INTRODUCTION
Precision medicine represents a new era in medicine - one that delivers the most appropriate treatment at the most appropriate time enabled, in large part, by the mapping of the human genome [1]. Its initial purpose is focused on cancer, by providing a new strategy for cancer treatment approaches.
Head and neck squamous cell carcinoma (HNSCC) is the sixth most common cancer all over the world. It remains a serious public health problem [2, 3]. More than 500,000 new head and neck cancer patients are diagnosed and approximately 300,000 deaths are caused by HNSCC every year. There have been 48,300 new cases and 9,570 deaths just in USA according to Rebecca L. Siegel’s report for 2016 [2]. Over 90 percent of cancers that arise from the oral mucosa are diagnosed as squamous cell carcinomas [4]. HNSCC also remains a serious public health problem in Asian countries [5]. However, the current therapies including surgery, radiation therapy, Cisplatin-based chemoradiotherapy and pharmacotherapy are often aggressive [6, 7] and carry considerable side effects [8]. In most cases these treatments lead to poor prognostic outcomes [6]. Only 40%-50% of patients with HNSCC survive for 5 years [9]. When accurate diagnosis and improved prognosis are required for oral cancer [10], precision medicine gives the HNSCC patients new hope.
In this paper, we will summarize the main steps in the emergence of precision medicine, and then focus on how precision medicine is used in HNSCC, especially for targeted therapy and gene therapy.
EMERGENCE OF PRECISION MEDICINE
Precision medicine is an emerging therapeutic strategy that aims at disease prevention and treatment through comprehensive analysis patient’s individual variations in genes, environment, and lifestyle [11]. It is beyond the scope of P4 medicine (predictive, personalized, preventive and participatory) [12]. The term “precision” has largely replaced “personalized”, because it considers the patient and health care system interaction in a more accurate manner. Precision medicine is a personalized intervention therapeutic method based on the big OMICS data including epigenetics, genomics, proteomics and metabolomics, and is involved in the classification, staging and grading of the disease depending on histopathology or immunology. Precision medicine may be used to predict the development of the diseases and patients’ response to therapies, facilitating clinical decision making to achieve the best curative effect [13]. It contains two key factors: (1) building a biological database; (2) utilizing the data to guide the individual disease diagnose and treatment. Five important steps are involved in the process of proposition precision medicine (Figure 1).
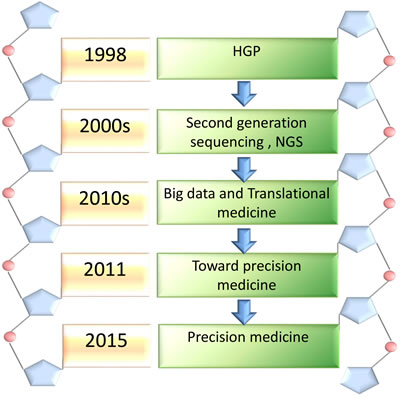
Figure 1: The major steps of the emergence of precision medicine. Precision medicine experienced five important steps from 1985 to 2015. HGP leads us to enter the genomic era in 1985. With the explosive development of biomedical technology, the second and next generation sequencing technology was invented in the 2000s, making gene sequencing much faster, cheaper and preciser. Big data and translational medicne promoted the development of precision medicine. A framework of the “toward presision medicine” established a conception of new therapeutic strategy in 2011. Precison medicion was presented officially by President Obama in 2015.
It has been over forty years since man’s first landing on the moon-Project Apollo. Human landed on the “moon” again-Human Genome Project (HGP), which was initiated in 1985 [14]. Five years later, HGP was launched with the approval of the congress of the United States, planning to spend at least $3 billion for the analysis of the human genome in 15 years. It was an important day when scientists who were from the United States, United Kingdom, France, Germany, Japan and China announced that they finished drawing the human genome sketches in 2000 [15]. The completion of HGP is an important step forward towards understanding the mysteries of the human beings.
With the completion of HGP and the rise of second-generation sequencing technology and next-generation sequencing technology (NGS) [16], bioinformatics analysis of large sets of data has been dramatically amplified in the 2000s. How should the big data be appropriately used for clinical applications? Translational medicine has partially solved this problem, but it is not sufficient [17, 18]. To address this question, the National Academy of Sciences made a systematic discussion of a new concept “Precision Medicine” in “Toward Precision Medicine: Building a Knowledge Network for Biomedical Research and a New Taxonomy of Disease” [19].
President Obama launched a new Precision Medicine Initiative, which aims to cure diseases like cancer based on using of personalized information to keep ourselves and our families away from diseases on January 30th, 2015. In his speech, he had infinite faith in that public health would significant progress with the development of scientific research. He established a research institution aiming to make the best use of genetically informed approaches for cancer treatment. Over 1 million volunteers are involved in this project. It is believed that it is time to realize his far-sighted initiative, and the National Institutes of Health (NIH) as the main leadership and other participators will join hands with each other to make precision medicine come true [20].
APPLICATION OF PRECISION MEDICINE TO CANCER THERAPY
Cancer is one of the major diseases leading to death. Recent studies have demonstrated that cancer can be caused by a series of molecular damage and that each cancer has individual and characteristic genomic codes with both common and tumor-specific features [20]. NIH announced their near-team goals - intensifying efforts to apply precision medicine to cancer, including innovating clinical trials of targeted drugs, using combination therapies, and overcoming drug resistance. President Obama unveiled the Precision Medicine initiative during the State of the Union address. Focusing on cancer, it aims to encourage more volunteers to join this research and build biomedical data by collecting and dealing with the genotypic, phenotypic, environment and lifestyle information [21]. Cancer Medicine Program (CMP) aims to utilize precise genome sequencing and information analysis for accurate diagnosis of cancer, molecular typing, treatment response prediction and interpretation for resistance to cancer drugs. In fact, the discovery of the most effective targeted cancer therapies comes from cancer genetics and genomics studies. Accordingly, cancer genome sequencing may provide cancer patients with effective personalized precision therapies [23]. Numerous previous studies have demonstrated the power of precision medicine in cancer treatment [24, 25].
PRECISION MEDICINE FOR HEAD AND NECK SQUAMOUS CELL CARCINOMA
Precision prediction and diagnosis of HNSCC
Oral potential malignant disease (OPMD)
Oral potential malignant disease (OPMD), such as oral leukoplakia, erythroplakia, is likely to cause oral squamous cell carcinoma (OSCC) since more than 62 percent of OSCC results from OPMD [27]. Therefore, the identification of the underlying relationship between OPMD and OSCC should enable early diagnosis of OSCC. A large number of studies have been performed to explore the potential relevance between OPMD and OSCC [28]. Sequencing results of the lesions that have high risk of malignancy may be used to predict and improve disease progression and outcome.
In the study by Jay O. Boyle [29], the incidence of p53 mutations was detected in 19 percent of non-invasive lesions and 43 percent of invasive carcinomas, indicating that p53 mutations increased in preceding invasion in primary head and neck cancer. In the study by Guozhong Qin et al. [30], the p53 mutations were found in both erythroplakia lesions and oral squamous cell carcinoma. Sciubba James J indicated that the p53 mutations increase the incidence of erythroplakia in developing malignancy and proved that p53 mutation is an early event during the development of OSCC [31]. Davide B. Gissi found that the lesions express p53 and Ki67 at the same time, indicating that the non-dysplastic oral leukoplakia may be at risk of developing oral cancer through a retrospective longitudinal study [32].
Nilva K. Cervigne [24] found that the DNA copy numbers underwent unpredictable change in the process of DNA replication. Some of the copy numbers increased like 1p (over 80%), 11q13.4 (68%) and 9q34.13 (64%), especially the quite obvious increasing on the segment of 1p35 and 1p36. He also observed that other regions of DNA copy number were missing in more than 20 percent of samples, especially on 5q31.2 (about 35%), 16p13.2 (about 30%) and so on. In sum, Nilva K. Cervigne concluded that there is a correlation between the DNA copy number alterations (CNAs) and the different grades of dysplasia in 70 percent of patients. These statistical data indicated that CNAs may be connected with OSCC progression. Recent studies reached similar conclusion that the DNA copy number change (gain and lose) can be used to predict the risk of OSCC progress in patients with OPMD [33-35].
By examining microRNA expression in twelve patients with progressive leukoplakias and four with non-progressive leukoplakias, Cervigne NK found that 109 microRNAs were specifically overexpressed in patients with progressive leukoplakia and invasive OSCC. The malignant transformation might be closely related to the high-expression of microRNA-21, microRNA-181b and microRNA-345. His findings suggested that the change of microRNA expression can predict the risk of malignant transformation of leukoplakia [36]. Guanghui Zhu also provided direct evidence that microRNA played an important role in the development of leukoplakia to oral cancer [37]. It was reported that about 37.5 percent of microRNAs were related to oral cancer in previous literature [37].
Head and neck squamous cell carcinoma (HNSCC)
Oncogenes and tumor suppressor genes play critical roles in cancer initiation and progression. The dysregulated expression and mutation of these genes are detected in all kinds of cancers and the related studies have been fruitful in the past decades. Oncogenes and tumor suppressor genes are now frequently used as molecular targets for cancer treatment [38]. Genetic alterations are considered as an important step in the development and progression of HNSCC, leading to the dysregulation of essential cellular signaling pathways involved in cell proliferation and differentiation [39]. Progressive allelic loss is one of the significant steps in the development of HNSCC. There have been many oncogenes and tumor association suppressor genes identified in the previous studies, including p53 [40], p16 (CDKN2A) [41, 42], NOTCH, EGFR [43], RAS, CCND1 [44], BRAF, and PIK3CA [45, 46]. The roles of these genes and their therapeutic potential in HNSCC have just begun to be revealed [26, 47-49]. The ratios of mutation or dysregulated expressing of key genes are listed in Table 1.
Table 1: Application of key genes and their ratios of mutation or expressing
Gene |
Ratios of mutation/ expressing |
Application |
Refs |
P53 |
50% mutation |
Precision prediction and diagnosis; Gene addition therapy; Precision surgery |
|
EGFR |
95% overexpressing |
Precision prediction and diagnosis; Targeted therapy; Gene disruption therapy; Targeted radionuclide therapy |
|
VEGFR |
33% overexpressing |
Targeted therapy; Gene disruption therapy; Targeted radionuclide therapy |
|
HER2 |
1%-2% overexpressing |
Targeted therapy |
|
NOTCH1 |
10%-18.6% mutation |
Precision prediction and diagnosis |
|
PI3K/Akt/mTOR |
30.5% mutation |
Targeted therapy |
|
RAS/RAF/MEK/ERK |
6% mutation |
Precision prediction and diagnosis; Targeted therapy |
|
JAK/STAT3 |
Rarely mutated |
Targeted therapy |
|
CDKN2A |
78.7% overexpressing |
Precision prediction and diagnosis |
[43] |
The ratios of mutations or expressing of several important genes are shown by this table. The table also demonstrates how gene targets are used in precision medicine.
As a tumor suppressor gene, p53 gene contributes to the arrest of tumor cell proliferation and tumor growth [50, 51]. P53 gene mutation can be detected in about 50 percent of HNSCC patient samples [50, 51]. The incidence of p53 mutation significantly increased when the pre-malignant diseases progressed to invasive cancer [52, 53].
NOTCH1 is another important tumor suppressor gene. Approximately 10 to 18.6 percent of HNSCC patients carry mutations in the NOTCH1 gene, making NOTCH1 the second most frequently mutated gene next to p53 [54, 55]. Nicolas Stransky’s findings indicted that the dysregulation of NOTCH signaling enhanced the genesis and progression of HNSCC [3].
It was found that HPV infection is an important risk factor associates with a large group of HNSCC patients [56]. HPV-positive and HPV-negative HNSCCs have distinct clinical features and dissimilar biology [9, 57]. Approximately 22-26 percent of HNSCCs were found to be HPV positive. The prevalence was even up to 35-41 percent in OSCCs [57, 58]. The treatment outcome of the HPV-positive patients was deficient [59]. It may be associated with HPV oncoproteins E6 and E7 [58].
Telomerase is also an important marker for early diagnosis and prognosis of cancer, including HNSCC [60, 61]. Promoter hypermethylation has been found to be related to the assessment of HNSCC patients’ prognosis. Examination of the hypermethylation status can be used for early diagnosis, invasive detection and tumor surveillance in clinic [62, 63].
Precision medicine for the treatment of HNSCC
Traditional therapeutic modalities such as surgery, chemotherapy, and radiation therapy have their own limitations [17, 64, 65]. Advanced understanding of the molecular mechanism of cancer has highlighted targeted therapies as more promising cancer therapeutics for patients with HNSCC (Figure 2).
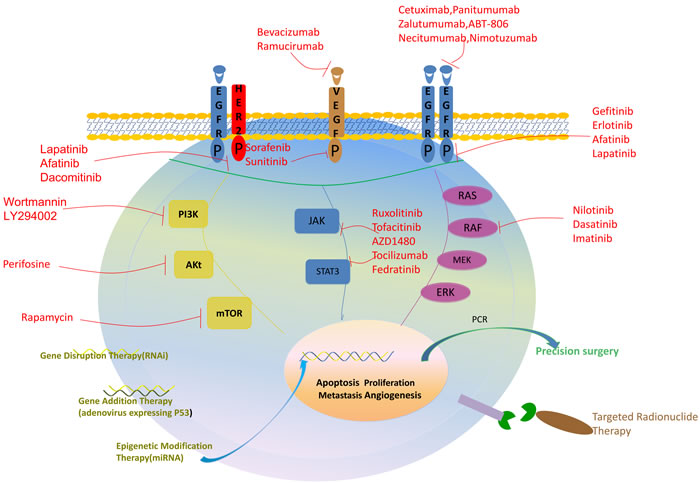
Figure 2: Precision medicine used for the treatment of HNSCC. This figure shows 5 major methods of precision medicine used in HNSCC patient: (1) inhibitors targeting cell surface signaling receptors, including EGFR, VEGFR, HER2 (2) inhibitors targeting cellular signaling pathways, including PI3K/Akt/mTOR, RAS/RAF/MEK/ERK, JAK/STAT3 (3) precision genetic manipulation therapies, including gene disruption therapy, gene addition therapy and epigenetic modification therapy (4) precision medicine combining with surgery (5) precision targeted radionuclide therapy.
Targeting cell surface signaling receptors
EGFR
Epidermal growth factor receptor (EGFR), a cell surface receptor, is one of the most important targets for HNSCC. EGFR is a member of the ErbB growth factor receptor tyrosine kinase family, which has four members: EGFR-ErbB1, Her2-neu-ErbB2, HER3-ErbB3 and HER4 -ErbB4 [7, 66-68]. Hyperactivated EGFR signaling has been frequently found in HNSCC cells. EGFR is involved in the regulation of cell proliferation, metastasis, migration, invasion, angiogenesis, and inhibition of apoptosis [69-71]. Although rarely mutated, EGFR is highly upregulated in many cancers with epithelial origins, including HNSCC in which EGFR upregulation is found in over 95% of the tumors [72]. There are many approaches used to inhibit or downregulate the EGFR, such as tyrosine kinase inhibitors (TKls), monoclonal antibodies (mAbs), ligand-linked toxins, and antisense oligonucleotides [67, 73-75]. Two of them are most widely used in clinical models: (1) monoclonal antibodies (mAbs) targeting extracellular surface receptor, including cetuximab, panitumumab, zalutumumab, ABT-806, necitumumab and nimotuzumab; (2) tyrosine kinase inhibitors (TKIs) functioning as a competitor of ATP, including gefitinib, erlotinib, afatinib and lapatinib [76-84]. Cetuximab is the only FDA approved molecular targeted agent in HNSCC, and it has a significant effect combined with radiotherapy, chemotherapy or radio-chemotherapy [85, 86]. Although a case was reported that using cetuximab as a single agent got ideal outcome, the current data did not provide solid evidence for its use as a monotherapy drug [86, 87]. Zalutumumab extended progression-free survival of patients with recurrent HNSCC and median overall survival was increased from 5.2 months to 6.7 months [78]. The patients with HNSCC were assessable for response to TKIs, with an observed response and a disease control, but further study is needed for the better understanding of their clinical application [76, 84, 85].
HER2
Human epidermal growth factor receptor-2 (HER2) is a member of the epidermal growth factor receptor family and activates a variety of signaling pathways, leading to cell proliferation and tumorigenesis [88, 89]. HER2 has been found to be overexpressed in a number of human cancers, including HNSCC, contributing to tumor development, cell cycle progression, cellular motility and growth [90]. A number of strategies are used to inhibit HER2, including mAbs and small-molecule TKIs. Trastuzumab and pertuzumab are the most successful mAbs [91]. Lapatinib, afatinib and dacomitinib are also effective therapeutic agents targeting HER2 [88, 92, 93].
VEGFR
Vascular endothelial growth factor (VEGF) is the most effective pro-angiogenic growth factor, which regulates the growth of blood vessels. VEGF has been identified as an independent prognostic factor for HNSCC [94, 95]. Angiogenesis is critical for tumor growth and metastasis, and becomes an important target of anticancer treatment. Anti-angiogenesis agents have shown promising therapeutic effect on several solid tumors including HNSCC [96], such as bevacizumab, bamucirumab, sorafenib and sunitinib. A combination of anti-angiogenesis agents with other agents achieved enhanced efficacy for HNSCC treatment [97-99].In Maria Vassilakopoulou’s study, it was reported that 3-year progression-free survival rate was improved from 60% to 86% in patients using bevacizumab combining with other antitumor therapies [96].
Cellular signaling pathways
PI3K/Akt/mTOR
Recent progress in understanding the oncogenesis of HNSCC has revealed multiple dysregulated signaling pathways [100]. One frequently altered axis is the PI3K/Akt/mTOR pathway observed in 30.5 percent of HNSCC patients [101-103]. It plays a crucial role in HNSCC progression. PI3K/Akt/mTOR pathway is involved in essential cellular functions including cell proliferation, apoptosis and differentiation. Wortmannin (a natural product) and LY294002 (a synthetic drug) are two representative first-generation PI3K inhibitors whose application has so far been primarily restricted to preclinical studies [100]. A well-established lipid-based Akt inhibitor, perifosine showed potent activity in the inhibition of cell proliferation in preclinical studies [104]. The first mTOR inhibitor rapamycin, a natural product, was originally used for its antifungal properties before its immunosuppressive and antineoplastic effects were discovered [100]. Enhanced efficacy was achieved when these agents were used in combination with other drugs (cetuximab, cisplatin, etc.) in patients with HNSCC [102].
RAS/RAF/MEK/ERK
RAS/RAF/MEK/ERK pathway is another major signaling pathway for HNSCC cell survival [71]. Most often used RAF inhibitors are nilotinib, dasatinib, and imatinib [105]. So far, however, the clinical potential of RAF inhibitors in HNSCC remains to be further explored [38].
JAK/STAT3
JAK/STAT pathway is a newly discovered signal transduction pathway which contributes to the proliferation and survival of tumor cells including HNSCC [106-108]. Several JAK/STAT kinase inhibitors have been identified and are currently being tested in clinical trials, including ruxolitinib, tofacitinib, AZD1480, tocilizumab, fedratinib and pyrimethamine, due to their contribution to treatment resistance and immune escape. [107]. AZD1480 was demonstrated to inhibit proliferation of cell lines andpatient-derived xenograft in HNSCC preclinical models [106].
Precision gene therapy
Gene disruption therapy
Gene disruption therapy is an effective approach in decreasing tumor cell proliferation, slowing tumor invasion and reducing drug resistance. RNA interfere technique (RNAi) was successfully used to target and disable a group of crucial genes for HNSCC treatment [109, 110], including EGFR [111], EpCAM [112], MET receptor [113], Cyclin D1, NF-κB, p65, VEGFR, telomerase reverse transcriptase and p63 [111].
Gene addition therapy
Gene addition therapy is a therapy of adding specific genes to tumor cells with lower or absent expression of such genes. Gene addition therapy has demonstrated its potential anti-tumor effects for HNSCC treatment. Gene addition therapy targeting p53 gene has been very well established [110, 114]. The treatment approaches of delivering p53 have been tested in HNSCC by direct injection of an adenoviral vector expressing wild-type p53 gene [115]. In Peng’s report [116], radiotherapy in combination with p53 addition therapy showed a dramatically improved efficacy (64% showing complete response) than using radiotherapy alone in more than 2000 patients with advanced HNSCC.
Epigenetic modification therapy
The differential expression of miRNAs between cancer and normal tissue has been well documented. Specific miRNAs expression was detected in HNSCC cells [118]. MiRNA involves in a variety of regulatory pathways, including cell proliferation, apoptosis and metabolism. Approaches specifically targeting on miRNAs demonstrated significant anti-cancer potential [118].
Precision medicine in combination with surgery
Most surgeons rely on histological examination like frozen biopsy to determine the surgical margins. However, the local recurrence of HNSCC is often reported in clinical practice [119, 120]. Therefore, a new method to determine the surgical margins is urgently needed. From the perspective of molecular biology, detecting mutation genes or abnormal protein to find rare cancer cells provides a new strategy [120-122]. Polymerase chain reaction (PCR) or immunohistochemistry can be used to detect rare cancer cells. Examination of p53 mutations has been used to detect such cells [35, 122]. Precision surgery with molecular detection can decrease the incidence of local recurrence.
Precision targeted radionuclide therapy
Targeted radionuclide therapy (TRT) is a new therapy of cancer with using the targeted bridging drugs to improve the sensitivity of ray to the tumor cells, which causes less collateral damage to normal tissues. Targeted bridging drugs that play a crucial part in the TRT have two connection ends. One of the ends carries radionuclides and the other shows high affinity to the surface antigens of tumor cell [123, 124]. EGFR mutation is most often observed in HNSCC and considered as a suitable target for TRT [125]. VEGF is also a potential TRT target [80, 82, 126].
FUTURE AND CHALLENGES
Big data does not directly translate genetic information into clinically useful interventions to benefit HNSCC patients [54, 127]. One of the limitations is insufficient large-scale genome sequencing for HNSCC. It’s necessary to build database of HNSCC with larger numbers of patients. In addition, the sharing of data is limited among different countries. Electronic health records will be essential for communication among providers, patients, and researchers regarding crucial medical information [11]. A better designed platform for appropriately sharing clinical information is needed.
There are very few desired solutions for drug resistance [128, 129]. The patients with drug-resistance can be better classified to identify their commonalities by using sequencing technology. With the help of precision medicine, delays in the patient’s diagnosis may be avoided, and more suitable drugs may be chosen.
Combination of anti-tumor therapies may lead to improved treatment outcome, but the underlying mechanism is poorly understood [130-132]. In the ongoing studies, the unknown tumor molecule and genetic pathogenesis will be identified to provide new precision therapeutic targets [8, 126].
CONCLUSIONs
Precision medicine raises everyone’s hopes. Precise treatment is based on gene sequencing, proteomics, genomics and molecular biology. Ideally, each patient should receive a unique optimized treatment regimen based on differentially expressed genes and proteins to achieve the best clinical outcome. Precision medicine gives us a new strategy with which to pursue a cure for HNSCC. Further understanding of the underlying molecular mechanisms will lead to the development of HNSCC therapies with significantly enhanced efficacy.
ACKNOWLEDGMENTS
We thank Xiaobo Luo, Ruixue Ai for scientific discussions.
CONFLICTs OF INTEREST
The authors declare no conflict of interest.
FUNDING
This work was supported by grants from the National Natural Science Foundation of China (No. 81321002 to Q.CH, 81372890 to M.F), and the Thousand Talents Program of China for Distinguished Young Scholars to M.F.
REFERENCES
1. Mirnezami R, Nicholson J, and Darzi A. Preparing for precision medicine. N Engl J Med. 2012; 366: 489-91.
2. Siegel RL, Miller KD, and Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016; 66: 7-30.
3. Stransky N, Egloff AM, Tward AD, Kostic AD, Cibulskis K, Sivachenko A, Kryukov GV, Lawrence MS, Sougnez C, McKenna A, Shefler E, Ramos AH, Stojanov P, et al. The mutational landscape of head and neck squamous cell carcinoma. Science. 2011; 333: 1157-60.
4. Neville BW, and Day TA. Oral cancer and precancerous lesions. CA Cancer J Clin. 2002; 52: 195-215.
5. Murugan AK, Munirajan AK, and Alzahrani AS. MicroRNAs: Modulators of the Ras Oncogenes in Oral Cancer. J Cell Physiol. 2016; 231: 1424-31.
6. Argiris A, Karamouzis MV, Raben D, and Ferris RL. Head and neck cancer. Lancet. 2008; 371: 1695-709.
7. Seiwert TY, Salama JK, and Vokes EE. The chemoradiation paradigm in head and neck cancer. Nat Clin Pract Oncol. 2007; 4: 156-71.
8. Martinez-Useros J, and Garcia-Foncillas J. The challenge of blocking a wider family members of EGFR against head and neck squamous cell carcinomas. Oral Oncol. 2015; 51: 423-30.
9. Leemans CR, Braakhuis BJ, and Brakenhoff RH. The molecular biology of head and neck cancer. Nat Rev Cancer. 2011; 11: 9-22.
10. Varelas X, and Kukuruzinska MA. Head and neck cancer: from research to therapy and cure. Ann N Y Acad Sci. 2014; 1333: 1-32.
11. Jaffe S. Planning for US Precision Medicine Initiative underway. Lancet. 2015; 385: 2448-9.
12. Hood L, and Friend SH. Predictive, personalized, preventive, participatory (P4) cancer medicine. Nat Rev Clin Oncol. 2011; 8: 184-7.
13. Peer D. Precision medicine—delivering the goods. Cancer Lett. 2014; 352: 2-3.
14. Collins FS, Patrinos A, Jordan E, Chakravarti A, Gesteland R, Walters L. New goals for the U.S. Human Genome Project: 1998-2003. Science. 1998. 282: 682-9.
15. Collins FS, Morgan M, and Patrinos A. The Human Genome Project: lessons from large-scale biology. Science. 2003; 300: 286-90.
16. Koboldt DC, Steinberg KM, Larson DE, Wilson RK, and Mardis ER. The next-generation sequencing revolution and its impact on genomics. Cell. 2013; 155: 27-38.
17. Razzouk S. Translational genomics and head and neck cancer: toward precision medicine. Clin Genet. 2014; 86: 412-21.
18. Littman BH, Di ML, Plebani M, and Marincola FM. What’s next in translational medicine. Clin Sci (Lond). 2007; 112: 217-27.
19. National Research Council. Toward precision medicine: building a knowledge network for biomedical research and a New Taxonomy of Disease. US: National Academies Press; 2011.
20. Collins FS, and Varmus H. A new initiative on precision medicine. N Engl J Med. 2015; 372: 793-5.
21. Fox JL. Obama catapults patient-empowered Precision Medicine. Nat Biotechnol. 2015; 33: 325.
22. Garraway LA, and Lander ES. Lessons from the cancer genome. Cell. 2013; 153: 17-37.
23. Xie J. Pathways towards Precision Medicine in Cancer Management Using Genomic Information. J Genet Genomics. 2015; 42: 515-6.
24. Rubin MA. Health: Make precision medicine work for cancer care. Nature. 2015; 520: 290-1.
25. Godbert Y, de Figueiredo B H, Bonichon F, Chibon F, Hostein I, Pérot G, Dupin C, Daubech A, Belleannée G, Gros A, Italiano A, and Soubeyran I. Remarkable Response to Crizotinib in Woman With Anaplastic Lymphoma Kinase-Rearranged Anaplastic Thyroid Carcinoma. J Clin Oncol. 2015; 33: e84-7.
26. Gaykalova DA, Mambo E, Choudhary A, Houghton J, Buddavarapu K, Sanford T, Darden W, Adai A, Hadd A, Latham G, Danilova LV, Bishop J, Li RJ, et al. Novel insight into mutational landscape of head and neck squamous cell carcinoma. PLoS One. 2014; 9: e93102.
27. Cervigne NK, Machado J, Goswami RS, Sadikovic B, Bradley G, Perez-Ordonez B, Galloni NN, Gilbert R, Gullane P, Irish JC, Jurisica I, Reis PP, and Kamel-Reid S. Recurrent genomic alterations in sequential progressive leukoplakia and oral cancer: drivers of oral tumorigenesis. Hum Mol Genet. 2014; 23: 2618-28.
28. Dionne KR, Warnakulasuriya S, Zain RB, and Cheong SC. Potentially malignant disorders of the oral cavity: current practice and future directions in the clinic and laboratory. Int J Cancer. 2015; 136: 503-15.
29. Boyle JO, Hakim J, Koch W, van der Riet P, Hruban RH, Roa RA, Correo R, Eby YJ, Ruppert JM, and Sidransky D. The incidence of p53 mutations increases with progression of head and neck cancer. Cancer Res. 1993; 53: 4477-80.
30. Qin GZ, Park JY, Chen SY, and Lazarus P. A high prevalence of p53 mutations in pre-malignant oral erythroplakia. Int J Cancer. 1999; 80: 345-8.
31. Sciubba JJ. Improving detection of precancerous and cancerous oral lesions. Computer-assisted analysis of the oral brush biopsy. U.S. Collaborative OralCDx Study Group. J Am Dent Assoc. 1999; 130: 1445-57.
32. Gissi DB, Gabusi A, Servidio D, Cervellati F, and Montebugnoli L. Predictive Role of p53 Protein as a Single Marker or Associated with ki67 Antigen in Oral Leukoplakia: A Retrospective Longitudinal Study. Open Dent J. 2015; 9: 41-5.
33. Zhang L, and Rosin MP. Loss of heterozygosity: a potential tool in management of oral premalignant lesions. J Oral Pathol Med. 2001; 30: 513-20.
34. Sudbø J, Kildal W, Risberg B, Koppang HS, Danielsen HE, and Reith A. DNA content as a prognostic marker in patients with oral leukoplakia. N Engl J Med. 2001; 344: 1270-8.
35. Day TA, Davis BK, Gillespie MB, Joe JK, Kibbey M, Martin-Harris B, Neville B, Richardson MS, Rosenzweig S, Sharma AK, Smith MM, Stewart S, and Stuart RK. Oral cancer treatment. Curr Treat Options Oncol. 2003; 4: 27-41.
36. Cervigne NK, Reis PP, Machado J, Sadikovic B, Bradley G, Galloni NN, Pintilie M, Jurisica I, Perez-Ordonez B, Gilbert R, Gullane P, Irish J, and Kamel-Reid S. Identification of a microRNA signature associated with progression of leukoplakia to oral carcinoma. Hum Mol Genet. 2009; 18: 4818-29.
37. Zhu G, He Y, Yang S, Chen B, Zhou M, and Xu XJ. Identification of Gene and MicroRNA Signatures for Oral Cancer Developed from Oral Leukoplakia. Biomed Res Int. 2015; 2015: 841956.
38. Hunt JL, Barnes L, Lewis JS, Mahfouz ME, Slootweg PJ, Thompson LD, Cardesa A, Devaney KO, Gnepp DR, Westra WH, Rodrigo JP, Woolgar JA, Rinaldo A, et al. Molecular diagnostic alterations in squamous cell carcinoma of the head and neck and potential diagnostic applications. Eur Arch Otorhinolaryngol. 2014; 271: 211-23.
39. Agrawal N, Frederick MJ, Pickering CR, Bettegowda C, Chang K, Li RJ, Fakhry C, Xie TX, Zhang J, Wang J, Zhang N, El-Naggar AK, Jasser SA, et al. Exome sequencing of head and neck squamous cell carcinoma reveals inactivating mutations in NOTCH1. Science. 2011; 333: 1154-7.
40. Poeta ML, Manola J, Goldenberg D, Forastiere A, Califano JA, Ridge JA, Goodwin J, Kenady D, Saunders J, Westra W, Sidransky D, and Koch WM. The Ligamp TP53 Assay for Detection of Minimal Residual Disease in Head and Neck Squamous Cell Carcinoma Surgical Margins. Clin Cancer Res. 2009; 15: 7658-65.
41. Demokan S, Chuang A, Suoğlu, Ulusan M, Yalnız, Califano JA, and Dalay N. Promoter methylation and loss of p16(INK4a) gene expression in head and neck cancer. Head Neck. 2012; 34: 1470-5.
42. Rieckmann T, Tribius S, Grob TJ, Meyer F, Busch CJ, Petersen C, Dikomey E, and Kriegs M. HNSCC cell lines positive for HPV and p16 possess higher cellular radiosensitivity due to an impaired DSB repair capacity. Radiother Oncol. 2013; 107: 242-6.
43. Bonner JA, Harari PM, Giralt J, Cohen RB, Jones CU, Sur RK, Raben D, Baselga J, Spencer SA, Zhu J, Youssoufian H, Rowinsky EK, and Ang KK. Radiotherapy plus cetuximab for locoregionally advanced head and neck cancer: 5-year survival data from a phase 3 randomised trial, and relation between cetuximab-induced rash and survival. Lancet Oncol. 2010; 11: 21-8.
44. Rothenberg SM, and Ellisen LW. The molecular pathogenesis of head and neck squamous cell carcinoma. J Clin Invest. 2012; 122: 1951-7.
45. Murugan AK, Hong NT, Fukui Y, Munirajan AK, and Tsuchida N. Oncogenic mutations of the PIK3CA gene in head and neck squamous cell carcinomas. Int J Oncol. 2008; 32: 101-11.
46. Lui VW, Hedberg ML, Li H, Vangara BS, Pendleton K, Zeng Y, Lu Y, Zhang Q, Du Y, Gilbert BR, Freilino M, Sauerwein S, Peyser ND, et al. Frequent mutation of the PI3K pathway in head and neck cancer defines predictive biomarkers. Cancer Discov. 2013; 3: 761-9.
47. Okami K, Wu L, Riggins G, Cairns P, Goggins M, Evron E, Halachmi N, Ahrendt SA, Reed AL, Hilgers W, Kern SE, Koch WM, Sidransky D, et al. Analysis of PTEN/MMAC1 alterations in aerodigestive tract tumors. Cancer Res. 1998; 58: 509-11.
48. Papadimitrakopoulou VA, Izzo J, Mao L, Keck J, Hamilton D, Shin DM, El-Naggar A, den Hollander P, Liu D, Hittelman WN, and Hong WK. Cyclin D1 and p16 alterations in advanced premalignant lesions of the upper aerodigestive tract: role in response to chemoprevention and cancer development. Clin Cancer Res. 2001; 7: 3127-34.
49. Califano J, van der Riet P, Westra W, Nawroz H, Clayman G, Piantadosi S, Corio R, Lee D, Greenberg B, Koch W, and Sidransky D. Genetic progression model for head and neck cancer: implications for field cancerization. Cancer Res. 1996; 56: 2488-92.
50. Tassone P, Old M, Teknos TN, and Pan Q. p53-based therapeutics for head and neck squamous cell carcinoma. Oral Oncol. 2013; 49: 733-7.
51. Lin Y, Mallen-St CJ, Luo J, Sharma S, Dubinett S, and St JM. p53 modulates NF-κB mediated epithelial-to-mesenchymal transition in head and neck squamous cell carcinoma. Oral Oncol. 2015; 51: 921-8.
52. Lobert S, Graichen ME, Hamilton RD, Pitman KT, Garrett MR, Hicks C, and Koganti T. Prognostic biomarkers for HNSCC using quantitative real-time PCR and microarray analysis: β-tubulin isotypes and the p53 interactome. Cytoskeleton (Hoboken). 2014; 71: 628-37.
53. Maruyama H, Yasui T, Ishikawa-Fujiwara T, Morii E, Yamamoto Y, Yoshii T, Takenaka Y, Nakahara S, Todo T, Hongyo T, and Inohara H. Human papillomavirus and p53 mutations in head and neck squamous cell carcinoma among Japanese population. Cancer Sci. 2014; 105: 409-17.
54. Sun W, and Califano JA. Sequencing the head and neck cancer genome: implications for therapy. Ann N Y Acad Sci. 2014; 1333: 33-42.
55. Sun W, Gaykalova DA, Ochs MF, Mambo E, Arnaoutakis D, Liu Y, Loyo M, Agrawal N, Howard J, Li R, Ahn S, Fertig E, Sidransky D, et al. Activation of the NOTCH pathway in head and neck cancer. Cancer Res. 2014; 74: 1091-104.
56. Krupar R, Hartl M, Wirsching K, Dietmaier W, Strutz J, and Hofstaedter F. Comparison of HPV prevalence in HNSCC patients with regard to regional and socioeconomic factors. Eur Arch Otorhinolaryngol. 2014; 271: 1737-45.
57. Polanska H, Heger Z, Gumulec J, Raudenska M, Svobodova M, Balvan J, Fojtu M, Binkova H, Horakova Z, Kostrica R, Adam V, Kizek R, and Masarik M. Effect of HPV on tumor expression levels of the most commonly used markers in HNSCC. Tumour Biol. 2015.
58. Dayyani F, Etzel CJ, Liu M, Ho CH, Lippman SM, and Tsao AS. Meta-analysis of the impact of human papillomavirus (HPV) on cancer risk and overall survival in head and neck squamous cell carcinomas (HNSCC). Head Neck Oncol. 2010; 2: 15.
59. Dorsey K, and Agulnik M. Promising new molecular targeted therapies in head and neck cancer. Drugs. 2013; 73: 315-25.
60. Mori K, Sato S, Kodama M, Habu M, Takahashi O, Nishihara T, Tominaga K, and Takenaka S. Oral cancer diagnosis via a ferrocenylnaphthalene diimide-based electrochemical telomerase assay. Clin Chem. 2013; 59: 289-95.
61. Sebastian S, Grammatica L, and Paradiso A. Telomeres, telomerase and oral cancer (Review). Int J Oncol. 2005; 27: 1583-96.
62. Shaw R. The epigenetics of oral cancer. Int J Oral Maxillofac Surg. 2006; 35: 101-8.
63. Arantes LM, de Carvalho AC, Melendez ME, Carvalho AL, and Goloni-Bertollo EM. Methylation as a biomarker for head and neck cancer. Oral Oncol. 2014; 50: 587-92.
64. Bose P, Brockton NT, and Dort JC. Head and neck cancer: from anatomy to biology. Int J Cancer. 2013; 133: 2013-23.
65. Echarri MJ, Lopez-Martin A, and Hitt R. Targeted Therapy in Locally Advanced and Recurrent/Metastatic Head and Neck Squamous Cell Carcinoma (LA-R/M HNSCC). Cancers (Basel). 2016; 8.
66. Grandis JR, and Tweardy DJ. Elevated levels of transforming growth factor alpha and epidermal growth factor receptor messenger RNA are early markers of carcinogenesis in head and neck cancer. Cancer Res. 1993; 53: 3579-84.
67. Hynes NE, and Lane HA. ERBB receptors and cancer: the complexity of targeted inhibitors. Nat Rev Cancer. 2005; 5: 341-54.
68. Bublil EM, and Yarden Y. The EGF receptor family: spearheading a merger of signaling and therapeutics. Curr Opin Cell Biol. 2007; 19: 124-34.
69. Ang KK, Berkey BA, Tu X, Zhang HZ, Katz R, Hammond EH, Fu KK, and Milas L. Impact of epidermal growth factor receptor expression on survival and pattern of relapse in patients with advanced head and neck carcinoma. Cancer Res. 2002; 62: 7350-6.
70. Rubin GJ, Melhem MF, Gooding WE, Day R, Holst VA, Wagener MM, Drenning SD, and Tweardy DJ. Levels of TGF-alpha and EGFR protein in head and neck squamous cell carcinoma and patient survival. J Natl Cancer Inst. 1998; 90: 824-32.
71. Yokota T. Is biomarker research advancing in the era of personalized medicine for head and neck cancer. Int J Clin Oncol. 2014; 19: 211-9.
72. Kalyankrishna S, and Grandis JR. Epidermal growth factor receptor biology in head and neck cancer. J Clin Oncol. 2006; 24: 2666-72.
73. Mendelsohn J, and Baselga J. Status of epidermal growth factor receptor antagonists in the biology and treatment of cancer. J Clin Oncol. 2003; 21: 2787-99.
74. Mao L, Hong WK, and Papadimitrakopoulou VA. Focus on head and neck cancer. Cancer Cell. 2004; 5: 311-6.
75. Conley BA, Enos RA, and Cheson BD. Clinical trials referral resource. Targeted therapy in squamous cell cancers of the head and neck. Oncology (Williston Park). 2002; 16: 621-2, 625-6.
76. Schmitz S, Ang KK, Vermorken J, Haddad R, Suarez C, Wolf GT, Hamoir M, and Machiels JP. Targeted therapies for squamous cell carcinoma of the head and neck: current knowledge and future directions. Cancer Treat Rev. 2014; 40: 390-404.
77. Seiwert TY, Fayette J, Cupissol D, Del CJM, Clement PM, Hitt R, Degardin M, Zhang W, Blackman A, Ehrnrooth E, and Cohen EE. A randomized, phase II study of afatinib versus cetuximab in metastatic or recurrent squamous cell carcinoma of the head and neck. Ann Oncol. 2014; 25: 1813-20.
78. Machiels JP, Subramanian S, Ruzsa A, Repassy G, Lifirenko I, Flygare A, Sørensen P, Nielsen T, Lisby S, and Clement PM. Zalutumumab plus best supportive care versus best supportive care alone in patients with recurrent or metastatic squamous-cell carcinoma of the head and neck after failure of platinum-based chemotherapy: an open-label, randomised phase 3 trial. Lancet Oncol. 2011; 12: 333-43.
79. Brizel DM. Targeting the future in head and neck cancer. Lancet Oncol. 2009; 10: 204-5.
80. Kundu SK, and Nestor M. Targeted therapy in head and neck cancer. Tumour Biol. 2012; 33: 707-21.
81. Siu LL, Soulieres D, Chen EX, Pond GR, Chin SF, Francis P, Harvey L, Klein M, Zhang W, Dancey J, Eisenhauer EA, and Winquist E. Phase I/II trial of erlotinib and cisplatin in patients with recurrent or metastatic squamous cell carcinoma of the head and neck: a Princess Margaret Hospital phase II consortium and National Cancer Institute of Canada Clinical Trials Group Study. J Clin Oncol. 2007; 25: 2178-83.
82. Cleary JM, Reardon DA, Azad N, Gandhi L, Shapiro GI, Chaves J, Pedersen M, Ansell P, Ames W, Xiong H, Munasinghe W, Dudley M, Reilly EB, et al. A phase 1 study of ABT-806 in subjects with advanced solid tumors. Invest New Drugs. 2015; 33: 671-8.
83. Stewart JS, Cohen EE, Licitra L, Van Herpen CM, Khorprasert C, Soulieres D, Vodvarka P, Rischin D, Garin AM, Hirsch FR, Varella-Garcia M, Ghiorghiu S, Hargreaves L, et al. Phase III study of gefitinib compared with intravenous methotrexate for recurrent squamous cell carcinoma of the head and neck [corrected]. J Clin Oncol. 2009; 27: 1864-71.
84. Ferrarotto R, and Gold KA. Afatinib in the treatment of head and neck squamous cell carcinoma. Expert Opin Investig Drugs. 2014; 23: 135-43.
85. Cohen RB. Current challenges and clinical investigations of epidermal growth factor receptor (EGFR)- and ErbB family-targeted agents in the treatment of head and neck squamous cell carcinoma (HNSCC). Cancer Treat Rev. 2014; 40: 567-77.
86. Knutzen G, and Subbiah S. Cetuximab Rechallenge and Monotherapy in Patients with Squamous Cell Carcinoma of the Head and Neck. Case Rep Oncol. 2015; 8: 503-8.
87. Reeves TD, Hill EG, Armeson KE, and Gillespie MB. Cetuximab therapy for head and neck squamous cell carcinoma: a systematic review of the data. Otolaryngol Head Neck Surg. 2011; 144: 676-84.
88. Pollock NI, and Grandis JR. HER2 as a therapeutic target in head and neck squamous cell carcinoma. Clin Cancer Res. 2015; 21: 526-33.
89. Kruser TJ, and Wheeler DL. Mechanisms of resistance to HER family targeting antibodies. Exp Cell Res. 2010; 316: 1083-100.
90. Yarden Y, and Sliwkowski MX. Untangling the ErbB signalling network. Nat Rev Mol Cell Biol. 2001; 2: 127-37.
91. Hudis CA. Trastuzumab—mechanism of action and use in clinical practice. N Engl J Med. 2007; 357: 39-51.
92. Medina PJ, and Goodin S. Lapatinib: a dual inhibitor of human epidermal growth factor receptor tyrosine kinases. Clin Ther. 2008; 30: 1426-47.
93. Williams JP, Kim I, Ito E, Shi W, Yue S, Siu LL, Waldron J, O’Sullivan B, Yip KW, and Liu FF. Pre-clinical characterization of Dacomitinib (PF-00299804), an irreversible pan-ErbB inhibitor, combined with ionizing radiation for head and neck squamous cell carcinoma. PLoS One. 2014; 9: e98557.
94. Carla C, Daris F, Cecilia B, Francesca B, Francesca C, and Paolo F. Angiogenesis in head and neck cancer: a review of the literature. J Oncol. 2012; 2012: 358472.
95. Olsson AK, Dimberg A, Kreuger J, and Claesson-Welsh L. VEGF receptor signalling - in control of vascular function. Nat Rev Mol Cell Biol. 2006; 7: 359-71.
96. Vassilakopoulou M, Psyrri A, and Argiris A. Targeting angiogenesis in head and neck cancer. Oral Oncol. 2015; 51: 409-15.
97. Hsu HW, Wall NR, Hsueh CT, Kim S, Ferris RL, Chen CS, and Mirshahidi S. Combination antiangiogenic therapy and radiation in head and neck cancers. Oral Oncol. 2014; 50: 19-26.
98. Machiels JP, Henry S, Zanetta S, Kaminsky MC, Michoux N, Rommel D, Schmitz S, Bompas E, Dillies AF, Faivre S, Moxhon A, Duprez T, and Guigay J. Phase II study of sunitinib in recurrent or metastatic squamous cell carcinoma of the head and neck: GORTEC 2006-01. J Clin Oncol. 2010; 28: 21-8.
99. Choong NW, Kozloff M, Taber D, Hu HS, Wade J, Ivy P, Karrison TG, Dekker A, Vokes EE, and Cohen EE. Phase II study of sunitinib malate in head and neck squamous cell carcinoma. Invest New Drugs. 2010; 28: 677-83.
100. Freudlsperger C, Burnett JR, Friedman JA, Kannabiran VR, Chen Z, and Van Waes C. EGFR-PI3K-AKT-mTOR signaling in head and neck squamous cell carcinomas: attractive targets for molecular-oriented therapy. Expert Opin Ther Targets. 2011; 15: 63-74.
101. Thariat J, Vignot S, Lapierre A, Falk AT, Guigay J, Van Obberghen-Schilling E, and Milano G. Integrating genomics in head and neck cancer treatment: Promises and pitfalls. Crit Rev Oncol Hematol. 2015; 95: 397-406.
102. Polivka J, and Janku F. Molecular targets for cancer therapy in the PI3K/AKT/mTOR pathway. Pharmacol Ther. 2014; 142: 164-75.
103. Nguyen SA, Walker D, Gillespie MB, Gutkind JS, and Day TA. mTOR inhibitors and its role in the treatment of head and neck squamous cell carcinoma. Curr Treat Options Oncol. 2012; 13: 71-81.
104. Clark C, Shah S, Herman-Ferdinandez L, Ekshyyan O, Abreo F, Rong X, McLarty J, Lurie A, Milligan EJ, and Nathan CO. Teasing out the best molecular marker in the AKT/mTOR pathway in head and neck squamous cell cancer patients. Laryngoscope. 2010; 120: 1159-65.
105. Mazumdar T, Sen B, Wang Y, Peng S, Nicholas C, Glisson BS, Myers JN, and Johnson FM. Drug-induced RAF dimerization is independent of RAS mutation status and does not lead to universal MEK dependence for cell survival in head and neck cancers. Anticancer Drugs. 2015; 26: 835-42.
106. Sen M, Pollock NI, Black J, DeGrave KA, Wheeler S, Freilino ML, Joyce S, Lui VW, Zeng Y, Chiosea SI, and Grandis JR. JAK kinase inhibition abrogates STAT3 activation and head and neck squamous cell carcinoma tumor growth. Neoplasia. 2015; 17: 256-64.
107. Geiger JL, Grandis JR, and Bauman JE. The STAT3 pathway as a therapeutic target in head and neck cancer: Barriers and innovations. Oral Oncol. 2016; 56: 84-92.
108. Lai SY, and Johnson FM. Defining the role of the JAK-STAT pathway in head and neck and thoracic malignancies: implications for future therapeutic approaches. Drug Resist Updat. 2010; 13: 67-78.
109. Martinez VD, Vucic EA, Thu KL, Pikor LA, Lam S, and Lam WL. Disruption of KEAP1/CUL3/RBX1 E3-ubiquitin ligase complex components by multiple genetic mechanisms: Association with poor prognosis in head and neck cancer. Head Neck. 2015; 37: 727-34.
110. Parsel SM, Grandis JR, and Thomas SM. Nucleic acid targeting: towards personalized therapy for head and neck cancer. Oncogene. 2015.
111. Nozawa H, Tadakuma T, Ono T, Sato M, Hiroi S, Masumoto K, and Sato Y. Small interfering RNA targeting epidermal growth factor receptor enhances chemosensitivity to cisplatin, 5-fluorouracil and docetaxel in head and neck squamous cell carcinoma. Cancer Sci. 2006; 97: 1115-24.
112. Yanamoto S, Kawasaki G, Yoshitomi I, Iwamoto T, Hirata K, and Mizuno A. Clinicopathologic significance of EpCAM expression in squamous cell carcinoma of the tongue and its possibility as a potential target for tongue cancer gene therapy. Oral Oncol. 2007; 43: 869-77.
113. Seiwert TY, Jagadeeswaran R, Faoro L, Janamanchi V, Nallasura V, El DM, Yala S, Kanteti R, Cohen EE, Lingen MW, Martin L, Krishnaswamy S, Klein-Szanto A, et al. The MET receptor tyrosine kinase is a potential novel therapeutic target for head and neck squamous cell carcinoma. Cancer Res. 2009; 69: 3021-31.
114. Chung CH, Parker JS, Karaca G, Wu J, Funkhouser WK, Moore D, Butterfoss D, Xiang D, Zanation A, Yin X, Shockley WW, Weissler MC, Dressler LG, et al. Molecular classification of head and neck squamous cell carcinomas using patterns of gene expression. Cancer Cell. 2004; 5: 489-500.
115. Chen GX, Zhang S, He XH, Liu SY, Ma C, and Zou XP. Clinical utility of recombinant adenoviral human p53 gene therapy: current perspectives. Onco Targets Ther. 2014; 7: 1901-9.
116. Peng Z. Current status of gendicine in China: recombinant human Ad-p53 agent for treatment of cancers. Hum Gene Ther. 2005; 16: 1016-27.
117. Chang SS, Jiang WW, Smith I, Poeta LM, Begum S, Glazer C, Shan S, Westra W, Sidransky D, and Califano JA. MicroRNA alterations in head and neck squamous cell carcinoma. Int J Cancer. 2008; 123: 2791-7.
118. Negrini M, Nicoloso MS, and Calin GA. MicroRNAs and cancer—new paradigms in molecular oncology. Curr Opin Cell Biol. 2009; 21: 470-9.
119. Graveland AP, Braakhuis BJ, Eerenstein SE, de Bree R, Bloemena E, de Maaker M, van den Brekel MW, Dijk F, Mesker WE, Tanke HJ, Leemans CR, and Brakenhoff RH. Molecular diagnosis of minimal residual disease in head and neck cancer patients. Cell Oncol (Dordr). 2012; 35: 367-75.
120. de Carvalho AC, Kowalski LP, Campos AH, Soares FA, Carvalho AL, and Vettore AL. Clinical significance of molecular alterations in histologically negative surgical margins of head and neck cancer patients. Oral Oncol. 2012; 48: 240-8.
121. Roh JL, Westra WH, Califano JA, Sidransky D, and Koch WM. Tissue imprint for molecular mapping of deep surgical margins in patients with head and neck squamous cell carcinoma. Head Neck. 2012; 34: 1529-36.
122. Yasui W, Tahara E, Tahara H, Fujimoto J, Naka K, Nakayama J, Ishikawa F, Ide T, and Tahara E. Immunohistochemical detection of human telomerase reverse transcriptase in normal mucosa and precancerous lesions of the stomach. Jpn J Cancer Res. 1999; 90: 589-95.
123. Gudkov SV, Shilyagina NY, Vodeneev VA, and Zvyagin AV. Targeted Radionuclide Therapy of Human Tumors. Int J Mol Sci. 2016; 17.
124. Pouget JP, Lozza C, Deshayes E, Boudousq V, and Navarro-Teulon I. Introduction to radiobiology of targeted radionuclide therapy. Front Med (Lausanne). 2015; 2: 12.
125. Sok JC, Coppelli FM, Thomas SM, Lango MN, Xi S, Hunt JL, Freilino ML, Graner MW, Wikstrand CJ, Bigner DD, Gooding WE, Furnari FB, and Grandis JR. Mutant epidermal growth factor receptor (EGFRvIII) contributes to head and neck cancer growth and resistance to EGFR targeting. Clin Cancer Res. 2006; 12: 5064-73.
126. Nestor MV. Targeted radionuclide therapy in head and neck cancer. Head Neck. 2010; 32: 666-78.
127. Martin D, Abba MC, Molinolo AA, Vitale-Cross L, Wang Z, Zaida M, Delic NC, Samuels Y, Lyons JG, and Gutkind JS. The head and neck cancer cell oncogenome: a platform for the development of precision molecular therapies. Oncotarget. 2014; 5: 8906-23. doi: 10.18632/oncotarget.2417.
128. Beloueche-Babari M, Box C, Arunan V, Parkes HG, Valenti M, De Haven Brandon A, Jackson LE, Eccles SA, and Leach MO. Acquired resistance to EGFR tyrosine kinase inhibitors alters the metabolism of human head and neck squamous carcinoma cells and xenograft tumours. Br J Cancer. 2015; 112: 1206-14.
129. Boeckx C, de Beeck K O, Wouters A, Deschoolmeester V, Limame R, Zwaenepoel K, Specenier P, Pauwels P, Vermorken JB, Peeters M, Van Camp G, Baay M, and Lardon F. Overcoming cetuximab resistance in HNSCC: the role of AURKB and DUSP proteins. Cancer Lett. 2014; 354: 365-77.
130. Ku BM, Yi SY, Koh J, Bae YH, Sun JM, Lee SH, Ahn JS, Park K, and Ahn MJ. The CDK4/6 inhibitor LY2835219 has potent activity in combination with mTOR inhibitor in head and neck squamous cell carcinoma. Oncotarget. 2016; 7: 14803-13. doi: 10.18632/oncotarget.7543.
131. Bozec A, Ebran N, Radosevic-Robin N, Sudaka A, Monteverde M, Toussan N, Etienne-Grimaldi MC, Nigro CL, Merlano M, Penault-Llorca F, and Milano G. Combination of mTOR and EGFR targeting in an orthotopic xenograft model of head and neck cancer. Laryngoscope. 2016; 126: E156-63.
132. Wang Z, Martin D, Molinolo AA, Patel V, Iglesias-Bartolome R, Degese MS, Vitale-Cross L, Chen Q, and Gutkind JS. mTOR co-targeting in cetuximab resistance in head and neck cancers harboring PIK3CA and RAS mutations. J Natl Cancer Inst. 2014; 106.

