Introduction
Gastric cancer (GCa) is the second commonly diagnosed cancer and also the second leading cause of cancer-related deaths in China [1]. According to recent cancer statistics, there were approximately 679,100 new cancer cases and 498,000 new deaths in 2015 in China, accounting for 15.8% of all cancer cases and 17.7% of all cancer deaths, respectively. Before diagnosis, GCa patients in the early stage did not have obvious symptoms usually, but the disease rapidly progressed, if its diagnosis was missed. Therefore, only prevention and earlier detection/diagnosis of GCa will reduce the incidence and mortality of GCa in China. Currently, the major treatment for GCa is still surgery, if the tumors are resectable at the time of diagnosis [2]. However, the high recurrence rate makes it the necessary to identify additional markers for prognosis, which may facilitate individualized treatment and management. Some clinical factors, such as Lauren’s classification, have been shown to be associated with GCa prognosis. For example, patients with intestinal type tumors may have a better prognosis than those with diffuse type tumors [3, 4]; however, there is lack of promising genetic predictors for GCa prognosis [5].
As a result of rapid advance in genomic technologies, genetic variants have been demonstrated by previous studies as potential factors that may affect both cancer etiology [6] and patients’ response to therapies [7] . For example, single nucleotide polymorphisms (SNPs), presented more than 1% in the general population, have been reported to be associated with GCa risk [8-10]. These findings provide tools for early detection, diagnosis and prevention at the genetic level in the at-risk populations. To date, five large published genome-wide association studies (GWASs) have revealed a series of genetic variants associated with susceptibility to GCa [11-15]. Some of these results have then been successfully reproduced by subsequent large case-control studies with sufficient statistical power [16, 17].
Recently, summary data on genetic susceptibility to GCa were pooled and comprehensively reviewed by a large meta-analysis [18]. Interestingly, it was reported that these GWAS-derived SNPs associated with GCa susceptibility are located in genes that are involved in carcinogenesis and tumor progression, which may affect the invasiveness and metastatic potential of GCa [19-21]. Moreover, the biological role of susceptibility genes in tumor progression may be reflected in the correlation of some susceptibility loci with poor prognosis [10, 22-25]. However, it has been reported that the somatic status of tumor cells is quit heterogeneous among patients, indicating that germline variants may also play a role in determining which somatic alterations is likely to be acquired [26], in addition to exposure to carcinogens. Therefore, tumors of patients with different inherited genetic background may act in different biological ways.
Taken together, there is one possibility that tumors in patients with different genetic risk of developing cancer derived from germline variants may have a different pattern of disease course. Therefore, we hypothesize that a well-defined genetic predisposition of patients with GCa is associated with patients’ prognosis. To test this hypothesis, we used the available genotyping data from an ongoing case-control study of GCa and survival data from the same patients in a single institution.
Results
Characteristics of the subjects are presented in Supplementary Tables 1 and 2. The 42 selected SNPs and their relevant information are presented in Supplementary Table 3. In the initial tests, the weighted genetic risk score (GRS) showed a normal distribution in all the subjects and subgroups as well (Figure 1). In the subsequent analyses, the results of multivariate fractional polynomial Cox proportional hazards regression models showed that the increase of GRS did not have an effect on the risk of death in the patients, although an upward trend was observed in the subgroup of stage IV and patients with Lauren’s mix-type tumors (Figure 2). According to the segmented trend of fractional polynomial regression model, a low, medium and high GRS was defined as the < 25%, 25%-75% and >75% quartiles, respectively. However, compared with patients with the medium GRS, those with a low and high GRS did have a similar survival [low vs medium vs high corresponding 74% vs 61.71% vs 59.09% of 5-year overall survival (OS); log-rank test, P = 0.879, Figure 3], and the results remained unchanged after adjustment for age, sex, stage, Lauren’s classification and differentiate degree of tumors, and treatment (low vs medium, HR = 0.87 and P = 0.349, and high vs medium, HR = 0.97 and P = 0.847). These results did not change substantially in the subgroups of either stage or Laruen’s classification (Table 1). Moreover, GRS was not helpful, in addition to clinical factors, to discriminate more patients with different outcomes in either the overall patients [Concordance index (C-index): 0.78 vs 0.78, P = 0.986] or subgroups of stage and Lauren’s classification (Table 2).
Table 1: Association of GRS with survival of GCa patients in an Eastern Chinese population |
||||
Group |
GRS |
Crude HR (95% CI), P value |
Adjusted HR (95% CI)a, P value |
Log-rank testb for 5-year OS, P value |
Overall |
Low |
1.01 (0.76, 1.36), 0.924 |
0.87 (0.64, 1.17), 0.349 |
60.74% vs 61.71% vs 59.09%, 0.879 |
Medium |
Reference |
Reference |
||
High |
1.08 (0.81, 1.45), 0.603 |
0.97 (0.72, 1.31), 0.847 |
||
Stage I |
Low |
0.66 (0.07, 5.94), 0.714 |
0.62 (0.06, 5.91), 0.677 |
97.44% vs 97.14% vs 95.00%, 0.546 |
Medium |
Reference |
Reference |
||
High |
1.89 (0.42, 8.46), 0.404 |
1.97 (0.40, 9.63), 0.402 |
||
Stage II |
Low |
0.86 (0.45, 1.64), 0.653 |
0.76 (0.49, 1.48), 0.425 |
76.00% vs 68.00% vs 75.00%, 0.804 |
Medium |
Reference |
Reference |
||
High |
0.79 (0.39, 1.60), 0.506 |
0.73 (0.35, 1.53), 0.401 |
||
Stage III |
Low |
0.91 (0.63, 1.32), 0.629 |
0.85 (0.59, 1.24), 0.413 |
34.85% vs 34.45% vs 34.92%, 0.880 |
Medium |
Reference |
Reference |
||
High |
0.94 (0.65, 1.36), 0.750 |
0.90 (0.60, 1.27), 0.467 |
||
Stage IV |
Low |
1.09 (0.45, 2.63), 0.848 |
0.91 (0.34, 2.41), 0.847 |
0% vs 5.88% vs 9.09%, 0.584 |
Medium |
Reference |
Reference |
||
High |
1.39 (0.62, 3.09), 0.420 |
1.99 (0.74, 5.35), 0.171 |
||
Intestinal |
Low |
0.96 (0.58, 1.59), 0.878 |
0.66 (0.39, 1.12), 0.128 |
67.19% vs 67.86% vs 62.71%, 0.425 |
Medium |
Reference |
Reference |
||
High |
1.33 (0.83, 2.14), 0.239 |
1.15 (0.70, 1.88), 0.584 |
||
Mix |
Low |
1.13 (0.66, 1.93), 0.659 |
1.06 (0.61, 1.86), 0.831 |
58.14% vs 58.33% vs 61.82%, 0.799 |
Medium |
Reference |
Reference |
||
High |
0.92 (0.55, 1.54), 0.743 |
0.93 (0.54, 1.60), 0.802 |
||
Diffuse |
Low |
1.06 (0.63, 1.77), 0.820 |
0.94 (0.54, 1.61), 0.812 |
52.00% vs 56.58% vs 48.65%, 0.930 |
Medium |
Reference |
Reference |
||
High |
1.10 (0.63, 1.92), 0.749 |
0.97 (0.53, 1.78), 0.919 |
||
GRS, genetic risk score; GCa, gastric cancer; HR, hazards ratio; OS, overall survival; aAdjusted for gender, age, disease stage, Lauren classification, tumor differentiation and treatment; bLog-rank test of 5-year OS, based on the comparison of Low versus medium versus high GRS. |
||||
Table 2: Dynamic predictive value (C-index) of considering clinical factors and GRS versus that of considering clinical factors alone in an Eastern Chinese population |
||
Group |
C-index |
P valueb |
Overall |
0.78 vs 0.78 |
0.986 |
Stage I |
0.79 vs 0.77 |
0.880 |
Stage II |
0.60 vs 0.60 |
0.999 |
Stage III |
0.56 vs 0.56 |
0.999 |
Stage IV |
0.68 vs 0.62 |
0.417 |
Intestinal |
0.82 vs 0.81 |
0.812 |
Mix |
0.75 vs 0.75 |
0.999 |
Diffuse |
0.78 vs 0.78 |
0.991 |
GRS, genetic risk score; C-index, concordance index; aClinical factors include age, gender, disease stage, Lauren classification, tumor differentiation and treatment; bP value for bootstrapping test (number of boots = 1000) |
||
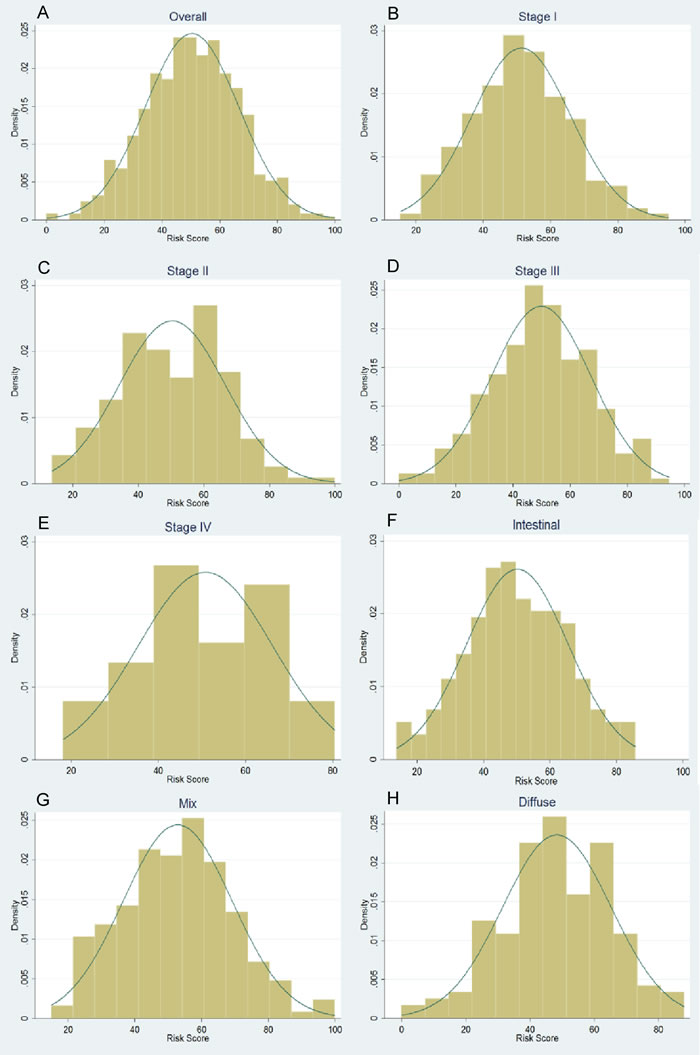
Figure 1: Normal distribution of genetic risk score (GRS) for overall patients (A), subgroup of stages (B-E) and Lauren’s classification (F-H)
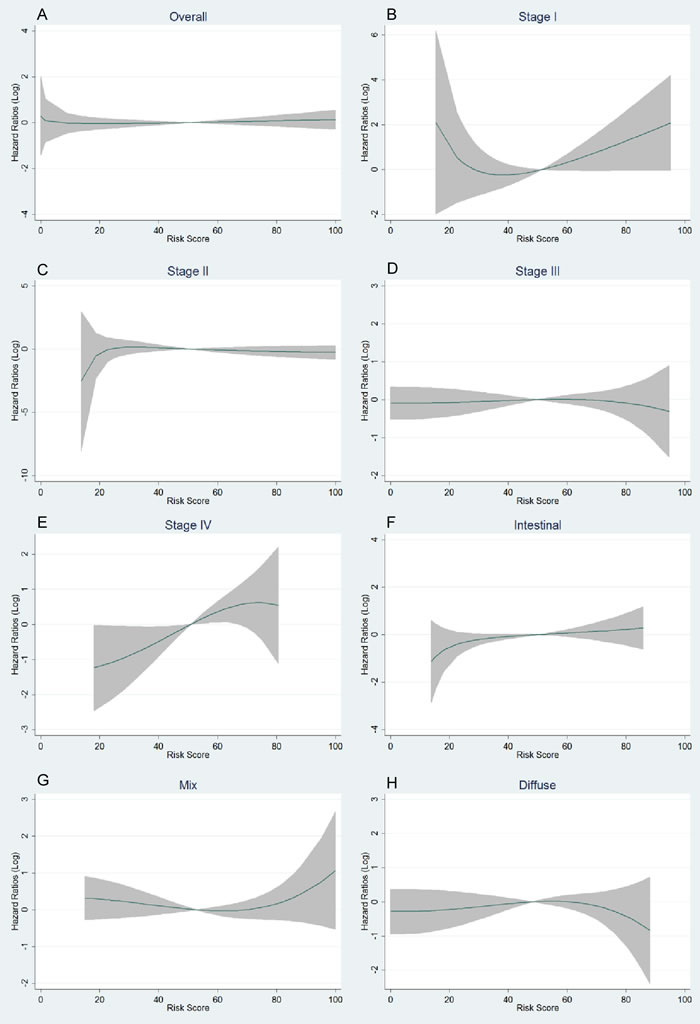
Figure 2: Trends of logarithmic HRs for death with the increase of genetic risk score (GRS) in multivariate fractional polynomial Cox proportional hazards regression models. Overall patients A.; subgroup of stages B.-E.; subgroup of Lauren’s classification F.-H. The continuous line is the point estimation of logarithmic HRs and the shadow area is the 95% confidence interval of logarithmic HRs.
Discussion
Recently, it has been shown that there is no association between GRS of prostate cancer and risk of radiotherapy toxicity, although some susceptibility genes were indeed involved in cellular radiation response [27]. In the present study, we tested the hypothesis that GRS, a genetic estimation of GCa risk, is also associated with survival in an Eastern Chinese population. We first used a case-control study to develop a population-specific GRS based on 42 well-established GCa susceptibility loci derived from published GWASs and meta-analysis and then used this GRS in a survival analysis of the patients from the same case-control study. Consistently with the reported prostate cancer study, we did not find any association between genetic risk of GCa and prognosis of the patients in this study population, nor did we find any evidence for an interaction between GRS and stage or Laruen’s classification, two clinical factors closely associated with prognosis of GCa. These data suggest that individuals who are susceptible to GCa are not necessarily have a poor clinical outcomes, which indicates that additional genetic variants that are responsible for poor clinical outcomes of GCa patients need to be further identified.
A number of studies indicated that a constitutional or germline background in cells, which is a primary genetic background inherited from parents, may indicate which somatic pattern in the tumor is more likely to be subsequently acquired. This phenomenon was further demonstrated in several cancer types. For example, it was reported that melanoma patients with germline variants in MC1R tended to acquire BRCA1 mutations more likely than those without genetic variants in MC1R [28]. Moreover, breast cancer patients with inherited BRCA1 variants have a higher frequency of copy number alterations than those without inherited BRCA1 variants [29]. On the other hand, studies have found that SNPs associated with GCa risk may regulate the expression of their nearby genes. These genes, such as PRKAA1 [14], PLCE1 [12] and MUC1 [30], also play an important role in cancer progression and tumorigenesis. For example, the PRKAA1 gene encodes catalytic α-subunit of 5′ AMP-activated protein kinase, which is involved in protecting tumors from energy deprivation [31] and thus promoting cancer metastasis [32, 33]. As a susceptible gene for GCa and esophageal squamous cell carcinoma, the effect of PLCE1 in tumorigenesis was also elucidated by a recent study, in which the PLCE1 gene targeted by miR-145 impaired tumor metastasis and proliferation [19]. Besides, several studies reported a biological role of MUC1 in cancer metastasis through disturbing cell adhesions [34-36]. Interestingly, the use of anti-MUC1 antibody was helpful for identifying poor differentiated cells in GCa tumors [20]. Tumorigenic effect of other susceptible genes, such as PKLR, on GCa was also reported [21]. Although all of these previous findings suggest a possibility that GCa patients with a different genetic susceptibility may have a different disease course that may contribute to the prognosis variation, we did not find any polygenic association with the prognosis of Chinese patients with GCa. The negative results can be explained as follows: first, not every germline variant plays an important role in determining which of somatic alterations is likely to be acquired, although some genetic variants acting as a key driver for somatic alterations have been identified by previous studies [28, 29]; second, environment factors may play a more important role in predicting prognosis than germline variants, which may have masked the effect of germline variants on prognosis of these GCa patients.
Previous studies have revealed a number of genetic variants associated with GCa prognosis. For example, SNPs of the TYMS gene, such as rs16430 and rs1059394, were found to be associated with GCa prognosis in North-American patients [37]. Genetic variants in genes involved in the nucleotide excision repair (NER) pathway were also shown to have an impact on survival of GCa patients in both Japanese and North-American patients [38]. Interestingly, a recent study found that the PSCA rs2294008 T allele was associated with either a poor prognosis or an increased susceptibility of Spanish GCa patients [10].
In the present study, GRS in the subgroup of stage IV patient showed the notable ability to help clinical factors to distinguish more patients with better and worse survival in a dynamic process (Cindex, 0.68 vs 0.62), although the result did not reach statistical significance (P = 0.417). However, the result from a limited sample size in stage IV patients (n = 36 only) without a sufficient statistical power needs to be validated in additional larger studies. Another interesting finding in the present study was that Lauren’s classification had no effect on the association between genetic risk and disease prognosis. According to Lauren’s classification, tumors can be defined as an intestinal, mix or diffuse type [39], and tumor cells with these three different Lauren’s types may act in different biological ways. Therefore, it is valuable to detect any association of genetic risk of GCa with prognosis in different Lauren’s types. Disease stage is another strong predictor for prognosis. However, it has the characteristic of time dependency, which is changeable in the development of cancer over time. Moreover, patients with an advanced stage may miss the opportunity of treatment due to the poor performance status. Because genetic risk is inherited and thus not changeable in the lifetime, elucidating the association between genetic risk and prognosis for GCa is helpful in making treatment decision at the time of presentation of the disease in different stages.
There are some limitations in the present study. First, due to a decrease in the number of patients in subgroup analysis, statistical power was greatly reduced. Second, although 42 well-established susceptibility loci selected from GWASs and confirmed in large meta-analyses were used, SNPs in some driver genes for tumorigenesis or information about somatic mutations in the same genes were missed in the analyses.
In conclusion, our results do not support our original hypothesis. It is likely that genetic risk of GCa, based at least on those susceptibility loci identified in previous GWASs and high-evidence meta-analysis, may not be associated with the prognosis, as we observed in the patients of an Eastern Chinese population. Due to limitations of the present study using retrospective data, these results needed to be validated by larger perspective studies. Likewise, our findings implicate that genetic risk of GCa may not affect the treatment decision in Chinese GCa patients, and additional genetic variants that may be associated with clinical outcomes in GCa patients need to be further identified.
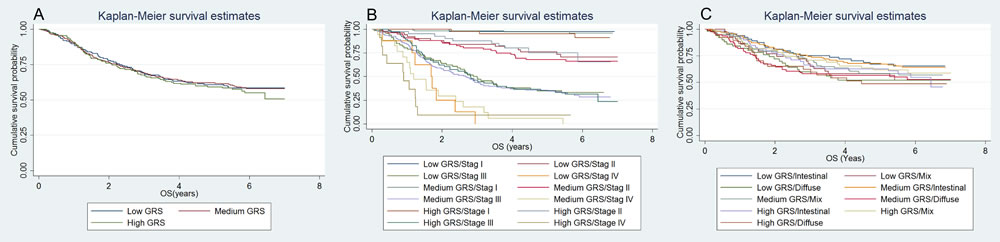
Figure 3: Kaplan-Meier (KM) curve for the survival comparison of patients with different GRS. Overall patients A.; subgroup of stages B.; subgroup of Lauren’s classification C.
Materials and Methods:
SNP selection
In the present study, the established susceptibility loci for GCa were selected by the following criteria: 1) GWAS-derived potential susceptibility loci that were reportedly associated with GCa risk [11-15]; and 2) susceptibility loci for GCa that was confirmed by a recent large meta-analysis as to be high-level evidence [18]. As a result, a total of 42 known SNPs were chosen for genotyping in the study population.
Study subjects
To construct the GRS in a case-control study, we first used 1,115 unrelated ethnic Han Chinese patients with newly diagnosed and histopathologically confirmed primary GCa, who were recruited from Fudan University Shanghai Cancer Center (FUSCC) in Eastern China between January 2009 and March 2011. Patients with neuroendocrine tumors and gastric stromal tumors were excluded. Then, we included an additional 1,172 cancer-free ethnic Han Chinese controls matched the cases by age, sex, smoking and drinking-matched, who were recruited from the Taizhou Longitudinal (TZL) study conducted at the same time period in Eastern China. For the subsequent survival analysis, we used the 633 GCa patients with available clinical data. Disease stage was categorized by the 7th AJCC staging system. A written informed consent for donating biological samples to scientific research was provided by all the subjects whose blood samples were stored at the biobanks at both FUSCC and TZJ studies. The study protocol was approved by the FUSCC institutional review board.
Genotyping and quality control
Blood samples of the GCa cases and controls were provided by the tissue banks at both FUSCC and TZL study. Demographic data of these subjects were collected. Clinical information with survival data was collected during the subsequent follow-up. DNA was extracted from blood samples, and the selected 42 SNPs were genotyped by Sequenom Mass Array in Mass ARRAY Analyzer 4 platform (Sequenom, CA, USA). One negative control and one duplicate sample were used for quality controls in each of the 96 plates. All the primers were designed by Assay Design Suite v2.0 from Mysequenom online software (https://www.mysequenom.com). Genotyping results of 5% patients were repeated, and the consistency was 100%.
Statistical methods
Initially, we used the penalty function in the iterative sure independence screening (ISIS) algorithm, in which a best-fit logistic regression model was generated through regularized log likelihood for the variables picked by ISIS [40]. Then, a weighted GRS was calculated by coefficients in the best-fit logistic regression model as follows:
![]()
Where, i is the i th subject, j is the j th SNP, coefficientj is the coefficient for the j th SNP in the best-fit logistic regression model, and nji is the risk allele dosage for the j th SNP in i th subject [27]. Then the GRS was translated to a range of 0-100, according to its distribution among the study subjects.
Then, fractional polynomial Cox proportional hazards regression model was used to evaluate the trends of logarithmic HRs for death with the increase of GRS. Overall survival was defined as the period between diagnosis and death. Survival data for patients alive or loss to follow-up were considered censored. Survival comparison was made by using the log-rank test and Kaplan-Meier curve. A dynamic predictive model for survival data, with the concordance index (C-index) as the estimation of the information in the predictive model, was adapted to reveal whether GRS for GCa was helpful, in addition to clinical factors, to discriminate more patients with different outcomes. The bootstrapping test was used for comparisons of the C-index.
All statistic work was achieved by R (version 3.2.4, R Foundation, Vienna, Austria) and STATA software (version 12, StataCrop, Texas, USA).
Acknowledgments and funding
This study was supported by the funds from China Recruitment Program of Global Experts at Fudan University, the Shanghai Committee of Science and Technology, China (Grant No. 12DZ2260100), Ministry of Science and Technology (2011BAI09B00) and Ministry of Health (201002007). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
ConflictS of interest
We declare that we have no conflict of interest
References
1. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J. Cancer statistics in China, 2015. CA Cancer J Clin. 2016; 66: 115-32. doi: 10.3322/caac.21338.
2. He W, Tu J, Huo Z, Li Y, Peng J, Qiu Z, Luo D, Ke Z, Chen X. Surgical interventions for gastric cancer: a review of systematic reviews. Int J Clin Exp Med. 2015; 8: 13657-69.
3. Toikkanen S, Helin H, Isola J, Joensuu H. Prognostic significance of HER-2 oncoprotein expression in breast cancer: a 30-year follow-up. J Clin Oncol. 1992; 10: 1044-8.
4. Yamashita K, Sakuramoto S, Katada N, Futawatari N, Moriya H, Hirai K, Kikuchi S, Watanabe M. Diffuse type advanced gastric cancer showing dismal prognosis is characterized by deeper invasion and emerging peritoneal cancer cell: the latest comparative study to intestinal advanced gastric cancer. Hepatogastroenterology. 2009; 56: 276-81.
5. Wadhwa R, Song S, Lee JS, Yao Y, Wei Q, Ajani JA. Gastric cancer-molecular and clinical dimensions. Nat Rev Clin Oncol. 2013; 10: 643-55. doi: 10.1038/nrclinonc.2013.170.
6. Bardai A, Overwater E, Aalfs CM. Germline Mutations in Predisposition Genes in Pediatric Cancer. N Engl J Med. 2016; 374: 1390-1. doi: 10.1056/NEJMc1600338#SA1.
7. Daniels J, Kadlubar S. Sulfotransferase genetic variation: from cancer risk to treatment response. Drug Metab Rev. 2013; 45: 415-22. doi: 10.3109/03602532.2013.835621.
8. Qiu LX, Cheng L, He J, Zhou ZR, Wang MY, Zhou F, Guo WJ, Li J, Sun MH, Zhou XY, Wang YN, Yang YJ, Wang JC, et al. PSCA polymorphisms and gastric cancer susceptibility in an eastern Chinese population. Oncotarget. 2016; 7: 9420-8. doi: 10.18632/oncotarget.7137.
9. Wang MY, He J, Zhu ML, Teng XY, Li QX, Sun MH, Wang XF, Yang YJ, Wang JC, Jin L, Wang YN, Wei QY. A Functional Polymorphism (rs2494752) in the AKT1 Promoter Region and Gastric Adenocarcinoma Risk in an Eastern Chinese Population. Sci Rep. 2016; 6: 20008. doi: 10.1038/srep20008.
10. Garcia-Gonzalez MA, Bujanda L, Quintero E, Santolaria S, Benito R, Strunk M, Sopena F, Thomson C, Perez-Aisa A, Nicolas-Perez D, Hijona E, Carrera-Lasfuentes P, Piazuelo E, et al. Association of PSCA rs2294008 gene variants with poor prognosis and increased susceptibility to gastric cancer and decreased risk of duodenal ulcer disease. Int J Cancer. 2015; 137: 1362-73. doi: 10.1002/ijc.29500.
11. Sakamoto H, Yoshimura K, Saeki N, Katai H, Shimoda T, Matsuno Y, Saito D, Sugimura H, Tanioka F, Kato S, Matsukura N, Matsuda N, Nakamura T, et al. Genetic variation in PSCA is associated with susceptibility to diffuse-type gastric cancer. Nat Genet. 2008; 40: 730-40. doi: 10.1038/ng.152.
12. Abnet CC, Freedman ND, Hu N, Wang Z, Yu K, Shu XO, Yuan JM, Zheng W, Dawsey SM, Dong LM, Lee MP, Ding T, Qiao YL, et al. A shared susceptibility locus in PLCE1 at 10q23 for gastric adenocarcinoma and esophageal squamous cell carcinoma. Nat Genet. 2010; 42: 764-7. doi: 10.1038/ng.649.
13. Wang LD, Zhou FY, Li XM, Sun LD, Song X, Jin Y, Li JM, Kong GQ, Qi H, Cui J, Zhang LQ, Yang JZ, Li JL, et al. Genome-wide association study of esophageal squamous cell carcinoma in Chinese subjects identifies susceptibility loci at PLCE1 and C20orf54. Nat Genet. 2010; 42: 759-63. doi: 10.1038/ng.648.
14. Shi Y, Hu Z, Wu C, Dai J, Li H, Dong J, Wang M, Miao X, Zhou Y, Lu F, Zhang H, Hu L, Jiang Y, et al. A genome-wide association study identifies new susceptibility loci for non-cardia gastric cancer at 3q13.31 and 5p13.1. Nat Genet. 2011; 43: 1215-8. doi: 10.1038/ng.978.
15. Hu N, Wang Z, Song X, Wei L, Kim BS, Freedman ND, Baek J, Burdette L, Chang J, Chung C, Dawsey SM, Ding T, Gao YT, et al. Genome-wide association study of gastric adenocarcinoma in Asia: a comparison of associations between cardia and non-cardia tumours. Gut. 2015. doi: 10.1136/gutjnl-2015-309340.
16. Song HR, Kim HN, Kweon SS, Choi JS, Shim HJ, Cho SH, Chung IJ, Park YK, Kim SH, Choi YD, Joo KW, Shin MH. Genetic variations in the PRKAA1 and ZBTB20 genes and gastric cancer susceptibility in a Korean population. Mol Carcinog. 2013; 52 Suppl 1: E155-60. doi: 10.1002/mc.22063.
17. Zhang H, Jin G, Li H, Ren C, Ding Y, Zhang Q, Deng B, Wang J, Hu Z, Xu Y, Shen H. Genetic variants at 1q22 and 10q23 reproducibly associated with gastric cancer susceptibility in a Chinese population. Carcinogenesis. 2011; 32: 848-52. doi: 10.1093/carcin/bgr051.
18. Mocellin S, Verdi D, Pooley KA, Nitti D. Genetic variation and gastric cancer risk: a field synopsis and meta-analysis. Gut. 2015; 64: 1209-19. doi: 10.1136/gutjnl-2015-309168.
19. Cui XB, Li S, Li TT, Peng H, Jin TT, Zhang SM, Liu CX, Yang L, Shen YY, Li SG, Li N, Li Y, Hu JM, et al. Targeting oncogenic PLCE1 by miR-145 impairs tumor proliferation and metastasis of esophageal squamous cell carcinoma. Oncotarget. 2016; 7: 1777-95. doi: 10.18632/oncotarget.6499.
20. Yonezawa S, Kitajima S, Higashi M, Osako M, Horinouchi M, Yokoyama S, Kitamoto S, Yamada N, Tamura Y, Shimizu T, Tabata M, Goto M. A novel anti-MUC1 antibody against the MUC1 cytoplasmic tail domain: use in sensitive identification of poorly differentiated cells in adenocarcinoma of the stomach. Gastric Cancer. 2012; 15: 370-81. doi: 10.1007/s10120-011-0125-2.
21. Nguyen A, Loo JM, Mital R, Weinberg EM, Man FY, Zeng Z, Paty PB, Saltz L, Janjigian YY, de Stanchina E, Tavazoie SF. PKLR promotes colorectal cancer liver colonization through induction of glutathione synthesis. J Clin Invest. 2016; 126: 681-94. doi: 10.1172/jci83587.
22. Wang YC, Chen CY, Chen SK, Chang YY, Lin P. p53 codon 72 polymorphism in Taiwanese lung cancer patients: association with lung cancer susceptibility and prognosis. Clin Cancer Res. 1999; 5: 129-34. doi:
23. Ding X, Gao S, Yang Q. rs621554 single nucleotide polymorphism of DLC1 is associated with breast cancer susceptibility and prognosis. Mol Med Rep. 2016; 13: 4095-100. doi: 10.3892/mmr.2016.4987.
24. Wang X, Ren H, Zhao T, Ma W, Dong J, Zhang S, Xin W, Yang S, Jia L, Hao J. Single nucleotide polymorphism in the microRNA-199a binding site of HIF1A gene is associated with pancreatic ductal adenocarcinoma risk and worse clinical outcomes. Oncotarget. 2016; 7:13717-29. doi: 10.18632/oncotarget.7263.
25. Zhang W, Zhang ZZ, Tang LY, Lin Y, Su FX, Xie XM, Su XF, Ren ZF. Genetic variants in EBV reactivation-related genes and the risk and survival of breast cancer. Tumour Biol. 2016. doi: 10.1007/s13277-015-4562-0.
26. Dworkin AM, Ridd K, Bautista D, Allain DC, Iwenofu OH, Roy R, Bastian BC, Toland AE. Germline variation controls the architecture of somatic alterations in tumors. PLoS Genet. 2010; 6: e1001136. doi: 10.1371/journal.pgen.1001136.
27. Ahmed M, Dorling L, Kerns S, Fachal L, Elliott R, Partliament M, Rosenstein BS, Vega A, Gomez-Caamano A, Barnett G, Dearnaley DP, Hall E, Sydes M, et al. Common genetic variation associated with increased susceptibility to prostate cancer does not increase risk of radiotherapy toxicity. Br J Cancer. 2016. doi: 10.1038/bjc.2016.94.
28. Landi MT, Bauer J, Pfeiffer RM, Elder DE, Hulley B, Minghetti P, Calista D, Kanetsky PA, Pinkel D, Bastian BC. MC1R germline variants confer risk for BRAF-mutant melanoma. Science. 2006; 313: 521-2. doi: 10.1126/science.1127515.
29. Jonsson G, Naylor TL, Vallon-Christersson J, Staaf J, Huang J, Ward MR, Greshock JD, Luts L, Olsson H, Rahman N, Stratton M, Ringner M, Borg A, et al. Distinct genomic profiles in hereditary breast tumors identified by array-based comparative genomic hybridization. Cancer Res. 2005; 65: 7612-21. doi: 10.1158/0008-5472.can-05-0570.
30. Jia Y, Persson C, Hou L, Zheng Z, Yeager M, Lissowska J, Chanock SJ, Chow WH, Ye W. A comprehensive analysis of common genetic variation in MUC1, MUC5AC, MUC6 genes and risk of stomach cancer. Cancer Causes Control. 2010; 21: 313-21. doi: 10.1007/s10552-009-9463-3.
31. Inoki K, Zhu T, Guan KL. TSC2 mediates cellular energy response to control cell growth and survival. Cell. 2003; 115: 577-90.
32. Legembre P, Schickel R, Barnhart BC, Peter ME. Identification of SNF1/AMP kinase-related kinase as an NF-kappaB-regulated anti-apoptotic kinase involved in CD95-induced motility and invasiveness. J Biol Chem. 2004; 279: 46742-7. doi: 10.1074/jbc.M404334200.
33. Lin H, Li N, He H, Ying Y, Sunkara S, Luo L, Lv N, Huang D, Luo Z. AMPK Inhibits the Stimulatory Effects of TGF-beta on Smad2/3 Activity, Cell Migration, and Epithelial-to-Mesenchymal Transition. Mol Pharmacol. 2015; 88: 1062-71. doi: 10.1124/mol.115.099549.
34. Sluiter N, de Cuba E, Kwakman R, Kazemier G, Meijer G, Te Velde EA. Adhesion molecules in peritoneal dissemination: function, prognostic relevance and therapeutic options. Clin Exp Metastasis. 2016. doi: 10.1007/s10585-016-9791-0.
35. Tamura Y, Higashi M, Kitamoto S, Yokoyama S, Osako M, Horinouchi M, Shimizu T, Tabata M, Batra SK, Goto M, Yonezawa S. MUC4 and MUC1 expression in adenocarcinoma of the stomach correlates with vessel invasion and lymph node metastasis: an immunohistochemical study of early gastric cancer. PLoS One. 2012; 7: e49251. doi: 10.1371/journal.pone.0049251.
36. Retterspitz MF, Monig SP, Schreckenberg S, Schneider PM, Holscher AH, Dienes HP, Baldus SE. Expression of {beta}-catenin, MUC1 and c-met in diffuse-type gastric carcinomas: correlations with tumour progression and prognosis. Anticancer Res. 2010; 30: 4635-41.
37. Shen R, Liu H, Wen J, Liu Z, Wang LE, Wang Q, Tan D, Ajani JA, Wei Q. Genetic polymorphisms in the microRNA binding-sites of the thymidylate synthase gene predict risk and survival in gastric cancer. Mol Carcinog. 2015; 54: 880-8. doi: 10.1002/mc.22160.
38. Li Y, Liu Z, Liu H, Wang LE, Onodera H, Suzuki A, Suzuki K, Wadhwa R, Elimova E, Sudo K, Shiozaki H, Estrella J, Lee JS, et al. Potentially functional variants in the core nucleotide excision repair genes predict survival in Japanese gastric cancer patients. Carcinogenesis. 2014; 35: 2031-8. doi: 10.1093/carcin/bgu142.
39. Lauren P. The two historical main types of gastric carcinoma: diffuse and so-called intestinal-type carcinoma. at attempt at a histo-clinical classification. Acta Pathol Microbiol Scand. 1965; 64: 31-49.
40. Fan J, Samworth R, Wu Y. Ultrahigh dimensional feature selection: beyond the linear model. J Mach Learn Res. 2009; 10: 2013-38.

