INTRODUCTION
Renal cell carcinoma (RCC) is estimated to be the ninth leading cause of cancers in the US [1]. Three subtypes taking up 95% cases of RCC are clear cell RCC (KIRC), kidney papillary carcinoma (KIRP) and kidney chromophobe (KICH) [2]. The five-year overall survival rate of RCC is about 74%. The prognosis of RCC patients is closely related to patients’ age, tumor grade, and TNM stage [3]. Recently, mutations in PBRM1, BAP1 and SETD2 are identified to be the molecular biomarkers for the prognosis of RCC [4, 5]. Beyond predicting prognosis, molecular biomarkers may also provide tumorigenic characteristics that are useful for the development of novel anti-RCC therapies [5].
Programmed cell death 1 ligand 1 (PD-L1, CD274, B7-H1) expressed on antigen presenting cells, B cells and other tissue cells can bind its receptor PD-1 on T cells, B cells and myeloid cells to negatively regulate immune responses [6]. In RCC patients, PD-L1 expressed on tumor cells detected by immunohistochemistry was considered to be a risk factor for prognosis, but other studies found that higher PD-L1 mRNA level in RCC tissues estimated by RNA-seq approach was recognized as an indicator of favorable prognosis [7–12]. Extensive studies are therefore required to compromise the contrary results.
The prognosis of locally advanced or metastatic RCC is poor. Targeting therapy directly inhibiting the specific molecules such as tyrosine kinase or mammalian target of rapamycin (mTOR) has better clinical responses than cytokine therapy, but many patients become refractory to these therapies after a period of the treatment [2]. Recently, checkpoint inhibitors targeting PD-1 or its ligand have been introduced and the clinical trial is ongoing [6, 13, 14]. Primary results indicate that clinical response rate to the checkpoint inhibitors ranges from 11.7-29% in RCC patients [6]. To improve the prognosis of advanced RCC patients, the optimized regimens of systemic therapies need to be explored.
In this study, we aimed to investigate the role of PD-L1 mRNA expression in tumors in predicting the outcome of RCC based on the analyses of the clinical and RNA-seq data presented in The Cancer Genome Atlas (TCGA) database. Gene-set enrichment analysis (GSEA) on the data in TCGA and Gene Expression Omnibus (GEO) databases contributes to comprehend the immunological changes in RCC and to provide potential strategies for systemic therapy of RCC.
RESULTS
Description of the integrated RCC data in TCGA
The integrated data of 522 KIRC, 259 KIRP and 66 KICH patients in TCGA were enrolled for analyses (Supplementary data 1). Demographic, clinical, follow-up and tumor pathological features of the three RCC subtypes are listed in Table 1. Among the three RCC subtypes, 174 (33.3%) KIRC patients, 41(15.8%) KIRP patients and 16 (24.2%) KICH patients died in the follow-up period (Table 1).
Table 1: Patient and tumor characteristics of the three RCC subtype cohorts in TCGA
Variable |
KIRC |
KIRP |
KICH |
|---|---|---|---|
Sample (n) |
522 |
259 |
66 |
Median age (year) |
61 (26-90) |
62 (28-88) |
50 (17-86) |
Median PD-L1 |
40.8 (0-5361.1) |
23.6 (0-640.6) |
67.4 (0.5-2930.8) |
Gender |
|||
Male |
337 (64.6%) |
191 (73.7%) |
39 (50.1%) |
Female |
185 (35.4%) |
68 (26.3%) |
27 (40.9%) |
Laterality |
|||
Left |
248 (47.5%) |
144 (55.6%) |
30 (45.5%) |
Right |
273 (52.3%) |
113 (43.6%) |
36 (54.5%) |
Others |
1 (0.2%) |
2 (0.8%) |
|
Clinical stage |
|||
Stage I |
260 (49.8%) |
172 (66.4%) |
21 (31.8%) |
Stage II |
56 (10.7%) |
21 (8.1%) |
25 (37.9%) |
Stage III |
123 (23.6%) |
51 (19.7%) |
14 (21.2%) |
Stage IV |
83 (15.9%) |
15 (5.8%) |
6 (9.1%) |
Tumor stage |
|||
T1 |
265 (50.8%) |
175 (67.6%) |
21 (31.8%) |
T2 |
68 (13.0%) |
24 (9.3%) |
25 (37.9%) |
T3 |
178 (34.1%) |
56 (21.6%) |
18 (27.3%) |
T4 |
11 (2.1%) |
2 (0.8%) |
2 (3.0%) |
Survival status |
|||
Alive |
348 (66.7%) |
218 (84.2%) |
50 (75.8%) |
Died |
174 (33.3%) |
41 (15.8%) |
16 (24.2%) |
RCC subtype, KIRC: kidney clear cell carcinoma; KIRP: kidney papillary carcinoma; KICH: kidney chromophobe
PD-L1 mRNA levels of the three RCC subtype cohorts were extracted from the RNA-seq2 data, which displayed continuous variables with a wide range of 0 to 5,361.1. The three RCC subtype cohorts were further divided into high PD-L1 group and low PD-L1 group based on the median PD-L1 mRNA value (Table 1) [15, 16].
PD-L1 mRNA level and survival status
In KIRC cohort, patients in high PD-L1 group had a median overall survival of 45.0 months (0-149.1 months) longer than the median overall survival of 37.1 months (0-133.6 months) in low PD-L1 group. The overall survival status is significantly different between high PD-L1 group and low PD-L1 group (HR=0.6, 95% CI 0.5-0.8, p=0.002; Figure 1). However, no significant difference in overall survival status was found between high PD-L1 group and low PD-L1 group in KIRP and KICH cohorts (Figure 1). Then we included the variables including age, gender, laterality, tumor grade, clinical stage, tumor stage, metastasis, and PD-L1 mRNA level into a multivariate Cox regression model and found that PD-L1 mRNA level was an independent predictor for overall survival status of KIRC patients (HR=0.7, 95% CI 0.5-0.9, p=0.007; Table 2).
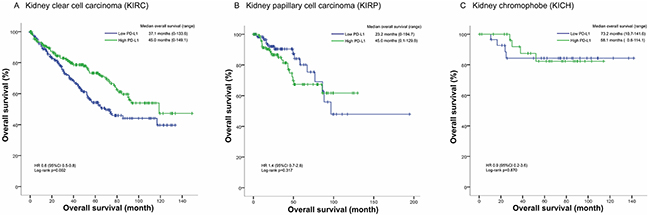
Figure 1: Kaplan-Meier plots of the KIRC, KIRP and KICH cohorts.
Table 2: Univariate and multivariate regression analyses for predicting overall survival in KIRC cohort
Variable |
Univariate |
Multivariate |
||
|---|---|---|---|---|
HR (95% CI) |
p-value |
HR (95% CI) |
p-value |
|
Age |
1.8 (1.3-2.4) |
<0.001* |
1.6 (1.2-2.1) |
0.003* |
Gender |
1.1 (0.8-1.4) |
0.741 |
1.2 (0.9-1.6) |
0.326 |
Laterality |
0.7 (0.5-1.0) |
0.024* |
0.8 (0.6-1.0) |
0.070 |
Tumor grade |
2.2 (1.8-2.7) |
<0.001* |
1.5 (1.2-1.9) |
<0.001* |
Clinical stage |
1.9 (1.6-2.1) |
<0.001* |
1.6 (1.4-1.9) |
<0.001* |
Tumor stage |
1.9 (1.6-2.2) |
<0.001* |
0.8 (0.6-1.1) |
0.109 |
Lymph node metastasis |
0.9 (0.8-1.1) |
0.267 |
0.9 (0.8-1.1) |
0.187 |
Distant metastasis |
2.3 (1.8-2.9) |
<0.001* |
1.1 (0.7-1.8) |
0.643 |
PD-L1 mRNA level |
0.6 (0.5-0.8) |
0.002* |
0.7 (0.5-0.9) |
0.007* |
*: statistically significant to predict overall survival rate
PD-L1 mRNA level and clinical features of KIRC cohort
In addition to the significant difference in overall survival status between low PD-L1 group and high PD-L1 group, no differences were detected in clinical characteristics including age, laterality, clinical stage, tumor stage, metastasis, and tumor grade, except for a higher male ratio in low PD-L1 group (p=0.026, Table 3).
Table 3: Comparison of clinical characteristics between low PD-L1 group and high PD-L1 group in KIRC cohort
Group |
p-Value |
||
|---|---|---|---|
Low PD-L1 |
High PD-L1 |
||
Sample (n) |
261 |
261 |
|
Age (year) |
0.381 |
||
≤61 |
133 (51.0%) |
143 (54.8%) |
|
>61 |
128 (49.0%) |
118 (45.2%) |
|
Gender |
0.022* |
||
Male |
181 (69.3%) |
156 (59.8%) |
|
Female |
80 (30.7%) |
105 (40.2%) |
|
Laterality |
0.601 |
||
Left |
123 (47.1%) |
125 (47.9%) |
|
Right |
137 (52.5%) |
136 (52.1%) |
|
Others |
1 (0.4%) |
0 |
|
Clinical stage |
0.082 |
||
Stage I |
133 (50.9%) |
127 (48.7%) |
|
Stage II |
19 (7.3%) |
37 (14.2%) |
|
Stage III |
66 (25.3%) |
57 (21.8%) |
|
Stage IV |
43 (16.5%) |
40 (15.3%) |
|
Tumor stage |
0.056 |
||
T1 |
135 (51.7%) |
130 (49.8%) |
|
T2 |
26 (10.0%) |
42 (16.1%) |
|
T3 |
97 (37.2%) |
81 (31.0%) |
|
T4 |
3 (1.1%) |
8 (3.1%) |
|
Lymph node metastasis |
0.208 |
||
N0 |
109 (41.8%) |
129 (49.4%) |
|
N1 |
9 (3.4%) |
7 (2.7%) |
|
NX |
143 (54.8%) |
125 (47.9%) |
|
Distant metastasis |
0.620 |
||
M0 |
207 (79.3%) |
214 (82.0%) |
|
M1 |
41 (15.7%) |
38 (14.6%) |
|
MX |
13 (5.0%) |
9 (3.4%) |
|
Tumor grade |
0.609 |
||
G1 |
6 (2.3%) |
6 (2.3%) |
|
G2 |
104 (39.8%) |
121 (46.4%) |
|
G3 |
110 (42.1%) |
95 (36.4%) |
|
G4 |
39 (15.0%) |
36 (13.8%) |
|
GX |
2 (0.8%) |
3 (1.1%) |
|
Survival status |
0.002* |
||
Alive |
157 (60.2%) |
191 (73.2%) |
|
Died |
104 (39.8%) |
70 (26.8%) |
|
*: statistically significant
Gene expression signature in high PD-L1 group and low PD-L1 group of KIRC cohort
We further analyzed the gene expression data in tumors to compare the differences in cell processes such as immune, proliferation, metabolism and DNA damage repair between high PD-L1 group and low PD-L1 group in the KIRC cohort (Table 4). We also performed the same analyses for the 72 KIRC cases in GSE53757 dataset of GEO database to confirm the differences of cell processes between the two groups in KIRC cohort. A total of 10 pathways were upregulated in high PD-L1 group, and a total of 3 pathways were upregulated in low PD-L1 group of KIRC patients in both TCGA and GEO databases. In high PD-L1 group, at least 8 of the 10 upregulated pathways are closely related to immunological functions. In contrast in low PD-L1 group, the 3 upregulated pathways are involved in tumor progression and metastasis (Figure 2).
Table 4: Pathway analyses for high PD-L1 group and low PD-L1 group in KIRC cohort from TCGA and GEO databases
KIRC from TCGA (522 cases) |
KIRC from GSEA 53757 in GEO (72 cases) |
|||||||
|---|---|---|---|---|---|---|---|---|
High PD-L1 |
q-val. |
Low PD-L1 |
q-val. |
High PD-L1 |
q-val. |
Low PD-L1 |
q-val. |
|
1 |
Interferon-γ response* |
<0.001 |
DNA repair |
<0.001 |
Allograft rejection* |
<0.001 |
Epithelial mesenchymal transition* |
<0.001 |
2 |
Allograft rejection* |
<0.001 |
MYC targets v2 |
<0.001 |
Interferon-γ response* |
<0.001 |
Uv response down |
<0.001 |
3 |
Interferon-α response* |
<0.001 |
Myogenesis* |
0.001 |
Interferon-α response* |
<0.001 |
Angiogenesis |
<0.001 |
4 |
Protein secretion |
<0.001 |
MYC targets v1 |
0.004 |
IL6 JAK Stat3 signaling* |
<0.001 |
Myogenesis* |
<0.001 |
5 |
Mitotic spindle |
<0.001 |
Glycolysis |
0.003 |
E2F targets |
<0.001 |
TGF–β signaling |
<0.001 |
6 |
Inflammatory response* |
<0.001 |
Epithelial mesenchymal transition* |
0.008 |
Inflammatory response* |
<0.001 |
Hypoxia |
<0.001 |
7 |
G2M checkpoint* |
0.001 |
Coagulation* |
0.041 |
G2M checkpoint* |
<0.001 |
Notch signaling |
0.002 |
8 |
Androgen response |
0.002 |
Oxidative phosphorylation |
0.043 |
TNF-α signaling via NF-κB* |
<0.001 |
Apical junction |
<0.001 |
9 |
IL6 JAK Stat3 signaling* |
0.006 |
Complement* |
<0.001 |
Wnt β-catenin signaling |
0.004 |
||
10 |
Kras signaling up |
0.005 |
IL2 STAT5 signaling* |
<0.001 |
Hedgehog signaling |
0.018 |
||
11 |
Complement* |
0.010 |
PI3K AKT mTOR signaling* |
0.004 |
Androgen response |
0.008 |
||
12 |
TNF-α signaling via NF-κB* |
0.010 |
Coagulation* |
0.002 |
||||
13 |
Uv response down |
0.014 |
Bile acid metabolism |
0.002 |
||||
14 |
IL2 Stat5 signaling* |
0.018 |
Fatty acid metabolism |
0.004 |
||||
15 |
PI3K AKT mTOR signaling* |
0.019 |
Adipogenesis |
0.004 |
||||
16 |
Xenobiotic metabolism |
0.006 |
||||||
17 |
Kras signaling up |
0.008 |
||||||
18 |
Estrogen response early |
0.012 |
||||||
*: upregulated both in KIRC patients in TCGA and GEO databases; q-val.: FDR q-value
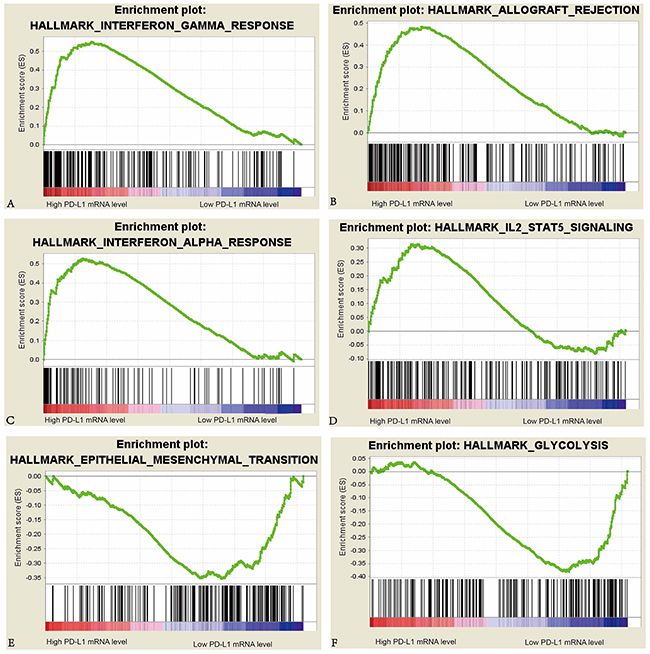
Figure 2: Enrichment plots of interferon-γ response, interferon-α response, epithelial mesenchymal transition, allograft rejection, IL2 Stat5 signaling, and glycolysis against PD-L1 mRNA level in the KIRC cohort.
DISCUSSION
In this study, we identified that PD-L1 mRNA level in tumor tissue was an independent prognosis predictor for KIRC patients and that the activation of functional pathways was different in KIRCs with different PD-L1 mRNA levels.
Previous studies using immunohistochemistry and ELISA to measure PD-L1 protein in tumors and sera reached the conclusion that higher PD-L1 level was associated with poor prognosis of the three subtypes of RCC [7, 8, 17]. Quantification of PD-L1 through the intensity of immunohistochemistry staining by different antibodies may bring ambiguous results [17], and may only represent the PD-L1 expression level in tumor cells. In our present study, we obtained the data of the three main subtypes of RCC from TCGA and processed by the same method. The results revealed that PD-L1 was an independent prognosis predictor for KIRC patients but not for KIRP and KICH patients. Recently, Messai et al. reported that mutations in von Hippel-Lindau (VHL) gene positively correlated with PD-L1 expression in KIRC cells but not in KIRP and KICH cells [18], suggesting that PD-L1 may play different role in different RCC subtypes and that anti-PD-1/PD-L1 therapy may not be suitable for all RCC patients.
It seems paradoxical that higher expression of immunosuppressive PD-L1 correlated with improved outcomes. This will be resolved if PD-L1 expression is viewed as a reflection of the presence of endogenous antitumor immunity [19]. In other words, higher PD-L1 mRNA level in tumors is the negative feedback to the activated antitumor responses such as IFN-γ response, IFN-α response and activated IL2-Stat5 signaling pathway in tumor microenvironment [19]. The outcome of a tumor is determined by the interaction between host antitumor immune responses and negative feedback to the immunological responses in tumor [19]. In the KIRC cohort, patients with active immune responses usually had higher PD-L1 mRNA level in tumors and better outcomes, while those with less active immune responses and increased glycolysis and epithelial-mesenchymal transition had lower PD-L1 mRNA level in tumors and shorter survival periods.
Treatment strategies for tumors with different status of endogenous immune responses should be different [20]. Recently, a prospective study revealed that tumors with higher PD-L1 expression had a better response to high-dose IL-2 than those with negative PD-L1 expression [21]. The tumors with active antitumor immune responses indicate that both innate and adaptive immune responses are strongly activated to eliminate tumor cells with specific antigens on their surfaces [22]. In view of the better response to high-dose IL-2 and the immunosuppressive effect of higher PD-L1 [21], the treatment of KIRC with higher PD-L1 expression should combine the therapies promoting host antitumor immune responses such as IL-2 and blocking the immunosuppressive status such as anti-PD-L1/PD-1 antibody therapies. In contrast in KIRC with lower response to high-dose IL-2 and lower PD-L1 mRNA expression, the weak immune response may attribute to the lack of tumor-specific antigens on tumor cells and the secretion of immunosuppressive cytokines such as VEGF and TGF-β [22]. Molecular target therapy may be useful for these patients. PD-L1 mRNA level in KIRC may be used as a reference for drug treatment strategies of KIRC patients. However, the different treatment regimen we propose for KIRC with different PD-L1 mRNA level in tumors must be tested further by random clinical trials. One limitation of this study is the lack of another independent cohort for validation. In addition, other factors which can also influence the outcome of KIRC patients are not taken into account due to the lack of the data, such as the time of disease recurrence after surgery and the treatment for the patients.
In conclusion, our study provides a new insight into the significance of PD-L1 in KIRC. Higher PD-L1 mRNA level was associated with a better outcome of the patients. The underlying mechanism may be the higher antitumor immune responses in the microenvironment of KIRC. PD-L1 mRNA level in tumor may be one of the factors affecting the outcome of KIRC patients, and may also be a reference for drug treatment strategy for these patients.
MATERIALS AND METHODS
Patients and data collection
Clinical, follow-up and RNA-seq data of the 536 KIRC, 291 KIRP and 66 KICH patients were obtained from TCGA by cBioportal platform and TCGA-Assembler [23, 24]. The patients with integrated clinical stage, T stage, overall survival information and mRNA levels in tumor were enrolled in this study. mRNA expression profiling by array of the 72 KIRC tumors in GSE53757 dataset in GEO were also included [25]. The data used in this study are opened to public for access without limitation and restriction. This study was performed according to the publication guidelines provided by TCGA (http://cancergenome.nih.gov/publications/publicationguidelines).
Pathway analysis
Gene-set enrichment analysis (GSEA) was used to identify the pathways in two different PD-L1 mRNA level groups [26]. RNA-seq data were processed by TCGA-Assembler, and a total of 20,486 genes were enrolled for GSEA analyses. In addition, 20,282 genes from GSE53757 dataset were used for validating the pathway analyses. In the processes of GSEA analyses, the hallmark gene sets (h.all.v5.1.symbols.gmt) were used [27]. The p value of GSEA was computed by 1,000-gene-set two-sided permutation test.
Statistical methods
KIRC, KIRP and KICH patients were divided into two groups according to the median value of PD-L1 mRNA level in tumors. Comparisons of demographic, clinical and pathological features between the two PD-L1 mRNA level groups were conducted using chi-square test or Fisher exact test. Overall survival was assessed using Kaplan-Meier method and log-rank test. Hazard ratio (HR) was calculated by Cox regression model and the result was provided as HR value and 95% confidence interval of the HR. In order to investigate whether PD-L1 level was an independent predictor for outcome in KIRC cohort, we included all the variables in multivariate Cox regression test using a backward conditional approach and eliminated the variables that the p value was >0.05. FDR q value was used for the evaluation of different pathways in different groups. Statistical tests were analyzed using SPSS 19.0 software (SPSS, Inc., Chicago, IL, USA). The p value of <0.05 was considered to be statistically significant.
ACKNOWLEDGMENTS
We thank Ding-fang Bu, Medical Experiment Center, Peking University First Hospital and Yuan-yuan Qi, Renal Division, Peking University First Hospital, for critical reading and revising the manuscript. The results shown here are in whole or part based upon data generated by the TCGA Research Network: http://cancergenome.nih.gov/.
CONFLICTS OF INTEREST
No potential conflicts of interest are disclosed.
GRANT SUPPORT
This research was supported by the grants from National Natural Science Foundation of China (Grant Number: 81172418) and Beijing Natural Science Foundation (Grant Number: 7142160).
REFERENCES
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. Cancer J Clin. 2016; 66: 7-30.
2. Shuch B, Amin A, Armstrong AJ, Eble JN, Ficarra V, Lopez-Beltran A, Martignoni G, Rini BI, Kutikov A. Understanding pathologic variants of renal cell carcinoma: distilling therapeutic opportunities from biologic complexity. Eur Urol. 2015; 67: 85-97.
3. Tan MH, Li H, Choong CV, Chia KS, Toh CK, Tang T, Tan PH, Wong CF, Lau W, Cheng C. The Karakiewicz nomogram is the most useful clinical predictor for survival outcomes in patients with localized renal cell carcinoma. Cancer. 2011; 117: 5314-24.
4. Kapur P, Pena-Llopis S, Christie A, Zhrebker L, Pavia-Jimenez A, Rathmell WK, Xie XJ, Brugarolas J. Effects on survival of BAP1 and PBRM1 mutations in sporadic clear-cell renal-cell carcinoma: a retrospective analysis with independent validation. Lancet Oncol. 2013; 14: 159-67.
5. Hakimi AA, Ostrovnaya I, Reva B, Schultz N, Chen YB, Gonen M, Liu H, Takeda S, Voss MH, Tickoo SK, Reuter VE, Russo P, Cheng EH, et al. Adverse outcomes in clear cell renal cell carcinoma with mutations of 3p21 epigenetic regulators BAP1 and SETD2: a report by MSKCC and the KIRC TCGA research network. Clin Cancer Res. 2013; 19: 3259-67.
6. Zou W, Wolchok JD, Chen L. PD-L1 (B7-H1) and PD-1 pathway blockade for cancer therapy: Mechanisms, response biomarkers, and combinations. Sci Transl Med. 2016; 8: 328rv4.
7. Thompson RH, Kuntz SM, Leibovich BC, Dong H, Lohse CM, Webster WS, Sengupta S, Frank I, Parker AS, Zincke H, Blute ML, Sebo TJ, Cheville JC, et al. Tumor B7-H1 is associated with poor prognosis in renal cell carcinoma patients with long-term follow-up. Cancer Res. 2006; 66: 3381-5.
8. Choueiri TK, Fay AP, Gray KP, Callea M, Ho TH, Albiges L, Bellmunt J, Song J, Carvo I, Lampron M, Stanton ML, Hodi FS, McDermott DF, et al. PD-L1 expression in nonclear-cell renal cell carcinoma. Ann Oncol. 2014; 25: 2178-84.
9. Peters I, Tezval H, Kramer MW, Wolters M, Grunwald V, Kuczyk MA, Serth J. [Implications of TCGA Network Data on 2nd Generation Immunotherapy Concepts Based on PD-L1 and PD-1 Target Structures]. Aktuelle Urol. 2015; 46: 481-5.
10. Thompson RH, Gillett MD, Cheville JC, Lohse CM, Dong H, Webster WS, Krejci KG, Lobo JR, Sengupta S, Chen L, Zincke H, Blute ML, Strome SE, et al. Costimulatory B7-H1 in renal cell carcinoma patients: Indicator of tumor aggressiveness and potential therapeutic target. Proc Natl Acad Sci U S A. 2004; 101: 17174-9.
11. Thompson RH, Gillett MD, Cheville JC, Lohse CM, Dong H, Webster WS, Chen L, Zincke H, Blute ML, Leibovich BC, Kwon ED. Costimulatory molecule B7-H1 in primary and metastatic clear cell renal cell carcinoma. Cancer. 2005; 104: 2084-91.
12. Chen F, Zhang Y, Senbabaoglu Y, Ciriello G, Yang L, Reznik E, Shuch B, Micevic G, De Velasco G, Shinbrot E, Noble MS, Lu Y, Covington KR, et al. Multilevel Genomics-Based Taxonomy of Renal Cell Carcinoma. Cell Rep. 2016; 14: 2476-89.
13. Motzer RJ, Escudier B, McDermott DF, George S, Hammers HJ, Srinivas S, Tykodi SS, Sosman JA, Procopio G, Plimack ER, Castellano D, Choueiri TK, Gurney H, et al. Nivolumab versus Everolimus in Advanced Renal-Cell Carcinoma. N Engl J Med. 2015; 373: 1803-13.
14. McDermott DF, Drake CG, Sznol M, Choueiri TK, Powderly JD, Smith DC, Brahmer JR, Carvajal RD, Hammers HJ, Puzanov I, Hodi FS, Kluger HM, Topalian SL, et al. Survival, Durable Response, and Long-Term Safety in Patients With Previously Treated Advanced Renal Cell Carcinoma Receiving Nivolumab. J Clin Oncol. 2015; 33: 2013-20.
15. Su H, Sun T, Wang H, Shi G, Zhang H, Sun F, Ye D. Decreased TCL6 expression is associated with poor prognosis in patients with clear cell renal cell carcinoma. Oncotarget. 2017; 8: 5789-5799. doi: 10.18632/oncotarget.11011.
16. Gyorffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, Li Q, Szallasi Z. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat. 2010; 123: 725-31.
17. Xu F, Xu L, Wang Q, An G, Feng G, Liu F. Clinicopathological and prognostic value of programmed death ligand-1 (PD-L1) in renal cell carcinoma: a meta-analysis. Int J Clin Exp Med. 2015; 8: 14595-603.
18. Messai Y, Gad S, Noman MZ, Le Teuff G, Couve S, Janji B, Kammerer SF, Rioux-Leclerc N, Hasmim M, Ferlicot S, Baud V, Mejean A, Mole DR, et al. Renal Cell Carcinoma Programmed Death-ligand 1, a New Direct Target of Hypoxia-inducible Factor-2 Alpha, is Regulated by von Hippel-Lindau Gene Mutation Status. Eur Urol. 2015.
19. Taube JM, Anders RA, Young GD, Xu H, Sharma R, McMiller TL, Chen S, Klein AP, Pardoll DM, Topalian SL, Chen L. Colocalization of inflammatory response with B7-h1 expression in human melanocytic lesions supports an adaptive resistance mechanism of immune escape. Sci Transl Med. 2012; 4: 127ra37.
20. Pardoll DM. The blockade of immune checkpoints in cancer immunotherapy. Nat Rev Cancer. 2012; 12: 252-64.
21. McDermott DF, Cheng SC, Signoretti S, Margolin KA, Clark JI, Sosman JA, Dutcher JP, Logan TF, Curti BD, Ernstoff MS, Appleman L, Wong MK, Khushalani NI, et al. The high-dose aldesleukin “select” trial: a trial to prospectively validate predictive models of response to treatment in patients with metastatic renal cell carcinoma. Clin Cancer Res. 2015; 21: 561-8.
22. Schreiber RD, Old LJ, Smyth MJ. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science. 2011; 331: 1565-70.
23. Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, Cerami E, Sander C, Schultz N. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013; 6: pl1.
24. Zhu Y, Qiu P, Ji Y. TCGA-assembler: open-source software for retrieving and processing TCGA data. Nat Methods. 2014; 11: 599-600.
25. von Roemeling CA, Radisky DC, Marlow LA, Cooper SJ, Grebe SK, Anastasiadis PZ, Tun HW, Copland JA. Neuronal pentraxin 2 supports clear cell renal cell carcinoma by activating the AMPA-selective glutamate receptor-4. Cancer Res. 2014; 74: 4796-810.
26. Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005; 102: 15545-50.
27. Liberzon A, Birger C, Thorvaldsdottir H, Ghandi M, Mesirov JP, Tamayo P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015; 1: 417-25.

