INTRODUCTION
Lung carcinoma is the top killer among cancers, claiming 1.4 million lives in the world yearly. Non-small-cell lungcarcinoma (NSCLC) is found in about 80% of primary lungcancer patients, who are usually diagnosed in advanced stage despite current efforts and improvements aimed at early diagnosis [1]. Although a variety of therapeutic options are available for lung cancer patients, e.g. surgery, chemotherapy and radiotherapy, five-year survival ratesremain critically low. In patients with epidermal growth factor receptor (EGFR) activatingmutation, treatment with EGFR tyrosine kinase inhibitors (TKIs), e.g. gefitinib, shows high efficacy [2]. However, acquired TKI resistance hampers the use of such molecules. Increasing evidence suggests that miRNAs may significantly affect the development and chemoresistance of lung cancer [3–6].
MiRNAs are endogenous single-stranded non-coding RNAsassociated with various types of cancer [7]. They have essential functions in gene regulation, andeffect many important pathophysiological processes such as differentiation, development and tumorigenesis [8–10]. MiRNAs mainly bind to the 3’-untranslated region (3’-UTR) of mRNA molecules, suppressing protein synthesis through mRNA degradation or translational repression [11]. MiR-30ewas described as a tumor suppressor gene in various cancers [12–14]. Here, we found that overexpressionof miR-30einlung cancer cells resulted inreduced cell proliferation and migration, reversing drug resistance to gefitinib. Known miR-30e targets include Bmi1, P4HA1, BCR-ABL and UBC9 [12–15]. However, how miR-30eregulates lung cancer tumorigenesisremains unclear.
HOX genes belong to a highly conserved subgroup of theHomeobox superfamily which characteristically encodea 60-amino acid long DNA-binding motif. HOX genes play critical roles in regulating cell fate as well as early developmental events and organogenesis [16–18]. Alterations in HOX genes are also associated with multiple cancers in humans, e.g. lung, breast, and hematological cancers [19–21]. Here, we demonstrated that miR-30e targetedHOXA1, whose expression was reduced in miR-30e treated cells and conversely enhanced after miR-30e inhibition.
It has been reported that miR-30e may play important roles in cancerby affecting different signaling pathways. The current data revealed that miR-30eamounts were reduced in PC9Gcells in comparison with PC9 cells. We further characterized miR-30e and explore its molecular mechanisms in lung cancer. Interestingly, ectopic expression of miR-30e resulted reduced cell proliferation andmigration, withinduced apoptosisin lung cancer cells by suppressing the key target HOXA1. In addition, miR-30e rendered PC9G cells more sensitive to gefitinib in vitro and in vivo. These findings revealed a novel mechanism for miR-30e, indicating that this miRNA could be further assessed for the development of lung cancer therapeutics.
RESULTS
MicroRNA-30e is markedly downregulated in lung carcinoma
To assessthe role of miR-30e in lung cancer, miR-30e amounts were assessed in 30 lung cancer tissues with the corresponding adjacent normal tissuespecimens. RT-qPCR showed that miR-30e was significant downregulated in lung cancer tissues (Figure 1A). Then, human lung cancer specimens were divided into 2 groups based on sensitivity of gefitinib; interestingly, loweramounts of miR-30ewere obtained in lung cancer patients with gefitinib resistance (Figure 1B). To evaluate the effect of miR-30e on NSCLC patientprognosis after treatment with EGFR-TKI, the Kaplan-Meier method and log-rank test were used to examine normalized miR-30e levelsand Disease free survival (DFS). Individuals displaying low miR-30e levels showed reduced DFS as well asoverall survival (OS) compared with thosedisplaying elevated miR-30e amounts (Figure 1C, 1D). Thesefindings suggested that loss of miR-30e may be associated with lung cancer disease progression, and should be considered a potential new biomarker for predicting poor prognosis in NSCLC.
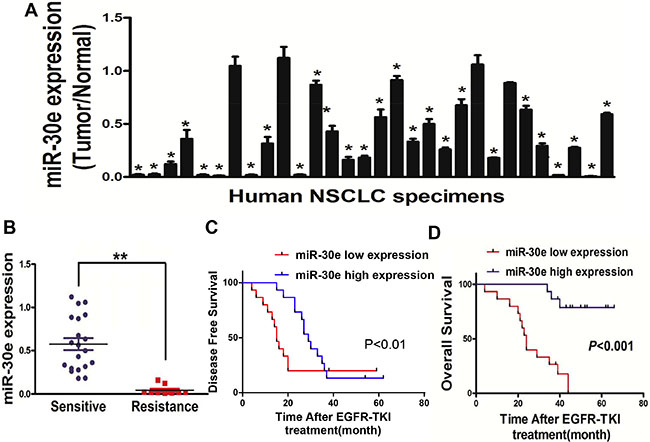
Figure 1: MiR-30e expression is markedlyreduced in lung cancer. (A) Relative miR-30e amounts analyzed by qRT-PCR in 30 human lung cancer tissues alongside adjacent non-cancerous specimens, with U6 employed for normalization. (B) MicroRNA-30e amounts werereduced in lung cancer patients resistant to gefitinib. (C, D) Kaplan-Meier curves showingdisease-free-and overallsurvival based on miR-30e amounts. High and low miR-30e amounts were defined based on the 50th percentile value. Data represent mean ± SD of three replicates. * and ** indicate significant differences at P < 0.05 and P < 0.01, respectively.
PC9G cells shows higher activity of proliferation and migration, and lower apoptosis rates compared with PC9 cells
Gefitinib-based chemotherapy is considered the cornerstone in treating advanced lung cancer. To mimic long-time exposure of patients to gefitinib, an in vitro model was established by transforming human lung cancer PC9 cells via exposure to lower concentrations ofgefitinib for 24 weeks (Figure 2A). Interestingly, miR-30e amounts in PC9 cells wereelevated compared with values obtained in theresistant PC9Gcell line (Figure 2B). PC9G cells had resistancefeatures, including enhanced cell proliferation andmigration, alongside lower apoptosis rates (Figure 2C–2E). In addition, we found that PC9G showedincreased cell proliferation andmigration, with reduced apoptosis compared with PC9 cells.
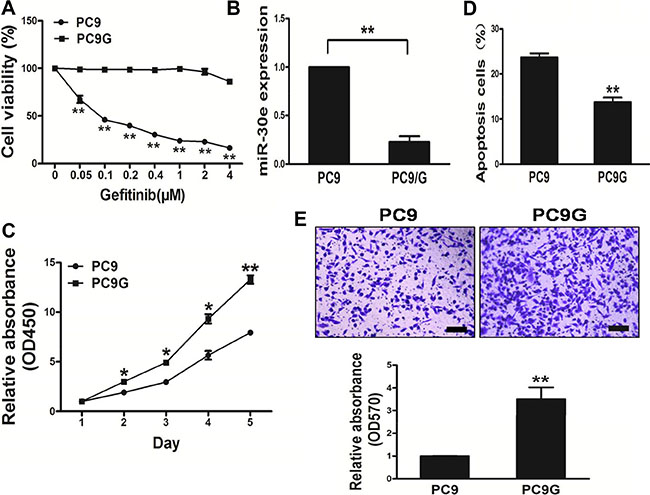
Figure 2: PC9Gcells show enhanced proliferation andmigration, and reduced apoptosis compared with PC9 cells. (A) Compared with PC9 cells, PC9G cells displayed reduced sensitivity to gefitinib. (B) MicroRNA-30e expression in PC9 and PC9G cells. (C) The CCK8 assay was used to determine cell viability of PC9 and PC9Gat various time points. (D) Transwell migration assays was conducted for respective cells. (E) Apoptosis assay was performed for PC9 and PC9G cells. Data are mean ± SDof 3 replicate experiments. * and ** indicate significant differences at P < 0.05 and P < 0.01, respectively.
High miR-30e levels in PC9G cells inhibit cancer aggressiveness and reverses drug resistance to gefitinib
MiR-30e amounts in resistant PC9G cells were lower thanin PC9 cells. Interestingly, cell growth was reduced in miR-30e-overexpressing lung cancer cells in comparison with those transfected with miR-NC (Figure 3A and 3B). We next assessed the impact of miR-30e on cell migration. As shown in Figure 3C, miR-30e re-expression starkly reduced the migrationability of lung cancer cells. What’s more, overexpressionof miR-30e promoted cell apoptosis (Figure 3D). We further found that miR-30eoverexpression reversed drug resistance to gefitinib in PC9G cells (Figure 3E). Thus, our results suggest miR-30e inhibited tumor aggressiveness, and reversed drug resistance to gefitinib.
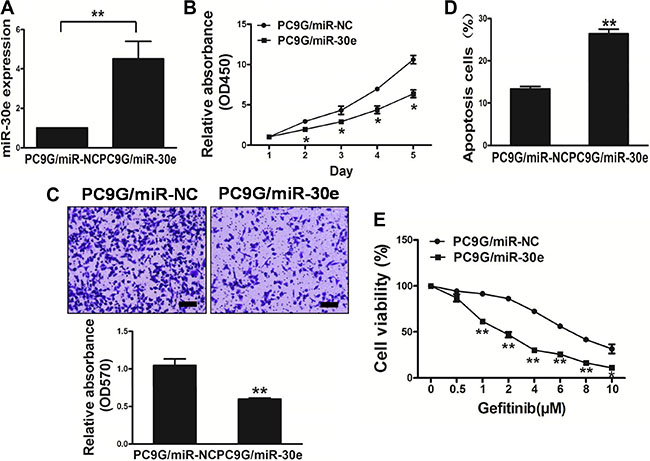
Figure 3: MicroRNA-30e overexpression in the PC9G cell linereduces cell proliferation andmigration, and reverses drug resistance to gefitinib. (A) Real-time PCR quantifying miR-30e amounts in PC9G cells. (B) The CCK8 assay was used to quantitate cell viability after transduction with miR-30e or miR-NC. (C) Transwell migration assays were conducted for respective cells. (D) Apoptosis Assay was carried outfor respective cells. (E) Gefitinib sensitivity of the PC9G/miR-NC and PC9G/miR-30e cell lines was tested by CCK-8 assay. Data are mean ± SD of 3 replicate experiments. * and ** indicate significant differences at P < 0.05 and P < 0.01, respectively.
Repression of miR-30e in PC9 cells significantly promotes cell growth and migration, also conferring resistance to gefitinib
To evaluate miR-30e function in lung cancer carcinogenesis, PC9 cell growthwas assessed after transfection with miR-30e-inhibitor. Interestingly, markedly decreased miR-30e levels were observed after silencing of the miRNA, resulting in increased PC9 cell growth compared with the miR-NC-inhibitorgroup (Figure 4A).
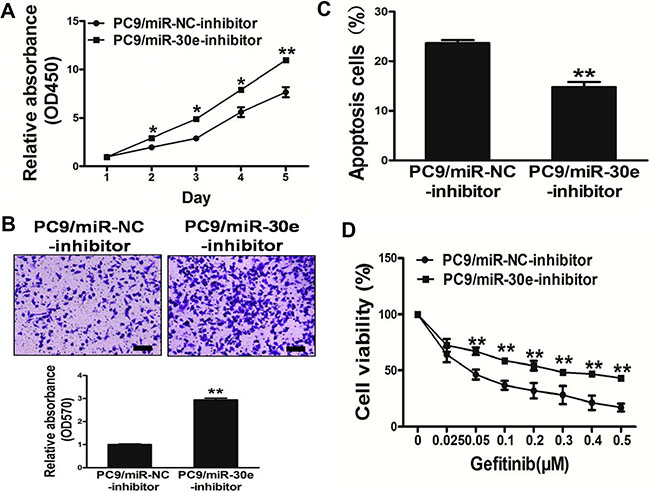
Figure 4: Repression of miR-30e in PC9 cells significantlypromotes cell growth and migration, and confers resistance to gefitinib. (A) The CCK8 assay of PC9 cellswas performed after transduction with miR-30e-inhibitor or miR-NC-inhibitor. (B) Transwell migration assays werecarried out for respective cells. (C) Apoptosis assay was performed in respective cells. (D) Gefitinib sensitivity in PC9/miR-NC-inhibitor and PC9/miR-30e-inhibitor cell lines was tested by the CCK-8 assay. Data are mean ± SD of 3 replicate experiments. * and ** indicate significant differences at P < 0.05 and P < 0.01, respectively.
Since migration is a very important malignancy feature, the effects ofmiR-30e on cellmigration was evaluated. As shown in Figure 4B, miR-30e-inhibition dramatically induced the normally strong migration capacity of lung cancer cells, promoting cell survival by inducing apoptosis (Figure 4C). Furthermore, inhibition of miR-30econferred resistance to gefitinib in PC9 cells (Figure 4D). Thus, these results suggest that repression of miR-30e in PC9 cells significantly promoted cell growth andmigration, while conferring resistance to gefitinib.
MiR-30e sensitizes HCC827/GR cells to gefitinib
Next, we adopted thehuman lung cancer cellline HCC827 with its variant HCC827/GR cells which exposure to indicated lower concentration gefitinib for 24 weeks (Figure 5A). Interestingly, miR-30e amounts in HCC827 cells wereincreased compare with the values of the resistant HCC827/GR cells (Figure 5B). Meanwhile, HCC827/GR cells had resistance features, including elevated cell proliferation and a reduced apoptosis rate (Figure 5C–5D). Furthermore, high miR-30e amounts reversed drug resistance to gefitinib in HCC827/GR cells (Figure 5E). Meanwhile, inhibition of miR-30epromotedchemoresistance to gefitinib in HCC827 cell lines (Figure 5F). Our results suggested that miR-30e sensitized HCC827/GR cells to gefitinib.
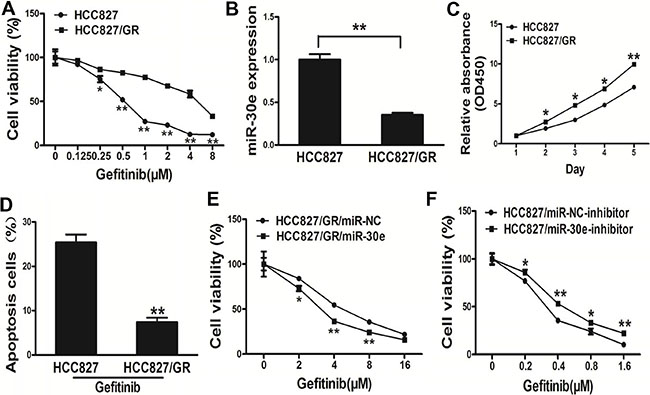
Figure 5: MicroRNA-30e renders HCC827/GR cells more sensitive to gefitinib. (A) In comparison with the HCC827 cell line, HCC827/GR cells displayed less sensitivity to gefitinib. (B) MicroRNA-30e amounts in HCC827 and resistant HCC827/GR cells. (C) Apoptosis Assay were conducted in HCC827 andHCC827/GR cells. (D) The CCK8 assay of HCC827 andHCC827/GR cells was carried out at various time points. (E) Gefitinib sensitivity in HCC827/GR/miR-NC and HCC827/GR/miR-30e cell lines evaluated by the CCK-8 assay. (F) Gefitinib sensitivity in HCC827/miR-NC-inhibitor and HCC827 /miR-30e-inhibitor cell lines assessed by the CCK-8 assay.* and ** indicate significant differences at P < 0.05 and P < 0.01, respectively.
HOXA1 is a direct target of miR-30e
To explore the underlying mechanism of miR-30e in lung cancer, the database TargetScan (www.targetscan.org) was searched. We found that miR-30e likely regulates the HOXA1 gene sinceits 3’-UTR harbored the binding site for the seed region of miR-30e.HOXA1 has a critical function in normal tissue growth and differentiation. Based on the putative binding site of miR-30e in the 3’UTR of the HOXA1 gene, we initially constructed two types of plasmids containingthe luciferase reporting gene with wild-type or mutantHOXA1 3’UTR, and co-transfected miR-30e mimics or inhibitorinto PC9G or PC9 cells; interestingly, cells co-transfected withmiR-30e mimics and wild-type HOXA1 3’UTR showeda significant decrease in luciferase activity, whilemiR-30e-inhibitor significantlyincreasedthe luciferase activity. However, in the mutant group, no detectablechange in luciferase activity was observed (Figure 6A–6B), suggesting that miR-30e suppressed thetranscription activity of the HOXA1 gene by targeting the putative3’UTR of HOXA1 mRNA independently. Western blot demonstrated that HOXA1 protein amounts were reduced in miR-30e treated PC9G cells, and increasedafter miR-30e-inhibition in PC9 cells (Figure 6C). Furthermore, HOXA1amounts were assessed in human lung cancersamples and adjacent normal tissue specimens, with markedly increased values found in cancer specimens (Figure 6D). Next, the association ofHOXA1content with miR-30e amounts in human lung cancer specimens was assessed by Spearman’s rank correlation. Interestingly, HOXA1content and miR-30e levels were inversely correlated in human lung cancer specimens (Spearman’s correlation r = –0.5382) (Figure 6E). These data demonstrated that miR-30e directly targeted HOXA1 in lung cancer cells.
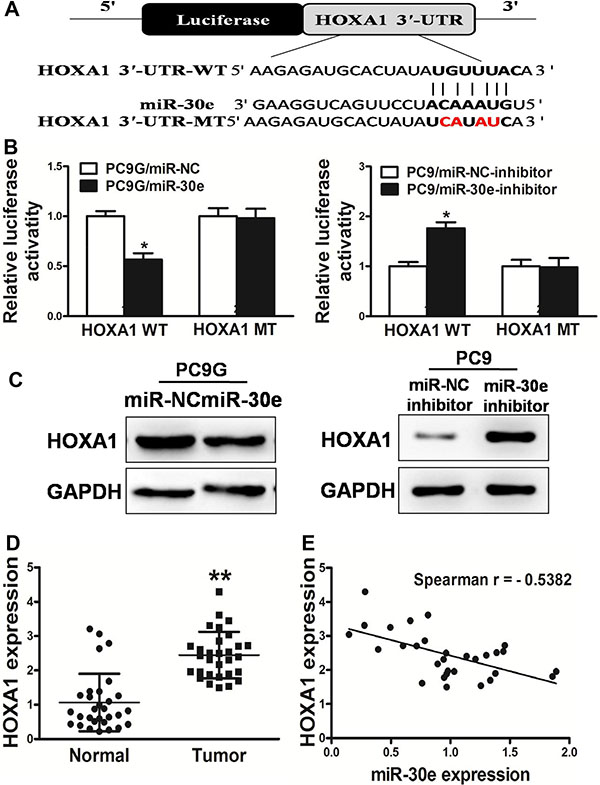
Figure 6: HOXA1 is a miR-30e target. (A) MicroRNA-30e binding site in the human HOXA1 3′-UTR and a reporter construct depicting the whole HOXA1 3′-UTR fragment as well as the mutant HOXA1 3′-UTR (mutated nucleotides are shown in red). (B) Luciferase assay on PC9G or PC9 cells, co-transfected with mimics or inhibitor and a luciferase reporter comprising full length HOXA1 3′-UTR (WT) or mutated (MT) sequence with 4 changed nucleotides in the miR-30e binding site. Luciferase activitywas assessed 24 h after transfection. MicroRNA-30e starklyreduced luciferase activity. Data are mean ± SD (n = 4). (C) The expression of HOXA1 in cells was determined by western blotting analysis. (D) HOXA1 amounts in non-cancerous tissues and human lung cancersamples were assessed by RT-qPCR; fold changes were derived for HOXA1based on GAPDH amounts. (E) The association of HOXA1 amounts and miR-30elevels was determined by Spearman′s correlation. Data are mean ± SD of three replicate experiments. * and ** indicate significant differences at P < 0.05 and P < 0.01, respectively.
MiR-30e enhances the chemosensitivity of gefitinib in vivo
To evaluate the function of miR-30e in canceraggressiveness in vivo, PC9G/miR-NC and PC9G/miR-30e cells were subcutaneously administered into both posterior flanks of male BALB/c nude mice. The formed tumors were measured every other day;gefitinib was administered by peritoneal injection. Interestingly, MiR-30e decreased tumor volumes and weights compared with the miR-NC group, and miR-30e plus gefitinibresulted in decreased tumor volumes and weights compared with miR-30e (Figure 7A–7C). To explore the molecular mechanisms by which miR-30e affects tumor growth, total protein samples were obtained for Western blot; as expected,HOXA1 protein amounts were reduced in miR-30eexpressing tumors (Figure 7D). These findings suggested that miR-30e enhanced lung cancer sensitivity to gefitinib in nude mice.
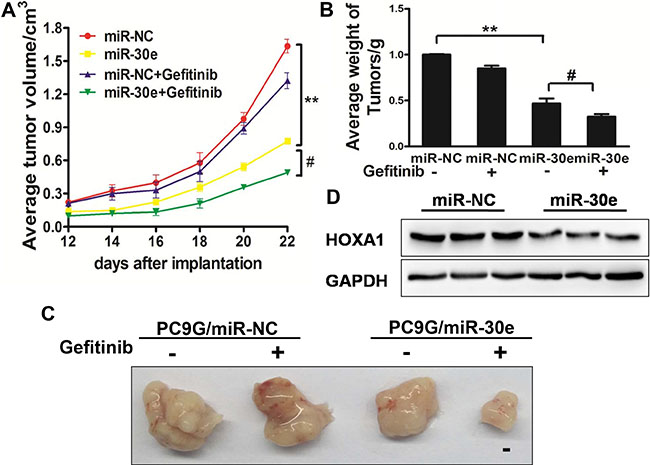
Figure 7: MiR-30e enhances chemosensitivitytogefitinib in a mouse model. (A–C) MicroRNA-30e affectsPC9G cell growth in nude mice. BALB/c nude mice were subcutaneously administered 5 × 106 cells transduced with lentiviruses carrying miR-NC or miR-30e. The tumorswere measured at different time points; gefitinibtreatment was carried out intraperitoneally. The tumorswere extracted and weighed at 24 days. MicroRNA-30e treatment resulted in decreased tumor volumes and weights compared with the miR-NC group; meanwhile, miR-30e plus gefitinibfurther inhibited tumor growth compared with the miR-30e group. Bar = 1 mm. (D) HOXA1amounts were assessed in tumor tissue specimens by immunoblotting. Data are mean ± SD. * and # indicate significant differences at P < 0.01 and P < 0.05, respectively.
DISCUSSION
MicroRNAs have significant functions in carcinogenesis, with some correlated with clinical characteristics and outcomes [22]. Meanwhile, lung cancertumorigenesis involves both genetic and epigenetic alterations, including the induction of oncogenes and/or suppression of tumor suppressors. Altered miRNA expression is commonly found in human carcinomas, e.g. NSCLC [23, 24]. Here, the role of miR-30e in lung carcinoma was evaluated, as well as the underlying molecular mechanisms.
Mounting evidence suggests that miR-30e is a potential tumor suppressor in multiple cancers. As shown above, reduced miR-30e levelswere obtained in lung cancer specimens, in comparison with adjacent non-cancerous tissue samples. The expression of miR-30e was reduced in the resistantlung carcinoma PC9G cell line in comparison with PC9 cells. In agreement, miR-30e overexpression resulted in decreased cell growth and migration, while inducingapoptosis in PC9G cells; conversely miR-30e repression markedly promoted cell growth and migration, and inhibited apoptosis in PC9 cells. This study is the first to reveal miR-30e overexpression in chemosensitivity. These findings could help developnovel therapeutic strategies for lung carcinoma treatment.
HOXA1, a member of the HOXA family which wasfirst identified in Drosophila, has been previously reported to significantly influence the normal growth and differentiationof mammalian tissues [25, 26]. HOXA1 re-expression in human mammary epithelial cells was shown to cause oncogenic transformation and tumorigenesis in vivo [27]. HOXA1 mutation results in decreased mammary cell proliferation, anchorageindependentgrowth, and loss of contact inhibition [28]. In addition, HOXA1 expression levels in squamous cell lung-and cervical cancer tissue samples are significantly elevated compared with adjacent normal tissue specimens [19, 29]. Here, the HOXA1 oncogene was further identified as anew miR-30e target both in vitro and in mice. First, luciferase reporter assay demonstrated that miR-30e directly recognized the 3′-UTR of HOXA1 mRNA. In addition, HOXA1 levels were significantly reduced after stable miR-30e expression. Thirdly, HOXA1 and miR-30ewere inversely correlated in clinical specimens. Taken together, these findings indicated that HOXA1 is anew miR-30e target.
MicroRNAs are considered to be involved in cancer chemoresistance; indeed, they are differentially expressed in chemo-sensitive and chemoresistant cells [30–32]. Interestingly, ectopic miR-34a was shown to sensitizecolorectal carcinoma cells to 5-fluorouracil [33]; meanwhile,miR-497 reduces tumor cell proliferation and sensitizes to 5-fluorouracil by inactivating KSR1 [34], and targets N-RAS to increase temozolomide-dependent apoptosis in gliomas [35]. As shown above,miR-30e enhances chemosensitivity togefitinib, both in vitro and in vivo, confirmingthatmiR-30e re-expression may constitute a novel strategy forovercoming chemoresistance to gefitinib in lung cancer. However, the majority of clinical samples did not undergoEGFR mutationassessmentbecause most patients were administeredgefitinib after unsuccessful chemotherapy and diseaseprogression. Thisis alimitation if this study. Therefore, it remains unclear whether theobserved response may be due to EGFR-mutations. We then retrospectively assessed samples with EGFR-mutation, which also showedelevatedmiR-30e amounts. These findings indicated that elevated miR-30e amounts may overlap to some extentwith EGFR-mutation, which is a very well-known predictive marker of response to EGFR-TKIs. Further research is required to testthis hypothesis.
In summary, the current findingsfirstly demonstrated that miR-30e played a significant role in suppressing lung cancer cell growth through HOXA1inhibition. Although we confirmed that miR-30e could inhibit lung cancer by targeting HOXA1, there might be other miR-30etargets, which could also affect tumor growth in lung cancer. Therefore, further studies are warranted for identifying additional targets and pathways modulated by miR-30e. Although the mechanisms underpinning lung cancer are currently more understood, treating this malignancy still constitutes a clinical challenge.
MATERIALS AND METHODS
Cell culture and clinical tissue specimens
Human lung cancer PC9、PC9G、HCC827 and HCC827/GR cells were maintained in RPMI 1640;the HEK-293T cell line was cultured in DMEM containing 10% fetal bovine serum (FBS), 100 IU/mL penicillinand 100 mg/mL streptomycin. Cell culture was carried out at 37°C in a humid atmosphere with 5% CO2. PC-9 and HCC-827 cellsharborthe activating EGFRmutation del E746-A750 in exon 19.
Lung cancer tissue samples and adjacent non-canceroustissue specimens were obtained from patients undergoing lung cancer resection, and snap-frozen in liquid nitrogen post-surgery.
Lentiviral packaging of miR-30e and stable cell line establishment
A lentiviral packagingkit was usedtostably overexpress miR-30e in lung cancer cells. Lentivirus carryingmiR-30e or negative control (miR-NC) was packaged according to the manufacturer’s instructions, in HEK-293T cells using polybrene (Sigma-Aldrich); selection was performed by treatment with puromycin (Sigma-Aldrich) for 2 weeks, to yield stablecell lines.
RNA purification and real-time reverse transcription polymerase chain reaction (qRT-PCR)
Total RNA was obtained from cultured cells with TRIzol reagent (Invitrogen, USA) as instructed by the manufacturer. Quantitative real-time RT-PCR detecting mature miR-30e was carried out in triplicate with RT Reagent Kit (Vazyme, Nanjing, China) as directed by the manufacturer, withAceQ SYBR Master Mix (Vazyme, Nanjing, China) on a 7900HT system. MicroRNA-30e levels in each group were determined relative to U6 amounts, by the 2-ΔΔCt) method.
Cell proliferation assay
Cell counting Kit-8 (CCK8 kit, Dojindo Laboratories, Japan) assay was used for cell viability assessment. A total of 2,000 cells were plated per well in 96-well plates, and cultured as described above for 48 h after transfection. After incubationforindicated times, CCK-8 reagent was supplemented per well and further incubated for 1–2 h. Absorbance was read at 450 nm.
Migration assay
The effects of miR-30e on cell migrationwere investigated in 24-wellMatrigel invasion chambers (BD Biosciences, UK) as instructed by the manufacturer. Transfected cells (5 × 104) were plated in upper wells in serum-free RPMI-1640, with RPMI-1640 containing 10% FBS in lower chambers. After 16-20 h, non-invading cells (top wells) were removed; invasive cells (bottom wells) were submitted to staining with 0.1% crystal violet after fixation (paraformaldehyde). Photomicrographs were captured in 3 randomly selected high power fields. After air drying, the membranes were treated with 33% acetic acid (300 μL/well) at room temperature for 15 minutes, and the resulting solutions transferred into 96-well plates. Absorbance at a wavelength of 570 nm was recorded.
Western blotting
Cells were treated as described above for 48 h, and lysed in RIPA buffer containing protease inhibitorson ice for 30 min. Total protein amounts were assessed by the BCA assay (Beyotime, China). Equal amounts of protein were then separated by 10% SDS-PAGE. Subsequently, protein bands were electrically transferred onto nitrocellulose membranes (Whatman, Germany), which were incubated with anti-HOXA1 (Proteintech Technology, USA) and anti-GAPDH (Bioworld Technology, USA) antibodies at 4°C overnight.
Luciferase reporter assay
TargetScan was employed to predict miR-30e binding sites. A fragment of the 3′-UTR of HOXA1 with the putative miR-30e binding site was cloned by PCR. To generate a construct harboring mutated miR-30e binding site, four nucleotides corresponding to the 5′-seeding region of this site were substituted in the wild type fragment. The complementary fragment in the 3′-UTR of HOXA1 (UGUUUAC) was mutated to UCAUAUC. PCR products were cut with SacI and HindIII, cloned inserted into pMIR-REPORTER, and validated by DNA sequencing. Constructs were then co-transfected with miR-30e or miR-NC into HEK-293 cells in 24-well plates for 24 h, followed by Luciferase assays with Dual Luciferase Reporter Assay System (Promega, WI, USA).
Apoptosis assay
Apoptosis was assessed flow-cytometrically, afterstaining with AnnexinV and propidium iodide (BD Pharmingen) according to the manufacturer’s instructions. Analysis was carried out on FACSCanto II (BD Biosciences) with the FlowJo software.
Xenograft studies
BALB/cA-nu (nu/nu) nude mice (male, 6-weeks old], purchased from Shanghai Laboratory Animal Center (Chinese Academy of Sciences, Shanghai, China), were housed in a specific pathogen-free (SPF) vivarium. Aliquots of cells (5 × 106) in 150 mL FBS-free RPMI 1640 was subcutaneously administered into both posterior flanks of the animals. Tumors were measured withVernier calipers every 2 days, with volumesderived as followed: Volume = 0.5 × Length × Width2. Ten days after implantation, gefitinib (5 μM) was intraperitoneal injected in the indicated mice. The animals were euthanized 24 days after implantation, extracting the tumors.
Statistical analysis
Experiments were carried out in triplicate, with data assessed using GraphPad Prism 5 (La Jolla, CA, USA). The association of miR-30e levels with HOXA1 amounts in human lung cancer wasassessed by Spearman’s rank test. Group comparison was carried out by t-test. Statistical significance level was set at P < 0.05.
ACKNOWLEDGMENTS AND FUNDING
This work was supported in part by the Science Foundation of FirstPeople’s Hospital of Wujiang, Suzhou (201607), theNational Natural Science Foundation of China (81202032 and 81172140), the Jiangsu 233 Province Clinical Science and Technology Projects (Clinical Research Center, BL2012008), and the Priority Academic ProgramDevelopment of Jiangsu Higher Education Institutions (JX10231801).
CONFLICTS OF INTEREST
The authors declare that they have no conflicts of interest.
REFERENCES
1. Siegel R, Naishadham D, Jemal A. Cancer statistics, 2012. CA Cancer J Clin. 2012; 62:10–29.
2. Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW, Harris PL, Haserlat SM, Supko JG, Haluska FG, Louis DN, Christiani DC, Settleman J, et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. The New England journal of medicine. 2004; 350:2129–2139.
3. Deng Z, Rong Y, Teng Y, Zhuang X, Samykutty A, Mu J, Zhang L, Cao P, Yan J, Miller D, Zhang HG. Exosomes miR-126a released from MDSC induced by DOX treatment promotes lung metastasis. Oncogene. 2016.
4. Pan B, Feng B, Chen Y, Huang G, Wang R, Chen L, Song H. MiR-200b regulates autophagy associated with chemoresistance in human lung adenocarcinoma. Oncotarget. 2015; 6:32805–32820. doi: 10.18632/oncotarget.5352.
5. Ye Z, Yin S, Su Z, Bai M, Zhang H, Hei Z, Cai S. Downregulation of miR-101 contributes to epithelial-mesenchymal transition in cisplatin resistance of NSCLC cells by targeting ROCK2. Oncotarget. 2016; 7:37524–37535. doi: 10.18632/oncotarget.6852.
6. Shen H, Yu X, Yang F, Zhang Z, Shen J, Sun J, Choksi S, Jitkaew S, Shu Y. Reprogramming of Normal Fibroblasts into Cancer-Associated Fibroblasts by miRNAs-Mediated CCL2/VEGFA Signaling. PLoS genetics. 2016; 12:e1006244.
7. Ambros V, Lee RC. Identification of microRNAs and other tiny noncoding RNAs by cDNA cloning. Methods in molecular biology. 2004; 265:131–158.
8. Pillai RS, Bhattacharyya SN, Filipowicz W. Repression of protein synthesis by miRNAs: how many mechanisms? Trends in cell biology. 2007; 17:118–126.
9. Thai TH, Christiansen PA, Tsokos GC. Is there a link between dysregulated miRNA expression and disease? Discovery medicine. 2010; 10:184–194.
10. German MA, Pillay M, Jeong DH, Hetawal A, Luo S, Janardhanan P, Kannan V, Rymarquis LA, Nobuta K, German R, De Paoli E, Lu C, Schroth G, et al. Global identification of microRNA-target RNA pairs by parallel analysis of RNA ends. Nature biotechnology. 2008; 26:941–946.
11. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004; 116:281–297.
12. Feng G, Shi H, Li J, Yang Z, Fang R, Ye L, Zhang W, Zhang X. MiR-30e suppresses proliferation of hepatoma cells via targeting prolyl 4-hydroxylase subunit alpha-1 (P4HA1) mRNA. Biochemical and biophysical research communications. 2016; 472:516–522.
13. Hershkovitz-Rokah O, Modai S, Pasmanik-Chor M, Toren A, Shomron N, Raanani P, Shpilberg O, Granot G. MiR-30e induces apoptosis and sensitizes K562 cells to imatinib treatment via regulation of the BCR-ABL protein. Cancer letters. 2015; 356:597–605.
14. Sugihara H, Ishimoto T, Watanabe M, Sawayama H, Iwatsuki M, Baba Y, Komohara Y, Takeya M, Baba H. Identification of miR-30e* regulation of Bmi1 expression mediated by tumor-associated macrophages in gastrointestinal cancer. PloS one. 2013; 8:e81839.
15. Wu F, Zhu S, Ding Y, Beck WT, Mo YY. MicroRNA-mediated regulation of Ubc9 expression in cancer cells. Clinical cancer research. 2009; 15:1550–1557.
16. Shah N, Sukumar S. The Hox genes and their roles in oncogenesis. Nature reviews Cancer. 2010; 10:361–371.
17. Cillo C, Cantile M, Faiella A, Boncinelli E. Homeobox genes in normal and malignant cells. Journal of cellular physiology. 2001; 188:161–169.
18. Grier DG, Thompson A, Kwasniewska A, McGonigle GJ, Halliday HL, Lappin TR. The pathophysiology of HOX genes and their role in cancer. The Journal of pathology. 2005; 205:154–171.
19. Abe M, Hamada J, Takahashi O, Takahashi Y, Tada M, Miyamoto M, Morikawa T, Kondo S, Moriuchi T. Disordered expression of HOX genes in human non-small cell lung cancer. Oncology reports. 2006; 15:797–802.
20. Brock A, Krause S, Li H, Kowalski M, Goldberg MS, Collins JJ, Ingber DE. Silencing HoxA1 by intraductal injection of siRNA lipidoid nanoparticles prevents mammary tumor progression in mice. Science translational medicine. 2014; 6:217ra212.
21. Rice KL, Licht JD. HOX deregulation in acute myeloid leukemia. The Journal of clinical investigation. 2007; 117:865–868.
22. Ambros V. MicroRNA pathways in flies and worms: growth, death, fat, stress, and timing. Cell. 2003; 113:673–676.
23. Du L, Pertsemlidis A. microRNAs and lung cancer: tumors and 22-mers. Cancer metastasis reviews. 2010; 29:109–122.
24. Gibson NW. Engineered microRNA therapeutics. The journal of the Royal College of Physicians of Edinburgh. 2014; 44:196–200.
25. Wang H, Liu G, Shen D, Ye H, Huang J, Jiao L, Sun Y. HOXA1 enhances the cell proliferation, invasion and metastasis of prostate cancer cells. Oncology reports. 2015; 34:1203–1210.
26. Wardwell-Ozgo J, Dogruluk T, Gifford A, Zhang Y, Heffernan TP, van Doorn R, Creighton CJ, Chin L, Scott KL. HOXA1 drives melanoma tumor growth and metastasis and elicits an invasion gene expression signature that prognosticates clinical outcome. Oncogene. 2014; 33:1017–1026.
27. Zhang X, Zhu T, Chen Y, Mertani HC, Lee KO, Lobie PE. Human growth hormone-regulated HOXA1 is a human mammary epithelial oncogene. The Journal of biological chemistry. 2003; 278:7580–7590.
28. Delval S, Taminiau A, Lamy J, Lallemand C, Gilles C, Noel A, Rezsohazy R. The Pbx interaction motif of Hoxa1 is essential for its oncogenic activity. PloS one. 2011; 6:e25247.
29. Shim C, Zhang W, Rhee CH, Lee JH. Profiling of differentially expressed genes in human primary cervical cancer by complementary DNA expression array. Clinical cancer research. 1998; 4:3045–3050.
30. Boren T, Xiong Y, Hakam A, Wenham R, Apte S, Chan G, Kamath SG, Chen DT, Dressman H, Lancaster JM. MicroRNAs and their target messenger RNAs associated with ovarian cancer response to chemotherapy. Gynecologic oncology. 2009; 113:249–255.
31. Xia L, Zhang D, Du R, Pan Y, Zhao L, Sun S, Hong L, Liu J, Fan D. miR-15b and miR-16 modulate multidrug resistance by targeting BCL2 in human gastric cancer cells. International journal of cancer. 2008; 123:372–379.
32. Shen H, Wang L, Ge X, Jiang CF, Shi ZM, Li DM, Liu WT, Yu X, Shu YQ. MicroRNA-137 inhibits tumor growth and sensitizes chemosensitivity to paclitaxel and cisplatin in lung cancer. Oncotarget. 2016; 7:20728–20742. doi: 10.18632/oncotarget.8011.
33. Siemens H, Jackstadt R, Kaller M, Hermeking H. Repression of c-Kit by p53 is mediated by miR-34 and is associated with reduced chemoresistance, migration and stemness. Oncotarget. 2013; 4:1399–1415. doi: 10.18632/oncotarget.1202.
34. Wang L, Jiang CF, Li DM, Ge X, Shi ZM, Li CY, Liu X, Yin Y, Zhen L, Liu LZ, Jiang BH. MicroRNA-497 inhibits tumor growth and increases chemosensitivity to 5-fluorouracil treatment by targeting KSR1. Oncotarget. 2016; 7:2660–2671. doi: 10.18632/oncotarget.6545.
35. Wang L, Shi ZM, Jiang CF, Liu X, Chen QD, Qian X, Li DM, Ge X, Wang XF, Liu LZ, You YP, Liu N, Jiang BH. MiR-143 acts as a tumor suppressor by targeting N-RAS and enhances temozolomide-induced apoptosis in glioma. Oncotarget. 2014; 5:5416–5427. doi: 10.18632/oncotarget.2116.

