INTRODUCTION
Rapid advances in cancer research have enabled the identification and characterization of driver mutations, expression changes and structural variations in genes that should translate into better selection of therapies for patients. Remarkable successes in this field include the FDA approval of Trastuzumab for breast cancer patients overexpressing the HER2 protein [1] and Erlotinib for metastatic non-small cell lung cancer patients (NSCLC) with exon 19 deletion or exon 21 (L858R) substitution in the EGFR gene [2]. However, not all tumors have actionable genomic alterations and/or available matching targeted therapies. For example, majority of pancreatic cancers that harbor activating KRAS mutations have no effective targeted therapies [3]. Therefore, despite the promise of targeted therapies, chemotherapy still remains the most widely used standard of treatment for both advanced, metastatic cancer patients, and in the adjuvant setting.
While the use of predictive biomarkers in personalizing targeted cancer therapies is common, it is not yet a standard approach for chemotherapy. This is primarily due to the lack of compelling evidence from biomarker-driven studies to support their clinical utility [4–6]. Therefore, there is a clear need to comprehensively curate and evaluate literature-based datasets within an evidence framework to determine the quantity and quality of evidence supporting or refuting the clinical utility of these biomarkers. Such evidence frameworks currently exist to determine the clinical utility of predictive biomarkers for targeted therapies [7–9]. However, to our knowledge, this approach has not been used to assess the clinical utility of chemopredictive biomarkers. Our goal was to perform an exhaustive literature review to assess the overall levels of evidence supporting the clinical utility of a shortlist of chemopredictive biomarkers, using an evidence framework that is based on widely accepted guidelines. We hope this work can serve as a reference to evaluate future predictive biomarkers published in the field.
Numerous studies have shown that the expression of DNA repair genes like ERCC1, β-tubulins or topoisomerases can predict response to platinum, taxanes and other cytotoxic agents respectively [10–12]. We identified 7 biomarkers that have been evaluated in a number of tumor types (Table 1) for their role in predicting response to commonly approved chemotherapy drugs in the first-line, advanced, metastatic and adjuvant treatment settings. These biomarker therapy combinations are ERCC1–platinum drugs, RRM1–gemcitabine, TUBB3–taxanes, TYMS–5-fluorocuracil (5-FU)/Capecitabine, MGMT–temozolomide, TOP1–irinotecan/topotecan and TOP2A–anthracyclines. Several CLIA and/or CAP certified molecular diagnostic assays measure expression levels of these biomarkers in cancer tissue to determine their sensitivity to chemotherapy drugs. Therefore, we specifically examined how gene or protein expression changes in these biomarkers affect sensitivity or resistance to the associated chemotherapy agents.
We conducted a comprehensive literature review from 1990-2015 on the predictive effect of gene or protein expression of these 7 biomarkers and associated chemotherapy drugs. The information from each study was systematically organized to minimize bias and maximize retrieval of relevant information in order to create a gold standard dataset (data available in supplementary tables). The level of evidence for each study was evaluated within an evidence framework which was adapted from widely accepted guidelines [13–15] Figure 1C. This dataset can inform future automation of information extraction through Natural Language Processing (NLP) approaches. Our results highlight a general need for more and higher quality level I evidence supporting clinical utility of chemopredictive biomarkers. Such an approach can help researchers and clinicians evaluate the clinical utility of chemopredictive biomarkers, thus enabling design of clinical trials and decision-making for patient care.
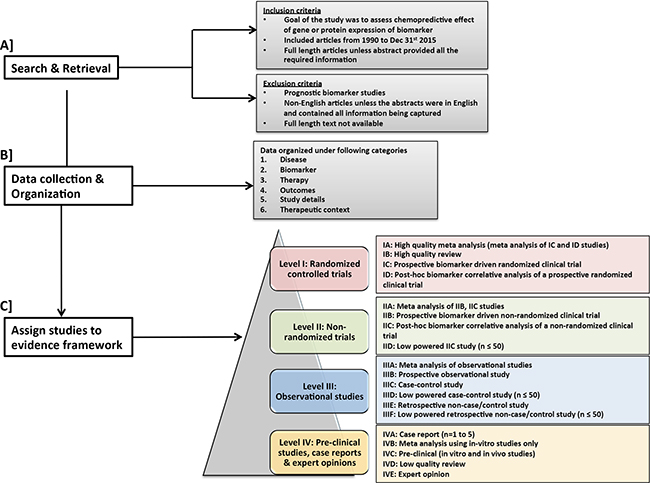
Figure 1: Workflow for search and retrieval and curation. A. Search & retrieval and selection criteria for studies B. data collection and organization and C. Assignment of studies in the proposed evidence framework.
METHODS
Figure 1 shows the overall workflow used to evaluate and summarize the evidence supporting the use of the 7 chemopredictive biomarkers for personalizing cancer treatment
Search and retrieval
We searched PubTator [16] for the following keyword combination: gene/protein name, the term “expression” and “drug name”; for example “TUBB3 expression and taxanes”. Studies published between 1/1/1990 – 12/31/2015 that focused on predictive biomarkers whose gene or protein expression influenced response to chemotherapy were shortlisted for manual curation, Figure 1A. Articles that primarily focused on prognostic biomarkers were excluded. Non-English articles were excluded unless the abstracts contained all the necessary information to populate the framework. Important citations from the selected articles were also curated.
Data collection
The following data elements were collected from each article and organized using controlled vocabulary wherever possible, Figure 1B:
• Disease – type of cancer and stage (early, advanced, metastatic, etc.)
• Biomarker – biomarker name, assay type used, other genes being studied, over and/or under expression status. The expression status is defined by the study and can be based on comparisons with all tumor samples, adjacent normal samples, or normal samples from unaffected individuals.
• Therapy – combinations of chemotherapy drugs that the patient/s was/were given and the therapy setting (first line, second line, adjuvant, etc.).
• Outcomes – outcome measures including, but not limited to, progression-free survival (PFS), disease-free survival, overall survival (OS), tumor response rate and tumor recurrence.
• Study details – model system being studied, for example cell line, animal models or human. For human studies we collected the inclusion, exclusion criteria and sample size, if available. Study metadata such as journal name, year of publication and publication source were also collected.
• Therapeutic context – predictive effect of biomarker expression on therapy outcome was interpreted from the results of the study.
Biocuration
Expert curation and analysis was conducted by a multi-disciplinary group including: oncologists, molecular biologists, translational researchers, a surgeon, biocurators, bioinformaticians and biostatisticians.
Predictive effect of biomarker
The predictive effect of the biomarker was determined from the results and conclusions reported in each study, captured under the following categories:
• Benefit: Over or under expression of the biomarker predicted sensitivity to therapy
• No Benefit: Over or under expression of the biomarker did not predict response to therapy
• Not Assessable: Results of the study were inconclusive for over and/or under expression of the biomarker.
Evidence framework
Evidence levels broadly ranging from I-IV were assigned to each publication based on its study design Figure 1C, adapted from widely accepted guidelines [13–15]. Randomized clinical trials were assigned the highest evidence level I, followed by level II evidence for non-randomized trials, level III evidence for observational studies and lowest evidence level IV for pre-clinical studies, expert opinions and case studies. Within each evidence level, we incorporated sub-levels of evidence based on additional characteristics of study type, including prospective and retrospective analysis and sample size. Meta-analyses were usually assigned the highest sub-level evidence, since they integrate all the available evidence pertaining to a scientific question of interest and quantitatively summarize the results [17]. Depending on the type of studies (e.g. randomized, non-randomized, etc.) included in the meta-analyses, they were categorized under the corresponding evidence levels I-IV. Furthermore, within each evidence category ranging from I-III, prospective studies were ranked higher than retrospective studies. Systematic reviews were carefully selected based on stringent study inclusion criteria and were assigned high evidence level (IB). The term “high quality” meta-analysis or review in the figure refers to these stringent criteria, not to journal impact factor. Expert opinions that had a built-in rigorous meta-analysis were assigned to level I. Reviews including different study types and lacking stringent patient and/or study inclusion criteria were assigned a lower evidence level (IVD). Each study was considered individually for evidence assignment and no effort was made to combine evidence across studies using meta-analysis approaches.
RESULTS
Results from the curation of the 7 chemopredictive biomarker-drug combinations are summarized in Table 1, presenting the total number of studies screened and shortlisted based on our inclusion criteria; the most commonly represented cancers for each biomarker-drug combination and a summary of the overall evidence supporting the predictive effect of each biomarker. Figure 2 shows a breakdown of the number of studies in each evidence level demonstrating the predictive effect of biomarker expression on response to corresponding chemotherapy drugs defined in terms of benefit, no benefit or not assessable. Further details on therapy setting, sample size, outcomes and other study details are available in the supplementary tables.
Table 1: Summary of data for each biomarker-drug combination
Biomarker – Drug combination |
# of studies screened (# of studies curated) |
Most common cancer studied (# of studies) |
Other cancers studied (# of studies, if ≥ 2) |
Overall Evidence |
|---|---|---|---|---|
ERCC1 – Platinum agents |
266 (85) |
Non-small cell lung cancer (43) |
Ovarian cancer (10), Esophageal cancer (5), Small-cell lung cancer (4), Squamous cell head and neck cancers (HNSCC) (3), Colorectal cancer (3), Pancreatic cancer (2), Bladder cancer (2) |
Consistent evidence from levels I-IV retrospective studies |
MGMT – Temozolomide |
366 (55) |
Gliomas (25) |
Pituitary tumors (9), Melanoma (6), Neuroendocrine tumors (2) |
Modest evidence from levels III-IV studies |
RRM1 – Gemcitabine |
131 (55) |
Non-small cell lung cancer (33) |
Pancreatic cancer (7), Breast cancer (2) |
Consistent evidence from levels I-IV retrospective studies |
TS – 5-fluorouracil (5-FU), Capecitabine |
617 (55) |
Colorectal cancer (27) |
Gastric cancer (13), esophageal cancer (5), Hepatocellular cancer (2), Pancreatic cancer (2) |
Modest evidence from levels III-IV studies |
TUBB3 – Taxanes |
61 (40) |
Non-small cell lung cancer (14) |
Breast cancer (9), Gastric cancer (7), Ovarian cancer (3) Melanoma (2) |
Modest evidence from levels III-IV studies |
TOPO1 – Irinotecan, Topotecan |
50 (11) |
Colorectal cancer (5) |
Weak evidence from few level III and IV studies |
|
TOP2A – Anthracyclines |
62 (17) |
Breast cancer (13) |
Hepatocellular carcinoma (2) |
Weak evidence from few level III and IV studies |
Total number of studies screened and curated based on our inclusion criteria; the most commonly represented cancers for each biomarker-drug combination and the overall evidence supporting the predictive effect of each biomarker.
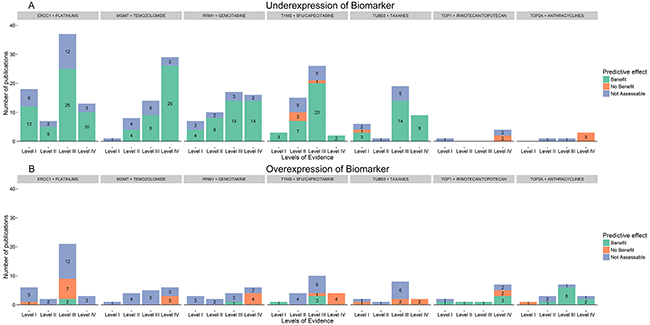
Figure 2. Overall evidence supporting the clinical utility of chemopredictive biomarkers. Overall evidence associated with the predictive effect of biomarker expression on response to corresponding chemotherapy drugs defined in terms of benefit, no benefit or not assessable. Studies where chemotherapy response was not assessable for both, over and under expression of biomarkers have been represented twice, in the over and under expression section. For example, in the RRM1-Gemcitabine plot, a total of six level III studies showed that gemcitabine response is not assessable for both over and under expression of RRM1 and the same six studies are plotted in both over and under expression histograms (blue).
Excision repair cross-complementing group 1 (ERCC1) - platinum drugs
Pre-clinical studies have suggested that underexpression of ERCC1 sensitizes cancer cells to platinum agents whereas overexpression induces resistance [18, 19]. 70% of the studies in our dataset confirm this hypothesis, where the majority of the evidence is from levels I-III retrospective studies. The predictive effect of ERCC1 underexpression on response to platinum agents has been widely studied in NSCLC (Table 1). Out of the two level IC clinical trials in our dataset, one study had no benefit for PFS. In this study, an internal control suggested the possibility of biased randomization, thereby confounding the result, and therefore we have determined the results of this study to be not assessable [20]. A second level IC study confirmed a modest response rate advantage (the primary endpoint), while failing to confirm PFS benefit (secondary endpoint). While we have counted this study as supporting the biomarker hypothesis, a study evaluating the clinical benefit endpoint as primary would be very helpful in determining the clinical significance of this result [21]. Two level III studies showed contradictory evidence in advanced NSCLC and metastatic pancreatic cancer patients where overexpression of ERCC1 showed benefit from platinum based chemotherapy [22, 23]. 22 studies in our dataset had results that were not assessable due to various reasons including different methods of biomarker quantification, disease type, stage and small sample size [24–28] (Supplementary Table S1). While the evidence supporting the chemosensitizing effect of ERCC1 underexpression in response to platinum agents was somewhat consistent across all retrospective studies in NSCLC, the evidence for other cancer types was often inconclusive or contradictory. The overall evidence for other cancer types was mainly from low-level III-IV studies [29–33]. Moreover, the predictive effect of immunostaining for ERCC1 protein has been difficult to confirm in validation studies. This can be attributed to discrepancies in the performance of antibody batches over the years and also a lack of understanding of the heterogeneous expression and function of different ERCC1 protein isoforms [34].
O6-methylguanine DNA methyltransferase (MGMT) - temozolomide
Underexpression of MGMT is thought to sensitize tumor cells to temozolomide (TMZ)-based therapies whereas overexpression induces resistance [35–38]. This hypothesis was widely tested and confirmed in gliomas and pituitary tumors (Table 1, Supplementary Table S2). We found that the overall evidence supporting the chemosensitizing effect of MGMT underexpression on TMZ response is mainly from level IV pre-clinical studies and a few level II-III retrospective studies (Figure 2). The results from 13 studies, including a high-level ID retrospective biomarker study were not assessable [39–42]. In addition to gene/protein expression, we also found several studies that evaluated the predictive effect of MGMT promoter methylation on TMZ response, including the landmark study by Hegi et.al [43] that drives the current clinical use of MGMT as a predictive biomarker for TMZ response in glioblastoma multiforme. Such studies were not included in our dataset since they did not meet our inclusion criteria, which focused only on predictive biomarkers whose gene/protein expression influenced response to chemotherapy. In studies that tested the predictive effect of both MGMT expression and promoter methylation on TMZ response, there was often poor correlation between the two, which resulted in inconclusive results [44].
Ribonucleotide reductase subunit M1 (RRM1) - gemcitabine
Pre-clinical studies have shown that RRM1 underexpression is associated with benefit of gemcitabine-based therapy whereas overexpression is associated with resistance [45–47]. This trend is consistent in 80% of the studies across all evidence levels I-IV. The predictive effect of RRM1 underexpression on benefit of response to gemcitabine was mainly studied in NSCLCs and confirmed by a level IC clinical trial [20]. In pancreatic cancers, the overall evidence supporting the predictive effect of RRM1 was equivocal and investigated only in levels III-IV studies [48–55].
A number of NSCLC studies analyzed the combined predictive effect of both RRM1 and ERCC1 expression, perhaps because treatment for NSCLCs usually includes gemcitabine in combination with platinum drugs. While a level IIA multi-trial NSCLC study confirmed that patients who received personalized first-line therapy based on their RRM1 and ERCC1 gene expression status had better survival than patients on standard therapy [56], a level IIIF study of a three drug concurrent regimen [57] produced results which were contradictory to previous studies, in that responders had low gene expression levels ERCC1 as expected but high levels of RRM1. It can be difficult to determine single marker outcomes from such multi-marker studies.
Thymidylate synthase (TYMS)-5-fluorouracil/capecitabine
Pre-clinical studies have demonstrated that underexpression of TYMS predicts benefit of response to 5-fluorouracil (5-FU)-based therapies whereas overexpression predicts no benefit [58–64]. Our dataset also included studies that tested TYMS expression and response to 5-FU’s prodrug, Capecitabine [10, 64–68]. We found that evidence supporting the chemosensitizing effect of TYMS under expression on 5-FU/Capecitabine is mainly from levels II-III retrospective biomarker analysis of clinical studies (Figure 2). This hypothesis was widely tested and confirmed in gastrointestinal cancers especially colorectal cancers (Table 1, Supplementary Table S4). However, the only level IC clinical trial in our dataset showed contradictory evidence where metastatic colorectal cancer patients overexpressing TYMS had a trend towards better overall survival than the underexpressing cases [69]. The chemosensitizing effect of TYMS underexpression was not assessable in 11 studies whereas 3 studies showed no benefit of TYMS underexpression on 5-FU therapy. Further prospective studies in large well-defined patient populations are necessary to determine the clinical utility of this biomarker. Moreover, ~27 studies in our dataset examined the predictive effect of other biomarkers like TP, DPD and ERCC1 in addition to TYMS in response to 5-FU based therapies. This makes it difficult to determine the predictive effect of TYMS alone on 5-FU response [70].
Class III beta-tubulin (TUBB3)-taxanes
Pre-clinical studies have suggested that underexpression of TUBB3 predicts sensitivity to taxanes whereas overexpression predicts resistance in breast cancer, NSCLC and gastric cancers [71–73]. This hypothesis was mainly supported by 11 level IV pre-clinical studies and 14 level III retrospective studies (Table 1, Figure 2). We did not find any evidence from prospective biomarker driven trials to support the predictive effect of TUBB3 underexpression on response to taxanes. The results from 9 studies that evaluated taxane response in patients under/over expressing TUBB3 were not assessable [74–82]. In a level ID breast cancer study [76], patients overexpressing TUBB3 had a higher probability of response to docetaxel but the predictive effect of TUBB3 underexpression was not assessable. This unusual predictive effect of TUBB3 expression has also been reported in patient populations that received taxanes in an adjuvant or advanced disease setting [83].
Type I topoisomerase (TOP1)-irinotecan/topotecan
Studies on gastrointestinal cancers have suggested that overexpression of TOP1 predicts benefit of camptothecin-based therapies [84–88]. However, the evidence associated with the predictive effect of TOP1 expression status and therapy outcome was mostly of low-level and studies often reported inconclusive results [89–91] (Figure 2). A level ID post-hoc biomarker correlative analysis of the FOCUS trial showed an overall survival benefit in a subset of colorectal cancer patients with moderate/high TOP1 levels measured by immunohistochemistry [92]. However, another similar level ID study conducted by the Dutch Colorectal Cancer Group showed no association between TOP1 expression and survival benefit [93]. Moreover, Meisenberg et al. reported an important correlation between TOP1 and TDP1, suggesting the possible role of other predictive markers in irinotecan/topotecan sensitivity [94].
Topoisomerase (DNA) II alpha (TOP2A) - anthracyclines
The TOP2A gene is overexpressed in several cancer types and is hypothesized to predict sensitivity to anthracycline-based therapies [95, 96]. However, our only high evidence study (level ID) on hepatocellular carcinoma patients showed no benefit of TOP2A overexpression on anthracycline based therapies [97]. 71% of the studies in our dataset showed levels II-IV evidence supporting this hypothesis, where either overexpression of TOP2A was associated with benefit or underexpression was associated with no benefit from anthracycline-based therapy (Figure 2, Supplementary Table S7). Since anthracyclines are frequently used in breast cancer treatments, we found this disease to be most widely represented in our dataset, especially in HER2+ breast cancer. However, we did not find any high evidence level I studies on TOP2A overexpression as a predictor of response to anthracyclines in this review and analysis.
DISCUSSION
Herein, we provided a high level quantitative perspective on the amount and quality of evidence supporting or contradicting the clinical utility of chemopredictive biomarkers. In our evaluation, we found that biomarker-driven prospective clinical trials (levels I-II) for these protein-drug pairs were few and often reported findings that were inconclusive or contradictory. We found somewhat consistent evidence from several retrospective biomarker analyses across levels I-III in NSCLC supporting the chemosensitizing effect of both ERCC1 and RRM1 underexpression in response to platinum and gemcitabine based treatments respectively. The evidence supporting the predictive role of gene/protein expression of TYMS–5FU/Capecitabine in colorectal cancers, TUBB3-taxanes in NSCLC and MGMT–temozolomide in brain tumors was modest and mainly from level III observational studies and level IV pre-clinical studies. There was sparse evidence from level III retrospective studies and level IV pre-clinical studies supporting the chemosensitizing effect of TOP1 and TOP2A overexpression in response to camptothecin-based therapies and anthracyclines in gastrointestinal and breast cancers respectively. Other studies that assessed promoter methylation in MGMT and amplification in TOP1 and TOP2A were not considered in this assessment. Our analysis highlights the need for more well-designed and higher quality level I evidence studies for the 7 chemopredictive biomarker – drug pairs.
Standardizing and organizing relevant information from different biomarker studies presents several challenges 1) author preferences in presenting the information 2) author bias resulting in discrepancies between the summary statements and the actual evidence 3) diversity of methods used to quantitatively measure biomarker levels 4) inconsistency in results and conclusions from different assays, statistical tests or for different endpoints, and 5) inconsistency in clinical information across tumor types. Moreover, since chemotherapy drugs are often given in combination, it may be difficult to determine the predictive effect of a single biomarker-drug combination.
The work summarized herein focuses the effort on a manageable number of studies, which need to be reviewed in greater detail, making such a task feasible. Level I evidence outweighs lower levels of evidence but within the level I evidence framework all studies cannot be considered equal. More detailed curation of such studies is required to rank them based on the strength of evidence they provide. Critical factors to assess within the level I evidence category include study population, endpoints, and appropriate controls; amount of missing data; assay validation; sample collection and processing; statistical and clinical significance of the results, and confounders. Evaluation of these criteria may help to resolve conflicts between studies if one provides stronger evidence than another. Limitations of the current level I studies hamper our ability to draw firm conclusions about the current clinical applicability of these biomarkers.
As more data emerges on novel “actionable” pathways in cancers, this evidence framework will need further development as outlined above to assess clinical actionability. The framework can be applied to enhance the value of the increasing volume of retrospective data collection. However, the framework will likely highlight the need for additional prospective studies and help guide their design. These critical future steps will allow the cancer precision medicine community to collectively evaluate and accept predictive biomarkers for cancer therapy.
ACKNOWLEDGMENTS
The authors would like to thank Dr. Orestis Panagiotou for valuable discussions and feedback on this manuscript. The authors would also like to acknowledge students - Samrat Mukesh Sanghvi and Trisha Miglani for assisting with the preliminary information retrieval on this project.
CONFLICTS OF INTEREST
The authors declare no conflict interest.
FINANCIAL SUPPORT
This research was partly funded by the NIH BD2K grant (1U01HG008390-01), 2015 Pancreatic Cancer Action Network American Association for Cancer Research Acceleration Network Grant (15-90-25-BROD), grant NIH-NCI R21 CA182692 01A1, research funding from the Ruesch Center for the Cure of Gastrointestinal Cancers, Perthera.Inc and the Georgetown University Center of Excellence in Regulatory Science and Innovation (CERSI; U01FD004319), a collaborative effort between the university and the U.S. Food and Drug Administration to promote regulatory science through innovative research and education. This research does not necessarily reflect the views of the FDA.
Author contributions
• Conception and design – SM, JRB, MJP, JLM
• Development of methodology – SRao, SM, RAB, SMB
• Acquisition of data – SRao, SRiazi, CSY
• Analysis and interpretation of data – SM, RAB, MJP, JRB, CSY, SRao
• Writing, review and/or revision of the manuscript – SRao, SM, RAB, MJP, JRB, SMB, SRiazi, CSY, JLM
• Administrative, technical, or material support – SRao, SM
• Study supervision - SM
REFERENCES
1. Slamon DJ, Leyland-Jones B, Shak S, Fuchs H, Paton V, Bajamonde A, Fleming T, Eiermann W, Wolter J, Pegram M, Baselga J and Norton L. Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N Engl J Med. 2001; 344:783-792.
2. Khozin S, Blumenthal GM, Jiang X, He K, Boyd K, Murgo A, Justice R, Keegan P and Pazdur R. U.S. Food and Drug Administration approval summary: Erlotinib for the first-line treatment of metastatic non-small cell lung cancer with epidermal growth factor receptor exon 19 deletions or exon 21 (L858R) substitution mutations. Oncologist. 2014; 19:774-779.
3. Bournet B, Buscail C, Muscari F, Cordelier P and Buscail L. Targeting KRAS for diagnosis, prognosis, and treatment of pancreatic cancer: Hopes and realities. Eur J Cancer. 2016; 54:75-83.
4. Hayes D. Daniel Hayes Leads Tour of Caris Website. The Cancer Letter. 2014; 40.
5. Besse B, Olaussen KA and Soria JC. ERCC1 and RRM1: ready for prime time? J Clin Oncol. 2013; 31:1050-1060.
6. Lustgarten DE, Deshpande C, Aggarwal C, Wang LC, Saloura V, Vachani A, Wang LP, Litzky L, Feldman M, Creaney J, Nowak AK, Langer C, Inghilleri S, et al. Thymidylate synthase and folyl-polyglutamate synthase are not clinically useful markers of response to pemetrexed in patients with malignant pleural mesothelioma. J Thorac Oncol. 2013; 8:469-477.
7. Meric-Bernstam F, Johnson A, Holla V, Bailey AM, Brusco L, Chen K, Routbort M, Patel KP, Zeng J, Kopetz S, Davies MA, Piha-Paul SA, Hong DS, Eterovic AK, Tsimberidou AM, Broaddus R, et al. A decision support framework for genomically informed investigational cancer therapy. J Natl Cancer Inst. 2015; 107.
8. Hayes DF, Bast RC, Desch CE, Fritsche H, Kemeny NE, Jessup JM, Locker GY, Macdonald JS, Mennel RG and Norton L. Tumor marker utility grading system: a framework to evaluate clinical utility of tumor markers. Journal of the National Cancer Institute. 1996; 88:1456-1466.
9. Simon RM, Paik S and Hayes DF. Use of archived specimens in evaluation of prognostic and predictive biomarkers. J Natl Cancer Inst. 2009; 101:1446-1452.
10. Lee S, Park YH, Kim KH, Cho EY, Ahn YC, Kim K, Shim YM, Ahn JS, Park K and Im YH. Thymidine synthase, thymidine phosphorylase, and excision repair cross-complementation group 1 expression as predictive markers of capecitabine plus cisplatin chemotherapy as first-line treatment for patients with advanced oesophageal squamous cell carcinoma. Br J Cancer. 2010; 103:845-851.
11. Stengel C, Newman SP, Leese MP, Potter BV, Reed MJ and Purohit A. Class III beta-tubulin expression and in vitro resistance to microtubule targeting agents. Br J Cancer. 2010; 102:316-324.
12. Pfister TD, Hollingshead M, Kinders RJ, Zhang Y, Evrard YA, Ji J, Khin SA, Borgel S, Stotler H, Carter J, Divelbiss R, Kummar S, Pommier Y, et al. Development and validation of an immunoassay for quantification of topoisomerase I in solid tumor tissues. PLoS One. 2012; 7:e50494.
13. Febbo PG, Ladanyi M, Aldape KD, De Marzo AM, Hammond ME, Hayes DF, Iafrate AJ, Kelley RK, Marcucci G, Ogino S, Pao W, Sgroi DC and Birkeland ML. NCCN Task Force report: Evaluating the clinical utility of tumor markers in oncology. J Natl Compr Canc Netw. 2011; 9:S1-32; quiz S33.
14. Canadian Task Force on the Periodic Health Examination. The periodic health examination. Can Med Assoc J. 1979;121:1193-254.
15. Howick J, Chalmers I, Glasziou P, Greenhalgh T, Heneghan C, Liberati A, Moschetti I, Phillips B, Thornton H, Goddard O, Hodgkinson M. OCEBM Levels of Evidence Working Group. “The Oxford Levels of Evidence 2”. Oxford Centre for Evidence-Based Medicine. 2011.
16. Wei CH, Kao HY and Lu Z. PubTator: a web-based text mining tool for assisting biocuration. Nucleic Acids Res. 2013; 41:W518-522.
17. Armitage P, Berry G and Matthews JNS. Statistical Methods in Medical Research. Blackwell Science. 1971; 641.
18. Gao Y, Su D, Ying L, Lv W and Ma S. [Enhanced cisplatin cytotoxicity by RNA interfering the excision repair cross-complementing gene 1 in lung cancer cell A549/DDP]. Zhongguo Fei Ai Za Zhi. 2010; 13:846-849.
19. Usanova S, Piee-Staffa A, Sied U, Thomale J, Schneider A, Kaina B and Koberle B. Cisplatin sensitivity of testis tumour cells is due to deficiency in interstrand-crosslink repair and low ERCC1-XPF expression. Mol Cancer. 2010; 9:248.
20. Bepler G, Williams C, Schell MJ, Chen W, Zheng Z, Simon G, Gadgeel S, Zhao X, Schreiber F, Brahmer J, Chiappori A, Tanvetyanon T, Pinder-Schenck M, et al. Randomized international phase III trial of ERCC1 and RRM1 expression-based chemotherapy versus gemcitabine/carboplatin in advanced non-small-cell lung cancer. J Clin Oncol. 2013; 31:2404-2412.
21. Cobo M, Isla D, Massuti B, Montes A, Sanchez JM, Provencio M, Vinolas N, Paz-Ares L, Lopez-Vivanco G, Munoz MA, Felip E, Alberola V, Camps C, et al. Customizing cisplatin based on quantitative excision repair cross-complementing 1 mRNA expression: a phase III trial in non-small-cell lung cancer. J Clin Oncol. 2007; 25:2747-2754.
22. Jian-Wei B, Yi-Min M, Yu-Xia S and Shi-Qing L. Expression levels of ERCC1 and RRM1 mRNA and clinical outcome of advanced non-small cell lung cancer. Pak J Med Sci. 2013; 29:1158-1161.
23. Fuereder T, Stift J, Kuehrer I, Stranzl N, Hoeflmayer D, Kornek G and Scheithauer W. Response to GEMOX plus erlotinib in pancreatic cancer is associated with ERCC1 overexpression. Eur J Clin Invest. 2014; 44:958-964.
24. Lee HW, Choi YW, Han JH, Kim JH, Jung JH, Jeong SH, Kang SY, Choi JH, Oh YT, Park KJ, Hwang SC and Sheen SS. Expression of excision repair cross-complementation group 1 protein predicts poor outcome in advanced non-small cell lung cancer patients treated with platinum-based doublet chemotherapy. Lung Cancer. 2009; 65:377-382.
25. Hoffmann AC, Wild P, Leicht C, Bertz S, Danenberg KD, Danenberg PV, Stohr R, Stockle M, Lehmann J, Schuler M and Hartmann A. MDR1 and ERCC1 expression predict outcome of patients with locally advanced bladder cancer receiving adjuvant chemotherapy. Neoplasia. 2010; 12:628-636.
26. Rubatt JM, Darcy KM, Tian C, Muggia F, Dhir R, Armstrong DK, Bookman MA, Niedernhofer LJ, Deloia J, Birrer M and Krivak TC. Pre-treatment tumor expression of ERCC1 in women with advanced stage epithelial ovarian cancer is not predictive of clinical outcomes: a Gynecologic Oncology Group study. Gynecol Oncol. 2012; 125:421-426.
27. Simon G, Sharma A, Li X, Hazelton T, Walsh F, Williams C, Chiappori A, Haura E, Tanvetyanon T, Antonia S, Cantor A and Bepler G. Feasibility and efficacy of molecular analysis-directed individualized therapy in advanced non-small-cell lung cancer. J Clin Oncol. 2007; 25:2741-2746.
28. Smit EF, Socinski MA, Mullaney BP, Myrand SP, Scagliotti GV, Lorigan P, Reck M, Ciuleanu T, von Pawel J, Karaseva NA, Szczesna A, Ohannesian D, Powell E, et al. Biomarker analysis in a phase III study of pemetrexed-carboplatin versus etoposide-carboplatin in chemonaive patients with extensive-stage small-cell lung cancer. Ann Oncol. 2012; 23:1723-1729.
29. Hsu DS, Lan HY, Huang CH, Tai SK, Chang SY, Tsai TL, Chang CC, Tzeng CH, Wu KJ, Kao JY and Yang MH. Regulation of excision repair cross-complementation group 1 by Snail contributes to cisplatin resistance in head and neck cancer. Clin Cancer Res. 2010; 16:4561-4571.
30. Li W, Sun Q and Lu M. [Correlation between expression of ERCC1 and the treatment of cisplatin-based chemotherapy in local advanced nasopharyngeal carcinoma]. Lin Chung Er Bi Yan Hou Tou Jing Wai Ke Za Zhi. 2015; 29:144-146.
31. Mancuso A, Sacchetta S, Saletti PC, Tronconi C, Milesi L, Garassino M, Martelli O, Leone A, Zivi A, Cerbone L, Recine F, Sollami R, Labianca R, et al. Clinical and molecular determinants of survival in pancreatic cancer patients treated with second-line chemotherapy: results of an Italian/Swiss multicenter survey. Anticancer Res. 2010; 30:4289-4295.
32. Lin K, Ye D and Xie X. Protein expression levels of excision repair cross-complementation group 1 and xeroderma pigmentosum D correlate with response to platinum-based chemotherapy in the patients with advanced epithelial ovarian cancer. Int J Gynecol Cancer. 2008; 18:1007-1012.
33. Lee HW, Hwang YH, Han JH, Choi JH, Kang SY, Jeong SH, Ann MS, Oh YT, Kim JH, Kim CH and Sheen SS. High expression of excision repair cross-complementation group 1 protein predicts poor outcome in patients with nasopharyngeal cancer. Oral Oncol. 2010; 46:209-213.
34. Friboulet L, Olaussen KA, Pignon JP, Shepherd FA, Tsao MS, Graziano S, Kratzke R, Douillard JY, Seymour L, Pirker R, Filipits M, Andre F, Solary E, et al. ERCC1 isoform expression and DNA repair in non-small-cell lung cancer. N Engl J Med. 2013; 368:1101-1110.
35. Kitange GJ, Carlson BL, Schroeder MA, Grogan PT, Lamont JD, Decker PA, Wu W, James CD and Sarkaria JN. Induction of MGMT expression is associated with temozolomide resistance in glioblastoma xenografts. Neuro Oncol. 2009; 11:281-291.
36. Happold C, Roth P, Wick W, Schmidt N, Florea AM, Silginer M, Reifenberger G and Weller M. Distinct molecular mechanisms of acquired resistance to temozolomide in glioblastoma cells. J Neurochem. 2012; 122:444-455.
37. Kulke MH, Hornick JL, Frauenhoffer C, Hooshmand S, Ryan DP, Enzinger PC, Meyerhardt JA, Clark JW, Stuart K, Fuchs CS and Redston MS. O6-methylguanine DNA methyltransferase deficiency and response to temozolomide-based therapy in patients with neuroendocrine tumors. Clin Cancer Res. 2009; 15:338-345.
38. Augustine CK, Yoo JS, Potti A, Yoshimoto Y, Zipfel PA, Friedman HS, Nevins JR, Ali-Osman F and Tyler DS. Genomic and molecular profiling predicts response to temozolomide in melanoma. Clin Cancer Res. 2009; 15:502-510.
39. Preusser M, Charles Janzer R, Felsberg J, Reifenberger G, Hamou MF, Diserens AC, Stupp R, Gorlia T, Marosi C, Heinzl H, Hainfellner JA and Hegi M. Anti-O6-methylguanine-methyltransferase (MGMT) immunohistochemistry in glioblastoma multiforme: observer variability and lack of association with patient survival impede its use as clinical biomarker. Brain Pathol. 2008; 18:520-532.
40. Querfeld C, Rosen ST, Guitart J, Rademaker A, Pezen DS, Dolan ME, Baron J, Yarosh DB, Foss F and Kuzel TM. Multicenter phase II trial of temozolomide in mycosis fungoides/sezary syndrome: correlation with O(6)-methylguanine-DNA methyltransferase and mismatch repair proteins. Clin Cancer Res. 2011; 17:5748-5754.
41. Pietanza MC, Kadota K, Huberman K, Sima CS, Fiore JJ, Sumner DK, Travis WD, Heguy A, Ginsberg MS, Holodny AI, Chan TA, Rizvi NA, Azzoli CG, et al. Phase II trial of temozolomide in patients with relapsed sensitive or refractory small cell lung cancer, with assessment of methylguanine-DNA methyltransferase as a potential biomarker. Clin Cancer Res. 2012; 18:1138-1145.
42. Rietschel P, Wolchok JD, Krown S, Gerst S, Jungbluth AA, Busam K, Smith K, Orlow I, Panageas K and Chapman PB. Phase II study of extended-dose temozolomide in patients with melanoma. J Clin Oncol. 2008; 26:2299-2304.
43. Hegi ME, Diserens AC, Gorlia T, Hamou MF, de Tribolet N, Weller M, Kros JM, Hainfellner JA, Mason W, Mariani L, Bromberg JE, Hau P, Mirimanoff RO, et al. MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med. 2005; 352:997-1003.
44. McCormack AI, Wass JA and Grossman AB. Aggressive pituitary tumours: the role of temozolomide and the assessment of MGMT status. Eur J Clin Invest. 2011; 41:1133-1148.
45. Bergman AM, Eijk PP, Ruiz van Haperen VW, Smid K, Veerman G, Hubeek I, van den Ijssel P, Ylstra B and Peters GJ. In vivo induction of resistance to gemcitabine results in increased expression of ribonucleotide reductase subunit M1 as the major determinant. Cancer Res. 2005; 65:9510-9516.
46. Oguri T, Achiwa H, Sato S, Bessho Y, Takano Y, Miyazaki M, Muramatsu H, Maeda H, Niimi T and Ueda R. The determinants of sensitivity and acquired resistance to gemcitabine differ in non-small cell lung cancer: a role of ABCC5 in gemcitabine sensitivity. Mol Cancer Ther. 2006; 5:1800-1806.
47. Wonganan P, Chung WG, Zhu S, Kiguchi K, Digiovanni J and Cui Z. Silencing of ribonucleotide reductase subunit M1 potentiates the antitumor activity of gemcitabine in resistant cancer cells. Cancer Biol Ther. 2012; 13:908-914.
48. Valsecchi ME, Holdbrook T, Leiby BE, Pequignot E, Littman SJ, Yeo CJ, Brody JR and Witkiewicz AK. Is there a role for the quantification of RRM1 and ERCC1 expression in pancreatic ductal adenocarcinoma? BMC Cancer. 2012; 12:104.
49. Ohhashi S, Ohuchida K, Mizumoto K, Fujita H, Egami T, Yu J, Toma H, Sadatomi S, Nagai E and Tanaka M. Down-regulation of deoxycytidine kinase enhances acquired resistance to gemcitabine in pancreatic cancer. Anticancer Res. 2008; 28:2205-2212.
50. Nakano Y, Tanno S, Koizumi K, Nishikawa T, Nakamura K, Minoguchi M, Izawa T, Mizukami Y, Okumura T and Kohgo Y. Gemcitabine chemoresistance and molecular markers associated with gemcitabine transport and metabolism in human pancreatic cancer cells. Br J Cancer. 2007; 96:457-463.
51. Nakahira S, Nakamori S, Tsujie M, Takahashi Y, Okami J, Yoshioka S, Yamasaki M, Marubashi S, Takemasa I, Miyamoto A, Takeda Y, Nagano H, Dono K, et al. Involvement of ribonucleotide reductase M1 subunit overexpression in gemcitabine resistance of human pancreatic cancer. Int J Cancer. 2007; 120:1355-1363.
52. Mori R, Ishikawa T, Ichikawa Y, Taniguchi K, Matsuyama R, Ueda M, Fujii Y, Endo I, Togo S, Danenberg PV and Shimada H. Human equilibrative nucleoside transporter 1 is associated with the chemosensitivity of gemcitabine in human pancreatic adenocarcinoma and biliary tract carcinoma cells. Oncol Rep. 2007; 17:1201-1205.
53. Jordheim LP, Seve P, Tredan O and Dumontet C. The ribonucleotide reductase large subunit (RRM1) as a predictive factor in patients with cancer. Lancet Oncol. 2011; 12:693-702.
54. Ashida R, Nakata B, Shigekawa M, Mizuno N, Sawaki A, Hirakawa K, Arakawa T and Yamao K. Gemcitabine sensitivity-related mRNA expression in endoscopic ultrasound-guided fine-needle aspiration biopsy of unresectable pancreatic cancer. J Exp Clin Cancer Res. 2009; 28:83.
55. Akita H, Zheng Z, Takeda Y, Kim C, Kittaka N, Kobayashi S, Marubashi S, Takemasa I, Nagano H, Dono K, Nakamori S, Monden M, Mori M, et al. Significance of RRM1 and ERCC1 expression in resectable pancreatic adenocarcinoma. Oncogene. 2009; 28:2903-2909.
56. Simon GR, Schell MJ, Begum M, Kim J, Chiappori A, Haura E, Antonia S and Bepler G. Preliminary indication of survival benefit from ERCC1 and RRM1-tailored chemotherapy in patients with advanced nonsmall cell lung cancer: evidence from an individual patient analysis. Cancer. 2012; 118:2525-2531.
57. Lv C, Ma Y, Feng Q, Fang F, Bai H, Zhao B, Yan S, Wu N, Zheng Q, Li S, Chen J, Wang J, Feng Y, et al. A pilot study: sequential gemcitabine/cisplatin and icotinib as induction therapy for stage IIB to IIIA non-small-cell lung adenocarcinoma. World J Surg Oncol. 2013; 11:96.
58. Ijichi K, Adachi M, Ogawa T, Hasegawa Y and Murakami S. Cell-cycle distribution and Thymidilate Synthatase (TS) expression correlate with 5-FU resistance in head and neck carcinoma cells. Anticancer Res. 2014; 34:2907-2911.
59. Kurata N, Fujita H, Ohuchida K, Mizumoto K, Mahawithitwong P, Sakai H, Onimaru M, Manabe T, Ohtsuka T and Tanaka M. Predicting the chemosensitivity of pancreatic cancer cells by quantifying the expression levels of genes associated with the metabolism of gemcitabine and 5-fluorouracil. Int J Oncol. 2011; 39:473-482.
60. Ligabue A, Marverti G, Liebl U and Myllykallio H. Transcriptional activation and cell cycle block are the keys for 5-fluorouracil induced up-regulation of human thymidylate synthase expression. PLoS One. 2012; 7:e47318.
61. Ide H, Kikuchi E, Hasegawa M, Kozakai N, Kosaka T, Miyajima A and Oya M. Prognostic significance of 5-fluorouracil metabolism-relating enzymes and enhanced chemosensitivity to 5-fluorouracil by 5-chloro 2,4-dihydroxy-pyridine in urothelial carcinoma. BMC Cancer. 2012; 12:420.
62. Brody JR, Hucl T, Gallmeier E, Winter JM, Kern SE and Murphy KM. Genomic copy number changes affecting the thymidylate synthase (TYMS) gene in cancer: a model for patient classification to aid fluoropyrimidine therapy. Cancer Res. 2006; 66:9369-9373.
63. Gu W, Fang FF, Li B, Cheng BB and Ling CQ. Characterization and resistance mechanisms of a 5-fluorouracil- resistant hepatocellular carcinoma cell line. Asian Pac J Cancer Prev. 2012; 13:4807-4814.
64. Meropol NJ, Gold PJ, Diasio RB, Andria M, Dhami M, Godfrey T, Kovatich AJ, Lund KA, Mitchell E and Schwarting R. Thymidine phosphorylase expression is associated with response to capecitabine plus irinotecan in patients with metastatic colorectal cancer. J Clin Oncol. 2006; 24:4069-4077.
65. Lu M, Gao J, Wang XC and Shen L. Expressions of Thymidylate Synthase, Thymidine Phosphorylase, Class III beta-tubulin, and Excision Repair Cross-complementing Group 1predict Response in Advanced Gastric Cancer Patients Receiving Capecitabine Plus Paclitaxel or Cisplatin. Chin J Cancer Res. 2011; 23:288-294.
66. Liu XF, Zhang H, Sun JQ, Yin C, Liu TF, Yang H and Chen LH. Correlation between expression of thymidylate synthase and clinical outcome of advanced gastric cancer treated with capecitabine alone chemotherapy. Tumour Biol. 2014; 35:12409-12414.
67. Lindebjerg J, Nielsen JN, Hoeffding LD and Jakobsen A. Immunohistochemical expression of thymidylate synthase as predictor of response to capecitabine in patients with advanced colorectal adenocarcinoma. APMIS. 2005; 113:600-602.
68. Lee J, Im YH, Cho EY, Hong YS, Lee HR, Kim HS, Kim MJ, Kim K, Kang WK, Park K and Shim YM. A phase II study of capecitabine and cisplatin (XP) as first-line chemotherapy in patients with advanced esophageal squamous cell carcinoma. Cancer Chemother Pharmacol. 2008; 62:77-84.
69. Kornmann M, Hebart H, Danenberg K, Goeb R, Staib L, Kron M, Henne-Bruns D, Danenberg P and Link KH. Response prediction in metastasised colorectal cancer using intratumoural thymidylate synthase: results of a randomised multicentre trial. Eur J Cancer. 2012; 48:1443-1451.
70. Showalter SL, Showalter TN, Witkiewicz A, Havens R, Kennedy EP, Hucl T, Kern SE, Yeo CJ and Brody JR. Evaluating the drug-target relationship between thymidylate synthase expression and tumor response to 5-fluorouracil. Is it time to move forward? Cancer Biol Ther. 2008; 7:986-994.
71. Kavallaris M, Kuo DY, Burkhart CA, Regl DL, Norris MD, Haber M and Horwitz SB. Taxol-resistant epithelial ovarian tumors are associated with altered expression of specific beta-tubulin isotypes. J Clin Invest. 1997; 100:1282-1293.
72. Derry WB, Wilson L, Khan IA, Luduena RF and Jordan MA. Taxol differentially modulates the dynamics of microtubules assembled from unfractionated and purified beta-tubulin isotypes. Biochemistry. 1997; 36:3554-3562.
73. Mozzetti S, Ferlini C, Concolino P, Filippetti F, Raspaglio G, Prislei S, Gallo D, Martinelli E, Ranelletti FO, Ferrandina G and Scambia G. Class III beta-tubulin overexpression is a prominent mechanism of paclitaxel resistance in ovarian cancer patients. Clin Cancer Res. 2005; 11:298-305.
74. Reiman T, Lai R, Veillard AS, Paris E, Soria JC, Rosell R, Taron M, Graziano S, Kratzke R, Seymour L, Shepherd FA, Pignon JP, Seve P and LACE-Bio Group. Cross-validation study of class III beta-tubulin as a predictive marker for benefit from adjuvant chemotherapy in resected non-small-cell lung cancer: analysis of four randomized trials. Ann Oncol. 2012; 23:86-93.
75. Kaira K, Takahashi T, Murakami H, Shukuya T, Kenmotsu H, Ono A, Naito T, Tsuya A, Nakamura Y, Endo M, Kondo H, Nakajima T and Yamamoto N. The role of betaIII-tubulin in non-small cell lung cancer patients treated by taxane-based chemotherapy. Int J Clin Oncol. 2013; 18:371-379.
76. Galmarini CM, Treilleux I, Cardoso F, Bernard-Marty C, Durbecq V, Gancberg D, Bissery MC, Paesmans M, Larsimont D, Piccart MJ, Di Leo A and Dumontet C. Class III beta-tubulin isotype predicts response in advanced breast cancer patients randomly treated either with single-agent doxorubicin or docetaxel. Clin Cancer Res. 2008; 14:4511-4516.
77. Dumontet C, Isaac S, Souquet PJ, Bejui-Thivolet F, Pacheco Y, Peloux N, Frankfurter A, Luduena R and Perol M. Expression of class III beta tubulin in non-small cell lung cancer is correlated with resistance to taxane chemotherapy. Bull Cancer. 2005; 92:E25-30.
78. Bernard-Marty C, Treilleux I, Dumontet C, Cardoso F, Fellous A, Gancberg D, Bissery MC, Paesmans M, Larsimont D, Piccart MJ and Di Leo A. Microtubule-associated parameters as predictive markers of docetaxel activity in advanced breast cancer patients: results of a pilot study. Clin Breast Cancer. 2002; 3:341-345.
79. Azuma K, Sasada T, Kawahara A, Takamori S, Hattori S, Ikeda J, Itoh K, Yamada A, Kage M, Kuwano M and Aizawa H. Expression of ERCC1 and class III beta-tubulin in non-small cell lung cancer patients treated with carboplatin and paclitaxel. Lung Cancer. 2009; 64:326-333.
80. Azuma K, Sasada T, Kawahara A, Hattori S, Kinoshita T, Takamori S, Ichiki M, Imamura Y, Ikeda J, Kage M, Kuwano M and Aizawa H. Expression of ERCC1 and class III beta-tubulin in non-small cell lung cancer patients treated with a combination of cisplatin/docetaxel and concurrent thoracic irradiation. Cancer Chemother Pharmacol. 2009; 64:565-573.
81. Hwang JE, Hong JY, Kim K, Kim SH, Choi WY, Kim MJ, Jung SH, Shim HJ, Bae WK, Hwang EC, Lee KH, Lee JH, Cho SH and Chung IJ. Class III beta-tubulin is a predictive marker for taxane-based chemotherapy in recurrent and metastatic gastric cancer. BMC Cancer. 2013; 13:431.
82. Pentheroudakis G, Batistatou A, Kalogeras KT, Kronenwett R, Wirtz RM, Bournakis E, Eleftheraki AG, Pectasides D, Bobos M, Papaspirou I, Kamina S, Gogas H, Koutras AK, et al. Prognostic utility of beta-tubulin isotype III and correlations with other molecular and clinicopathological variables in patients with early breast cancer: a translational Hellenic Cooperative Oncology Group (HeCOG) study. Breast Cancer Res Treat. 2011; 127:179-193.
83. Seve P and Dumontet C. Is class III beta-tubulin a predictive factor in patients receiving tubulin-binding agents? Lancet Oncol. 2008; 9:168-175.
84. Cubillo A, Hernando-Requejo O, Garcia-Garcia E, Rodriguez-Pascual J, De Vicente E, Morelli P, Rubio C, Lopez-Rios F, Muro A, Lopez U, Prados S, Quijano Y and Hidalgo M. A prospective pilot study of target-guided personalized chemotherapy with intensity-modulated radiotherapy in patients with early rectal cancer. Am J Clin Oncol. 2014; 37:117-121.
85. Shen J, Wei J, Wang H, Yue G, Yu L, Yang Y, Xie L, Zou Z, Qian X, Ding Y, Guan W and Liu B. A three-gene signature as potential predictive biomarker for irinotecan sensitivity in gastric cancer. J Transl Med. 2013; 11:73.
86. Zhang W, Shannon WD, Duncan J, Scheffer GL, Scheper RJ and McLeod HL. Expression of drug pathway proteins is independent of tumour type. J Pathol. 2006; 209:213-219.
87. Sakai A, Kasahara K, Ohmori T, Kimura H, Sone T, Fujimura M and Nakao S. MET increases the sensitivity of gefitinib-resistant cells to SN-38, an active metabolite of irinotecan, by up-regulating the topoisomerase I activity. J Thorac Oncol. 2012; 7:1337-1344.
88. Horisberger K, Erben P, Muessle B, Woernle C, Stroebel P, Kaehler G, Wenz F, Hochhaus A, Post S, Willeke F and Hofheinz RD. Topoisomerase I expression correlates to response to neoadjuvant irinotecan-based chemoradiation in rectal cancer. Anticancer Drugs. 2009; 20:519-524.
89. Gilbert DC, Chalmers AJ and El-Khamisy SF. Topoisomerase I inhibition in colorectal cancer: biomarkers and therapeutic targets. Br J Cancer. 2012; 106:18-24.
90. Nieves-Neira W and Pommier Y. Apoptotic response to camptothecin and 7-hydroxystaurosporine (UCN-01) in the 8 human breast cancer cell lines of the NCI Anticancer Drug Screen: multifactorial relationships with topoisomerase I, protein kinase C, Bcl-2, p53, MDM-2 and caspase pathways. Int J Cancer. 1999; 82:396-404.
91. Goldwasser F, Bae I, Valenti M, Torres K and Pommier Y. Topoisomerase I-related parameters and camptothecin activity in the colon carcinoma cell lines from the National Cancer Institute anticancer screen. Cancer Res. 1995; 55:2116-2121.
92. Braun MS, Richman SD, Quirke P, Daly C, Adlard JW, Elliott F, Barrett JH, Selby P, Meade AM, Stephens RJ, Parmar MK and Seymour MT. Predictive biomarkers of chemotherapy efficacy in colorectal cancer: results from the UK MRC FOCUS trial. J Clin Oncol. 2008; 26:2690-2698.
93. Koopman M, Knijn N, Richman S, Seymour M, Quirke P, van Tinteren H, van Krieken JHJM, Punt CJA and Nagtegaal ID. The correlation between topoisomerase-I (Topo1) expression and outcome of treatment with capecitabine and irinotecan in advanced colorectal cancer (ACC) patients (pts) treated in the CAIRO study of the Dutch Colorectal Cancer Group (DCCG). Eur J of Cancer Suppl. 2009; 7:321.
94. Meisenberg C, Gilbert DC, Chalmers A, Haley V, Gollins S, Ward SE and El-Khamisy SF. Clinical and cellular roles for TDP1 and TOP1 in modulating colorectal cancer response to irinotecan. Mol Cancer Ther. 2015; 14:575-585.
95. AbuHammad S and Zihlif M. Gene expression alterations in doxorubicin resistant MCF7 breast cancer cell line. Genomics. 2013; 101:213-220.
96. Burgess DJ, Doles J, Zender L, Xue W, Ma B, McCombie WR, Hannon GJ, Lowe SW and Hemann MT. Topoisomerase levels determine chemotherapy response in vitro and in vivo. Proc Natl Acad Sci U S A. 2008; 105:9053-9058.
97. Wong N, Yeo W, Wong WL, Wong NL, Chan KY, Mo FK, Koh J, Chan SL, Chan AT, Lai PB, Ching AK, Tong JH, Ng HK, et al. TOP2A overexpression in hepatocellular carcinoma correlates with early age onset, shorter patients survival and chemoresistance. Int J Cancer. 2009; 124:644-652.

