zhang L, Xiao LJ, Wan DC, Zhang JF, Waye MM. The lncRNA H19 promotes epithelial to mesenchymal transition by functioning as miRNA sponges in colorectal cancer. Oncotarget. 2015; 6: 22513-25. doi: 10.18632/oncotarget.4154.
27. Vennin C, Spruyt N, Dahmani F, Julien S, Bertucci F, Finetti P, Chassat T, Bourette RP, Le Bourhis X, Adriaenssens E. H19 non coding RNA-derived miR-675 enhances tumorigenesis and metastasis of breast cancer cells by downregulating c-Cbl and Cbl-b. Oncotarget. 2015; 6: 29209-23. doi: 10.18632/oncotarget.4976.
28. Yang C, Tang R, Ma X, Wang Y, Luo D, Xu Z, Zhu Y, Yang L. Tag SNPs in long non-coding RNA H19 contribute to susceptibility to gastric cancer in the Chinese Han population. Oncotarget. 2015; 6: 15311-20. doi: 10.18632/oncotarget.3840.
29. Han D, Gao X, Wang M, Qiao Y, Xu Y, Yang J, Dong N, He J, Sun Q, Lv G, Xu C, Tao J, Ma N. Long noncoding RNA H19 indicates a poor prognosis of colorectal cancer and promotes tumor growth by recruiting and binding to eIF4A3. Oncotarget. 2016; 7: 22159-73. doi: 10.18632/oncotarget.8063.
30. Li S, Hua Y, Jin J, Wang H, Du M, Zhu L, Chu H, Zhang Z, Wang M. Association of genetic variants in lncRNA H19 with risk of colorectal cancer in a Chinese population. Oncotarget. 2016; 7: 25470-7. doi: 10.18632/oncotarget.8330.
31. Wu T, Qu L, He G, Tian L, Li L, Zhou H, Jin Q, Ren J, Wang Y, Wang J, Kan X, Liu M, Shen J, et al. Regulation of laryngeal squamous cell cancer progression by the lncRNA H19/miR-148a-3p/DNMT1 axis. Oncotarget. 2016; 7: 11553-66. doi: 10.18632/oncotarget.7270.
32. Zhang K, Sun X, Zhou X, Han L, Chen L, Shi Z, Zhang A, Ye M, Wang Q, Liu C, Wei J, Ren Y, Yang J, et al. Long non-coding RNA HOTAIR promotes glioblastoma cell cycle progression in an EZH2 dependent manner. Oncotarget. 2015; 6: 537-46. doi: 10.18632/oncotarget.2681.
33. Zeng Z, Bo H, Gong Z, Lian Y, Li X, Li X, Zhang W, Deng H, Zhou M, Peng S, Li G, Xiong W. AFAP1-AS1, a long noncoding RNA upregulated in lung cancer and promotes invasion and metastasis. Tumour Biol. 2016; 37: 729-37. doi: 10.1007/s13277-015-3860-x.
34. Zhao J, Liu Y, Zhang W, Zhou Z, Wu J, Cui P, Zhang Y, Huang G. Long non-coding RNA Linc00152 is involved in cell cycle arrest, apoptosis, epithelial to mesenchymal transition, cell migration and invasion in gastric cancer. Cell Cycle. 2015; 14: 3112-23. doi: 10.1080/15384101.2015.1078034.
35. Leemans CR, Braakhuis BJ, Brakenhoff RH. The molecular biology of head and neck cancer. Nat Rev Cancer. 2011; 11: 9-22. doi: 10.1038/nrc2982.
36. Gao W, Chan JY, Wong TS. Long non-coding RNA deregulation in tongue squamous cell carcinoma. Biomed Res Int. 2014; 2014: 405860. doi: 10.1155/2014/405860.
37. Zhang H, Zhao L, Wang YX, Xi M, Liu SL, Luo LL. Long non-coding RNA HOTTIP is correlated with progression and prognosis in tongue squamous cell carcinoma. Tumour Biol. 2015; 36: 8805-9. doi: 10.1007/s13277-015-3645-2.
38. Liang J, Liang L, Ouyang K, Li Z, Yi X. MALAT1 induces tongue cancer cells’ EMT and inhibits apoptosis through Wnt/beta-catenin signaling pathway. J Oral Pathol Med. 2017; 46: 98-105. doi: 10.1111/jop.12466.
39. Ouyang KX, Zou R, Liang J, Bai ZB, Li ZQ, Zhao JJ. TUC338 Overexpression Leads to Enhanced Proliferation and Reduced Apoptosis in Tongue Squamous Cell Carcinoma Cells
40. Fang Z, Wu L, Wang L, Yang Y, Meng Y, Yang H. Increased expression of the long non-coding RNA UCA1 in tongue squamous cell carcinomas: a possible correlation with cancer metastasis. Oral Surg Oral Med Oral Pathol Oral Radiol. 2014; 117: 89-95. doi: 10.1016/j.oooo.2013.09.007.
41. Ji J, Tang J, Deng L, Xie Y, Jiang R, Li G, Sun B. LINC00152 promotes proliferation in hepatocellular carcinoma by targeting EpCAM via the mTOR signaling pathway. Oncotarget. 2015; 6: 42813-24. doi: 10.18632/oncotarget.5970.
42. Pang Q, Ge J, Shao Y, Sun W, Song H, Xia T, Xiao B, Guo J. Increased expression of long intergenic non-coding RNA LINC00152 in gastric cancer and its clinical significance. Tumour Biol. 2014; 35: 5441-7. doi: 10.1007/s13277-014-1709-3.
43. Yu J, Liu Y, Guo C, Zhang S, Gong Z, Tang Y, Yang L, He Y, Lian Y, Li X, Deng H, Liao Q, Li X, et al. Upregulated long non-coding RNA LINC00152 expression is associated with progression and poor prognosis of tongue squamous cell carcinoma. J Cancer. 2017. doi: 10.7150/jca.17510.
44. Han X, Wang L, Ning Y, Li S, Wang Z. Long non-coding RNA AFAP1-AS1 facilitates tumor growth and promotes metastasis in colorectal cancer. Biol Res. 2016; 49: 36. doi: 10.1186/s40659-016-0094-3.
45. Wang Y, Mo Y, Gong Z, Yang X, Yang M, Zhang S, Xiong F, Xiang B, Zhou M, Liao Q, Zhang W, Li X, Li X, et al. Circular RNAs in human cancer. Mol Cancer. 2017; 16: 25. doi: 10.1186/s12943-017-0598-7.
46. Lee M, Kim HJ, Kim SW, Park SA, Chun KH, Cho NH, Song YS, Kim YT. The long non-coding RNA HOTAIR increases tumour growth and invasion in cervical cancer by targeting the Notch pathway. Oncotarget. 2016; 7: 44558-71. doi: 10.18632/oncotarget.10065.
47. Chang L, Li C, Lan T, Wu L, Yuan Y, Liu Q, Liu Z. Decreased expression of long non-coding RNA GAS5 indicates a poor prognosis and promotes cell proliferation and invasion in hepatocellular carcinoma by regulating vimentin. Mol Med Rep. 2016; 13: 1541-50. doi: 10.3892/mmr.2015.4716.
48. Huang HB, Liang F, Xiong W, Li XL, Zeng ZY, Li GY. Bioinformatics Accelerates Drug Repositioning. Progress in Biochemistry and Biophysics. 2012; 39: 35-44. doi: 10.3724/Sp.J.1206.2011.00453.
49. Gong ZJ, Huang HB, Xu K, Liang F, Li XL, Xiong W, Zeng ZY, Li GY. Advances in microRNAs and TP53 Gene Regulatory Network. Progress in Biochemistry and Biophysics. 2012; 39: 1133-44. doi: 10.3724/Sp.J.1206.2012.00015.
50. Zeng Z, Huang H, Zhang W, Xiang B, Zhou M, Zhou Y, Ma J, Yi M, Li X, Li X, Xiong W, Li G. Nasopharyngeal carcinoma: advances in genomics and molecular genetics. Sci China Life Sci. 2011; 54: 966-75. doi: 10.1007/s11427-011-4223-5.
51. Zhang W, Zeng Z, Zhou Y, Xiong W, Fan S, Xiao L, Huang D, Li Z, Li D, Wu M, Li X, Shen S, Wang R, et al. Identification of aberrant cell cycle regulation in Epstein-Barr virus-associated nasopharyngeal carcinoma by cDNA microarray and gene set enrichment analysis. Acta Biochim Biophys Sin. 2009; 41: 414-28. doi: 10.1093/abbs/gmp025.
52. Zhou Y, Liao Q, Li X, Wang H, Wei F, Chen J, Yang J, Zeng Z, Guo X, Chen P, Zhang W, Tang K, Li X, et al. HYOU1, Regulated by LPLUNC1, Is Up-Regulated in Nasopharyngeal Carcinoma and Associated with Poor Prognosis. J Cancer. 2016; 7: 367-76. doi: 10.7150/jca.13695.
53. Xu K, Xiong W, Zhou M, Wang H, Yang J, Li X, Chen P, Liao Q, Deng H, Li X, Li G, Zeng Z. Integrating ChIP-sequencing and digital gene expression profiling to identify BRD7 downstream genes and construct their regulating network. Mol Cell Biochem. 2016; 411: 57-71. doi: 10.1007/s11010-015-2568-y.
54. Xiao K, Yu Z, Li X, Li X, Tang K, Tu C, Qi P, Liao Q, Chen P, Zeng Z, Li G, Xiong W. Genome-wide Analysis of Epstein-Barr Virus (EBV) Integration and Strain in C666-1 and Raji Cells. J Cancer. 2016; 7: 214-24. doi: 10.7150/jca.13150.
55. Zeng ZY, Zhou YH, Zhang WL, Xiong W, Fan SQ, Li XL, Luo XM, Wu MH, Yang YX, Huang C, Cao L, Tang K, Qian J, et al. Gene expression profiling of nasopharyngeal carcinoma reveals the abnormally regulated Wnt signaling pathway. Hum Pathol. 2007; 38: 120-33. doi: 10.1016/j.humpath.2006.06.023.
56. Zeng Z, Fan S, Zhang X, Li S, Zhou M, Xiong W, Tan M, Zhang W, Li G. Epstein-Barr virus-encoded small RNA 1 (EBER-1) could predict good prognosis in nasopharyngeal carcinoma. Clin Transl Oncol. 2016; 18: 206-11. doi: 10.1007/s12094-015-1354-3.
57. Zeng Z, Zhou Y, Xiong W, Luo X, Zhang W, Li X, Fan S, Cao L, Tang K, Wu M, Li G. Analysis of gene expression identifies candidate molecular markers in nasopharyngeal carcinoma using microdissection and cDNA microarray. J Cancer Res Clin Oncol. 2007; 133: 71-81. doi: 10.1007/s00432-006-0136-2.
58. Song Y, Li X, Zeng Z, Li Q, Gong Z, Liao Q, Li X, Chen P, Xiang B, Zhang W, Xiong F, Zhou Y, Zhou M, et al. Epstein-Barr virus encoded miR-BART11 promotes inflammation-induced carcinogenesis by targeting FOXP1. Oncotarget. 2016; 7: 36783-99. doi: 10.18632/oncotarget.9170.
59. Liang F, Li Q, Li X, Li Z, Gong Z, Deng H, Xiang B, Zhou M, Li X, Li G, Zeng Z, Xiong W. TSC22D2 interacts with PKM2 and inhibits cell growth in colorectal cancer. Int J Oncol. 2016; 49: 1046-56. doi: 10.3892/ijo.2016.3599.
60. He B, Li W, Wu Y, Wei F, Gong Z, Bo H, Wang Y, Li X, Xiang B, Guo C, Liao Q, Chen P, Zu X, et al. Epstein-Barr virus-encoded miR-BART6-3p inhibits cancer cell metastasis and invasion by targeting long non-coding RNA LOC553103. Cell Death Dis. 2016; 7: e2353. doi: 10.1038/cddis.2016.253.
61. Yan Q, Zeng Z, Gong Z, Zhang W, Li X, He B, Song Y, Li Q, Zeng Y, Liao Q, Chen P, Shi L, Fan S, et al. EBV-miR-BART10-3p facilitates epithelial-mesenchymal transition and promotes metastasis of nasopharyngeal carcinoma by targeting BTRC. Oncotarget. 2015; 6: 41766-82. doi: 10.18632/oncotarget.6155.
62. Duan Z, Zheng H, Xu S, Jiang Y, Liu H, Li M, Hu D, Li W, Bode AM, Dong Z, Cao Y. Activation of the Ig Ialpha1 promoter by the transcription factor Ets-1 triggers Ig Ialpha1-Calpha1 germline transcription in epithelial cancer cells. Cell Mol Immunol. 2014; 11: 197-205. doi: 10.1038/cmi.2013.52.
63. Hu D, Duan Z, Li M, Jiang Y, Liu H, Zheng H, Li L, Bode AM, Dong Z, Cao Y. Heterogeneity of aberrant immunoglobulin expression in cancer cells. Cell Mol Immunol. 2011; 8: 479-85. doi: 10.1038/cmi.2011.25.
64. Hu D, Zheng H, Liu H, Li M, Ren W, Liao W, Duan Z, Li L, Cao Y. Immunoglobulin expression and its biological significance in cancer cells. Cell Mol Immunol. 2008; 5: 319-24. doi: 10.1038/cmi.2008.39.
65. Li M, Zheng H, Duan Z, Liu H, Hu D, Bode A, Dong Z, Cao Y. Promotion of cell proliferation and inhibition of ADCC by cancerous immunoglobulin expressed in cancer cell lines. Cell Mol Immunol. 2012; 9: 54-61. doi: 10.1038/cmi.2011.40.
66. Yang Y, Zhou H, Yang Y, Li W, Zhou M, Zeng Z, Xiong W, Wu M, Huang H, Zhou Y, Peng C, Huang C, Li X, et al. Lipopolysaccharide (LPS) regulates TLR4 signal transduction in nasopharynx epithelial cell line 5-8F via NFkappaB and MAPKs signaling pathways. Mol Immunol. 2007; 44: 984-92. doi: 10.1016/j.molimm.2006.03.013.
67. Zheng H, Li M, Ren W, Zeng L, Liu HD, Hu D, Deng X, Tang M, Shi Y, Gong J, Cao Y. Expression and secretion of immunoglobulin alpha heavy chain with diverse VDJ recombinations by human epithelial cancer cells. Mol Immunol. 2007; 44: 2221-7. doi: 10.1016/j.molimm.2006.11.010.
68. Zhao R, Liu Y, Wang H, Yang J, Niu W, Fan S, Xiong W, Ma J, Li X, Phillips JB, Tan M, Qiu Y, Li G, et al. BRD7 plays an anti-inflammatory role during early acute inflammation by inhibiting activation of the NF-small ka, CyrillicB signaling pathway. Cell Mol Immunol. 2016. doi: 10.1038/cmi.2016.31.

