Introduction
Head and neck cancer (HNC) is a class of malignant tumors that arise in the tissues of the upper neck, such as the tongue, gingiva, nasopharynx, larynx, and thyroid [1-3]. Head and neck squamous-cell carcinoma (HNSCC) constitutes the majority (nearly 90%) of HNCs, with smoking and alcohol consumption as the major risk factors. In recent years, accumulating evidence suggests that human papillomavirus infection is closely related to the occurrence of HNC [4]. HNC constitutes an important component of cancer incidence and mortality, leading to approximately 225,100 new patients and 77,500 deaths in China every year [5]. Although advanced therapeutics have been applied to treat HNC, the 5-five-year survival rate has not increased significantly and the mortality rate has not decreased significantly in recent decades [6]. HNC is usually discovered and diagnosed in the advanced stages, a factor that significantly contributes to the high mortality rates; however, current treatments may be ineffective in suppressing HNC progression and have many side effects. Early diagnosis and effective therapy play an important role in HNC treatment [7-9]. Elucidation of the mechanisms of HNC carcinogenesis and progression may enable the discovery of early diagnostic biomarkers and effective therapeutic targets.
Long noncoding RNAs (lncRNAs) are a focus of current research, particularly cancer research. lncRNAs are noncoding polyadenylated RNAs longer than 200 bps. The transcription of lncRNAs is typically executed by RNA polymerase II, and their expression is generally tissue-specific [10]. Previously, lncRNAs have been considered byproducts of a leaky transcriptional system, but recent findings suggest a different role. As shown in numerous reports, lncRNAs are indispensable and are involved in a spectrum of biological processes, including embryogenesis, stem-cell biology and cellular differentiation. Anomalous lncRNAs are suggested to be closely associated with multiple human diseases, including cancer [10-15]. For HNC, a distinct lncRNA expression profile between HNC tumors and normal human tissues has been identified and it evolves as the HNC progresses [16]. Research into the cellular functions of lncRNAs has verified that lncRNAs play significant roles in HNC carcinogenesis and that lncRNA-mediated biology is related to HNC progression. lncRNAs may serve as oncogenes or tumor suppressors by participating in the modulation of HNC cell proliferation, differentiation, migration and invasion [16, 17]. Some cancer-associated lncRNAs have well-characterized roles in HNC development, suggesting that lncRNAs can be used as new biomarkers for HNC diagnosis and prognosis and as therapeutic targets for HNC intervention [17].
In this review, we will focus on the lncRNAs that are dysregulated in HNC, including oral cancer, laryngeal cancer, nasopharynx cancer and thyroid cancer. Additionally, we will summarize the latest findings regarding the biological function and molecular mechanisms of lncRNAs in HNC carcinogenesis and progression and explore the clinical application of lncRNAs as targets for HNC diagnosis, prognosis and therapy.
lncRNAs in HNC
Many lncRNAs are dysregulated in HNC and play important roles in HNC biology. Some lncRNAs that are up-regulated in HNC may act as oncogenes to promote cancer cell growth, migration, and invasion and to inhibit apoptosis. Some lncRNAs possess tumor-inhibiting functions and are down-regulated in HNC, accelerating oncogenesis and tumor development [16-18]. lncRNAs regulate gene transcription by associating with enzymes or protein complexes, thus participating in chromosome remodeling and epigenetic regulation. lncRNAs also regulate the transcription and translation of mRNAs by repressing miRNAs or altering mRNA modifications [10, 19, 20]. The biological functions and molecular mechanisms of specific lncRNAs in HNC are reviewed below (Table 1).
HOTAIR
HOTAIR (HOX antisense intergenic RNA) is a 2158-nucleotide mRNA-like lncRNA located at chromosome 12q13.13 that is transcribed from the antisense strand of the HOXC gene cluster by RNA polymerase II [21]. HOTAIR is not well conserved across species, and human HOTAIR contains 6 exons [22]. HOTAIR is up-regulated in many types of cancer and is related to oncogenesis, metastasis and poor prognosis in HNC, cervical cancer, and colon cancer [23-25]. HOTAIR recruits polycomb repressive complex 2 (PRC2) through its 5’-end binding domain and lysine-specific demethylase 1A (LSD1) through its 3’-end binding domain to targets to repress gene expression. The histone methylase enhancer of zeste homolog 2 (EZH2) is one component of PRC2, and the histone demethylase LSD1 is a flavin-dependent monoamine oxidase. HOTAIR interacts with EZH2 or LSD1, leading to gene silencing via H3K27-methylation or H3K4-demethylation, respectively [26, 27].
Recently, HOTAIR was shown to be dysregulated in HNC, including laryngeal squamous-cell carcinoma (LSCC), nasopharyngeal carcinoma (NPC) and oral squamous-cell carcinoma (OSCC) [28-32]. HOTAIR expression in LSCC tissues is 16-fold higher than in normal tissues, and increase in HOTAIR expression is statistically correlated with advanced tumor grade, lymph node metastasis (LNM), poor differentiation and advanced clinical stages. HOTAIR knockdown reduces cell invasion and increases cell apoptosis in vitro and inhibits LSCC xenograft growth in vivo. As revealed by a mechanistic study, HOTAIR serves as an oncogene by repressing phosphatase and tensin homolog deleted on chromosome ten (PTEN) expression to up-regulate DNA methylation. HOTAIR accumulation in cancer cells induces the hypermethylation of CpG islands in the PTEN gene, reducing the expression of PTEN at the mRNA and protein levels [28]. Because PTEN acts as a tumor repressor by inhibiting the phosphatidylinositol 3-kinase (PI3K) signaling pathway [33], the increased expression of HOTAIR in LSCC promotes cancer progression by activating the PI3K pathway [28]. HOTAIR expression is significantly increased in HNC and is correlated with cancer progression [29-32]. Additionally, in OSCC tumors, HOTAIR recruits EZH2 to the promoter region of E-cadherin and silences the expression of E-cadherin by increasing the levels of the H3K27me3 modification [30, 31]. HOTAIR is abundantly expressed in NPC tumor tissues and promotes tumor angiogenesis and growth. HOTAIR stimulates NPC angiogenesis by increasing the expression levels of the angiogenic factors vascular endothelial growth factor-A (VEGF-A) and glucose-regulated protein 78 (GRP78) [32]. GRP78, a member of the heat-shock protein 70 (HSP70) family, induces tumor angiogenesis and regulates the tumor microenvironment during tumor growth and metastasis [34, 35]. Furthermore, the expression of GRP78 in NPC increases the transcription of the angiogenic factors VEGF-A and angiopoietin 2 (Ang2) [32]. Thus, increased HOTAIR expression promotes cancer progression, and HOTAIR functions as an oncogene in HNC.
H19
The discovery of H19 has attracted the attention of many scientists and prompted them to study lncRNAs. H19 contains 5 exons and 4 introns and encodes a 2.3-kb noncoding RNA [11]. Insulin-like growth factor 2 (IGF2), a growth factor, stimulates tumor growth via autocrine or endocrine pathways [36]. Genomic imprinting is a type of hereditary epigenetic regulation that ensures the expression of parent-specific genes. Thus, only one of the parental imprinted alleles can be transcribed and translated [37]. H19 and IGF2 are a pair of imprinted genes on chromosome 11p15.5 that participate in embryonic development and growth regulation [11]. H19 gene is maternally expressed, and IGF2 is transcribed from the paternal allele. High H19 expression has been observed during embryogenesis, but the transcripts are not expressed in most tissues after birth, with the exception of the heart and skeletal muscle [38]. However, loss of imprinting (LOI) and loss of heterozygosity (LOH) have been observed at the IGF2-H19 locus in many cancers [39-45]. LOI at H19 leads to the transcription of H19 from the maternal and paternal alleles, and LOH at H19 inhibits the expression of H19. Hypomethylation in the H19 promoter region may be the main cause of H19 LOI. Moreover, c-Myc also activates H19 transcription in cancer by binding to the regulatory region in its gene. Mutation leads to H19 LOH, and the expression of H19 in cancer is down-regulated by p53 [40, 41]. Both LOI and LOH of H19 are associated with cancer carcinogenesis and progression. Re-expression of H19 in NPC suggests that H19 might be an oncogene [39], and the down-regulation of H19 in some HNSCC samples implies that H19 might be a tumor suppressor [42]. Although H19 has been enthusiastically studied for several years, the details of the mechanisms by which H19 functions as a tumor inhibitor or oncogene in cancer and its precise biological functions require further clarification.
Genomic imprinting plays important roles in organism development. LOI at the IGF2-H19 locus and aberrant H19 expression have been observed in HNC and may be related to HNC oncogenesis and progression [39, 42]. Among 64 HNSCC cases, 12 of 32 cases (37.5%) were showed a loss of imprinting at the H19 gene and 11 of 27 cases (40.7%) showed a for loss of imprinting at the IGF2 gene [44]. Among 27 samples from patients with Juvenile Nasopharyngeal Angiofibroma (JNA), 8 of 22 samples (36.4%) exhibited H19 over-expression and 7 of 19 samples (36.8%) exhibited IGF2 over-expression [43]. In another cohort of 35 HNSCC patients, 11 exhibited H19 re-expression. Among patients showing H19 expression, 6 had T2-grade tumors and 5 eventually presented recurrence and/or metastasis. In contrast, in patients without H19 expression, including 5 with T2-grade tumors, only 1 exhibited regional recurrence [45]. Up-regulation of H19 has been confirmed in undifferentiated NPC biopsies and LSCC tumor tissues and is related to low LSCC survival rates and cancer progression [39]. Hypomethylation of CpG dinucleotides in the H19 promoter region may be the main epigenetic event that induces H19 transcription [39]. The level of DNA methylation in the H19 promoter region regulates H19 expression, and the level of cellular DNA methylation is also regulated by H19. miR-148a-3p is a target of inhibition by H19, and miR-148a-3p reduces the levels of the DNA methyltransferase enzyme 1 (DNMT1) mRNA and protein, affecting cellular DNA methylation. Thus, H19 regulates the level of cellular DNA methylation through the H19/miR-148a-3p/DNMT1 cascade, and H19 may promote HNC cell migration, invasion and proliferation by increasing DNA methylation [46].
MEG3
Maternally Expressed Gene 3 (MEG3), a tumor suppressor, is involved in the development and progression of many cancers [47-49]. MEG3 is an imprinted gene located at chromosome 14q32 that encodes a 1.6-kb noncoding RNA from the maternal allele [50]. It is broadly distributed in many normal human tissues; however, MEG3 is not expressed in a series of human tumors and tumor cell lines, such as tongue squamous-cell carcinoma (TSCC), brain cancer, and hepatocellular cancer [47, 48, 51]. One of the main mechanisms of MEG3 silencing is hypermethylation of the MEG3 promoter region [52]. The loss of MEG3 expression promotes tumor progression through specific molecular mechanisms, and re-expression of MEG3 in tumor cells inhibits proliferation and induces apoptosis [51]. MEG3 inhibits tumorigenesis by increasing the expression of P53 and Growth Differentiation Factor 15 (GDF15). p53 is a tumor suppressor protein, and GDF15 reduces tumor cell proliferation and tumor formation. MEG3 increases p53 levels by reducing its ubiquitin-proteasome-mediated degradation [53]. MEG3 can down-regulate the E3 ubiquitin ligase MDM2, which mediates p53 ubiquitination and increase its stability and expression levels in the cell. GDF15 is also a target of MEG3, which can increase GDF15 expression in a p53-dependent manner [53].
Low MEG3 expression is associated with TSCC progression. MEG3 levels are significantly reduced in TSCC tissues compared with adjacent noncancerous tissues. Low MEG3 levels are associated with high TSCC mortality and poor overall survival. Over-expression of MEG3 in SCC-15 and CAL27 cell lines inhibits cell proliferation, arrests the cell cycle and induces apoptosis [51]. The reduction of the miR-26a levels in TSCC down-regulates MEG3 by increasing the DNA methylation levels. DNA methyltransferase 3B (DNMT3B) is a key enzyme in the DNA methylation process. DNMT3B mRNA contains a miR-26a target site in its 3’untranslated region (3’UTR), and miR-26a reduces the translation of DNMT3B by binding to its mRNA. Down-regulated of miR-26a in TSCC increases the level of DNMT3B, promoting DNA methylation and inhibiting MEG3 transcription [51, 54].
MALAT1
Metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) was first identified in non-small cell lung cancer (NSCLC). Up-regulation of MALAT1 in NSCLC is associated with cancer metastasis and is a biological marker used to predict disease outcomes in patients with early-stage NSCLC [55]. A MALAT1 transcript longer than 8,000 bps is abundantly expressed in the nucleus and is also known as nuclear-enriched abundant transcript 2 (NEAT2) [56]. MALAT1 was reported to be specifically up-regulated in many cancers, such as ovarian cancer, gastric cancer, and breast cancer, in several studies [57-59]. High MALAT1 expression levels in cancer promote cell proliferation, migration and invasion. MALAT1 accelerates cancer progression by regulating the expression of some ‘metastatic signature’ genes [60]. MALAT1 regulates gene expression at the transcriptional level by affecting the alternative splicing process and modulating the location and phosphorylation states of serine/arginine (SR) splicing factors [61].
Higher MALAT1 expression levels have been observed in LSCC tumor tissues and are significantly associated with a poor histological grade or an advanced clinical stage [30]. MALAT1 is also highly expressed in NPC tissues and it is closely related to LNM and advanced NPC stages [62]. Interference with MALAT1 expression in cancer inhibits tumor growth and metastasis and increases tumor cell apoptosis and sensitivity to radiotherapy [62, 63]. The Slug protein plays a considerable role in elevating cancer stem-cell (CSC) activity and tumor radioresistance. MALAT1 increases the levels of the slug mRNA and protein by acting as a competing endogenous RNA. The binding of MALAT1 to miR-1 reduces the levels of miR-1 and the slug mRNA. MALAT1 eliminates miR1-induced slug gene silencing [64].
Other lncRNAs in HNC
lncRNA urothelial cancer-associated 1 (UCA1) is involved in cancer invasion and metastasis. UCA1 is expressed at a dramatically higher level in TSCC tumor tissues than in neighboring non-tumor tissues, and patients with LNM express much higher levels of UCA1. High UCA1 expression in TSCC increases cell migration [65]. The specific mechanisms by which UCA1 contributes to TSCC progression remain to be elucidated.
Forkhead box C1 (FOXC1) is a crucial cancer-related gene. FOXC1 promotes cancer progression by regulating the epithelial-mesenchymal transition (EMT), cell proliferation and migration [66]. FOXC1 regulates the expression of matrix metalloproteinases (MMPs) and VEGF-A. MMPs regulate cancer cell migration and VEGF-A promotes tumor angiogenesis. FOXC1 upstream transcript (FOXCUT) is a lncRNA that is substantially over-expressed in OSCC. Over-expression of FOXCUT in OSCC promotes tumor cell proliferation and migration by regulating FOXC1 expression. The levels of the FOXC1 mRNA are positively associated with the levels of the FOXCUT transcript, and FOXCUT silencing substantially reduces FOXC1 expression at both the mRNA and protein levels [67].
The lncRNA nuclear paraspeckle assembly transcript1 (NEAT1) is an oncogene in many malignancies, including LSCC [68-71]. High NEAT1 expression is closely related to LSCC tumorigenesis, and patients with neck nodal metastasis exhibit high NEAT1 expression. NEAT1 knockdown in LSCC suppresses tumor growth, increases cell apoptosis and arrests the cell cycle. NEAT1 promotes LSCC tumorigenesis by regulating the miR-107/cyclin-dependent kinase 6 (CDK6) pathway [71]. The CDK6 protein belongs to the CDK family, regulating the transition from G1 to S phase of the cell cycle [72]. miR-107 represses CDK6 expression by targeting its 3’UTR. NEAT1 increases CDK6 expression by down-regulating miR-107 expression [71].
The lncRNA actin filament-associated protein 1 antisense RNA1 (AFAP1-AS1) is up-regulated in NPC and is related to NPC progression and poor patient survival. AFAP1-AS1 knockdown represses NPC cell invasion and migration in vitro and inhibits NPC lung metastasis in nude mice. AFAP1-AS1 regulates AFAP1 and some cytoskeleton-regulated proteins, including proteins in the small GTPase-signaling Rho/Rac pathway [73]. AFAP1 regulates actin filament integrity and stimulates lamellipodia formation. Small GTPases modulate actin cytoskeleton formation, which is associated with cell shape and migration [74]. AFAP1-AS1 promotes NPC cell metastasis by affecting the integrity of actin filaments [73].
The lncRNA hepatocyte nuclear factor 1A-antisense RNA (HNF1A-AS) may modulate tumorigenesis in NPC. HNF1A-AS expression is much higher in NPC tumor tissues than in matched normal tissues. Over-expression of HNF1A-AS in NPC increases tumor cell proliferation, migration and cell cycle progression. HNF1A-AS may stimulate NPC tumor growth and metastasis by accelerating the EMT. In NPC, HNF1A-AS increases the levels of the mesenchymal proteins N-cadherin and vimentin and reduces the level of epithelial protein E-cadherin [75].
Radioresistance and chemoresistance are key drivers of cancer mortality, and the lncRNA-ROR is related to NPC chemoresistance. lncRNA-ROR expression is significantly increased in NPC tissues compared with in normal tissues. lncRNA-ROR expression in NPC cells is positively correlated with cell proliferation, metastasis and suppression of apoptosis. High lncRNA-ROR expression increase the ability of NPC cells to resist chemotherapy. lncRNA-ROR facilitates NPC chemoresistance and progression, possibly by inhibiting the p53 signaling pathway. lncRNA-ROR may constitute a medical target by which to decrease NPC chemoresistance [76].
lncRNA-LET (low expression in tumor) is a tumor suppressor that is down-regulated in NPC tumor tissue and NPC cell lines compared with paired non-cancer tissues and nasopharyngeal epithelial cells. LET down-regulation is significantly associated with an advanced NPC clinical stage, a greater tumor volume, an increased lymph node tumor burden, and a lower overall survival rate. Forced LET expression in CNE2 cells inhibits cell proliferation and increases cell apoptosis. Injection of LET-over-expressing cells into nude mice reduces tumor growth, and low lncRNA-LET levels are critically involved in NPC tumor cell proliferation. Further research suggests that low levels of LET expression are induced by EZH2-mediated H3K27 histone methylation in the LET promoter region [77].
A study of gene mutations in 402 Chinese NPC samples revealed that the growth arrest-specific 8-antisense RNA 1 (GAS8-AS1) is the second most commonly mutated gene and the lncRNA GAS8-AS1 may serve as a tumor inhibitor. The levels of the lncRNA GAS8-AS1 are significantly decreased in PTC. Increased GAS8-AS1 expression substantially inhibits PTC cell viability. Furthermore, GAS8-AS1 mutations in PTC are associated with an advanced clinical stage, and the wild-type lncRNA GAS8-AS1 is a stronger suppressor of tumor growth than mutated lncRNA GAS8-AS1 [78].
Recently, a novel lncRNA LOC401317 was shown to be up-regulated in human NPC cell lines over-expressing TP53, and the transcription of the lncRNA LOC401317 is directly regulated by p53. As revealed in functional studies, LOC401317 is a tumor suppressor and LOC401317 transfection in NPC cells represses tumor cell proliferation, arrests the cell cycle and increases cell apoptosis. Mechanistic investigations revealed that LOC401317 inhibits the cell cycle in cancer cells by increasing the level of p21 and reducing the levels of cyclins D1 and E1. LOC401317 expedites apoptosis by inducing the activation of Poly (ADP-ribose) polymerase (PARP) and caspase-3 [79].
The levels of the lncRNA plasmacytoma variant translocation 1 (PVT1) are significantly increased in thyroid cancer (TC) tissues compared to neighboring normal tissues. PVT1 silencing in TC cell lines obviously inhibits cell proliferation and arrests the cell cycle at the G0/G1 phase. PVT1 increases the mRNA and protein levels of thyroid-stimulating hormone receptor (TSHR) and the cell cycle-associated protein cyclin D1 [80]. Thyroid-stimulating hormone (TSH) and TSHR are associated with cell proliferation [81]. Furthermore, the lncRNA PVT1 recruits EZH2 to target genes, repressing the expression of some tumor suppressor genes. Recruitment of EZH2 and regulation of the TSHR levels may be the two mechanisms by which the lncRNA PVT1 facilitates TC oncogenesis [80].
A novel lncRNA, non-protein coding RNA, associated with MAP kinase pathway and growth arrest (NAMA), is, as its name suggests, related to the MAP kinase pathway and cell-growth arrest. NAMA levels are reduced in papillary thyroid carcinoma (PTC). NAMA over-expression in PTC cells promotes cell apoptosis and arrests the cell cycle. NAMA expression is induced by BRAF knockdown, suppression of the MAP kinase pathway, growth arrest and DNA damage. NAMA induction may be a molecular event or secondary signal that induces growth arrest and apoptosis [82]. However, further research is required to explore the functions and potential targets of NAMA.
Papillary thyroid cancer susceptibility candidate 2 (PTCSC2) and 3 (PTCSC3) are two novel thyroid-specific lncRNAs that may serve as tumor suppressors. The expression levels of PTCSC2 and PTCSC3 are strongly down-regulated in PTC tumor tissues and in TC tissues, respectively, compared with neighboring normal tissues. Over-expression of PTCSC2 in PTC cells influences the expression of some cancer-related and cell-cycle-related genes. Re-expression of PTCSC3 in TC cells represses tumor growth and influences the transcription of some genes associated with DNA replication, recombination, repair, cellular movement, tumor morphology, and cell death. Down-regulation of PTCSC2 or PTCSC3 is related to PTC or TC and increases the risk of developing PTC or TC. PTCSC2 and PTCSC3 may be involved in the genetic predisposition to PTC and TC. The polymorphisms rs965513 and rs944289 are related to PTC and TC risk via PTCSC2 and PTCSC3, respectively. The risk allele (A) of rs965513 is significantly associated with down-regulation of PTCSC2 in normal thyroid tissues. The risk allele (T) of rs944289 is strongly related to the down-regulation of PTCSC3 in TC tumor tissues [83, 84].
The connection between the level of the lncRNA HOXA transcript at the distal tip (HOTTIP) and patients’ clinicopathological characteristics has been explored. High HOTTIP expression has been confirmed in TSCC and is positively correlated with high T classification, advanced clinical stages and distant metastasis [85]. The lncRNA ENST00000438550 was identified in an analysis of the differences in the lncRNA expression profiles between metastatic and primary NPC tumors. ENST00000438550 is strongly associated with the progression of NPC patients, and ENST00000438550 might be a prognostic biomarker and a potential treatment target in NPC [86]. An investigation of 100 PTC samples shows that focally amplified lncRNA on chromosome 1 (FAL1) is expressed at higher levels in PTC tissues than in surrounding healthy tissues. FAL1 over-expression greatly increases the risk of multifocality, and FAL1 might be associated with PTC malignancy [87]. The lncRNA AB209630 may be a tumor suppressor in hypopharyngeal squamous cell carcinoma (HSCC). Down-regulation of AB209630 promotes HSCC tumor growth, metastasis and invasion. HSCC patients with high AB209630 levels have a better prognosis than patients with low AB209630 levels. AB209630 may be a valuable target in HSCC therapy [88]. lncRNAs play an important role in promoting HNC tumorigenesis and development. The known mechanisms by which some lncRNAs function in HNC are shown in Figure 1, and the as yet unknown mechanisms require further research.
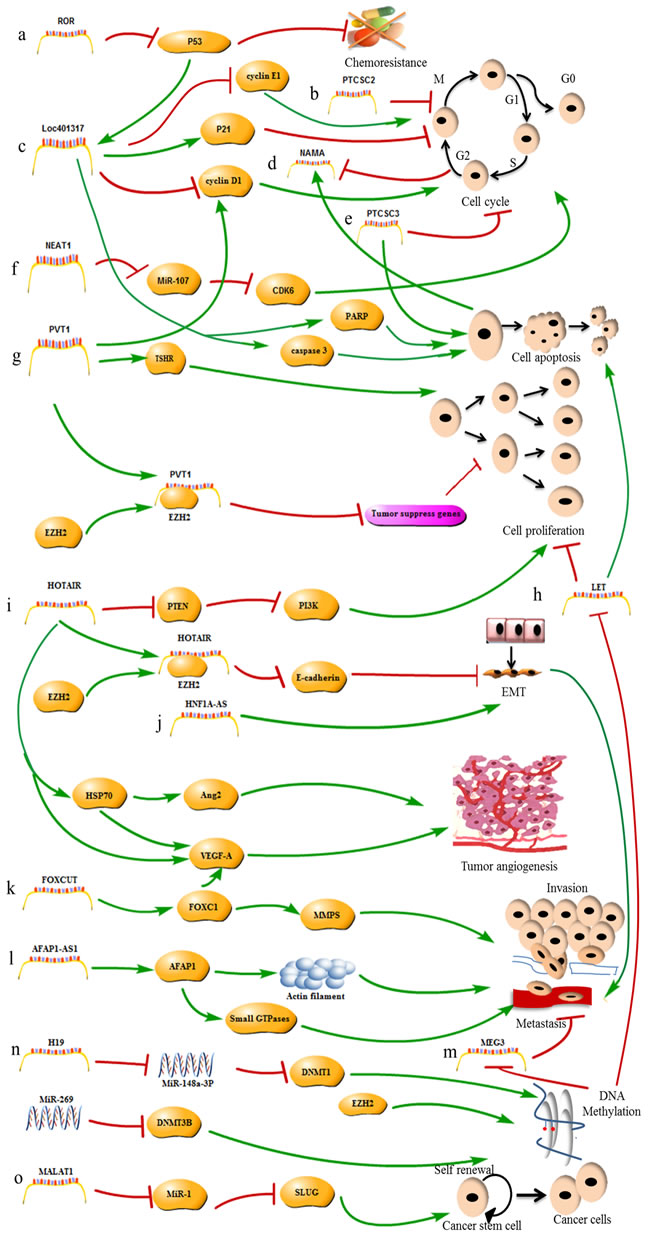
Figure 1: The mechanisms of lncRNAs in HNC. a. The lncRNA-ROR promotes tumor chemoresistance by repressing P53 expression. b. The lncRNA PTCSC2 inhibits tumor cell cycle progression. c. The lncRNA LOC401317 inhibits tumor cell cycle progression by inducing P21 expression and repressing cyclin D1 and E1 expression, and promotes apoptosis by inducing PARP and caspase 3 expression; lncRNA LOC401317 expression is induced by p53 activity. d. The lncRNA NAMA represses tumor growth, and its expression is induced by cell cycle arrest and apoptosis. e. The lncRNA PTCSC2 inhibits tumor cell cycle progression and promotes apoptosis; f. The lncRNA NEAT1 promotes cell cycle progression by inhibiting the miR-107/ CDK6 pathway. g. The lncRNA PVT1 promotes tumor cell cycle progression by inducing cyclin D1 expression and promotes cell proliferation by inducing TSHR expression and recruiting EZH2 to repress the expression of tumor-suppressor genes. h. lncRNA-LET inhibits tumor cell proliferation and promotes apoptosis. LET is inhibited by EZH2-mediated H3K27 methylation. i. The lncRNA HOTAIR promotes cell proliferation by inducing PTEN expression, promotes metastasis by recruiting EZH2 to accelerate the EMT, and promotes tumor angiogenesis by inducing VEGF-A and HSP70 expression. j. The lncRNA HNF1A-AS promotes metastasis by accelerating the EMT. k. The lncRNA FOXCUT promotes invasion and metastasis by inducing FOXC1 expression. l. The lncRNA AFAP1-AS1 promotes metastasis by inducing AFAP1 expression. m. The lncRNA MEG3 promotes metastasis and is inhibited by miR-269-induced DNA methylation. n. The lncRNA H19 promotes tumorigenesis by regulating DNA methylation. o. The lncRNA MALAT1 increases CSC activity by inhibiting miR-1 expression.
Clinical applications of lncRNAs in HNC
lncRNAs are broadly involved in HNC oncogenesis and progression. The key roles of lncRNAs in HNC imply that lncRNAs may serve as biomarkers or therapeutic targets for HNC diagnosis, prognosis and treatment.
Tumor diagnosis
The early diagnosis of HNC improves overall survival. lncRNAs play a vital role in the physiological and pathological progression of HNC, and the presence of lncRNAs in peripheral blood and body fluids suggests that they may be used as diagnostic biomarkers [89, 90].
Diagnostic biomarkers in OSCC
Some well-documented lncRNAs, such as MALAT-1 and HOTAIR, are present in the saliva of OSCC patients. Patients with LNM exhibit higher HOTAIR expression in their saliva than patients with non-metastatic cancer. The presence of some lncRNAs in patients’ saliva may be associated with OSCC development, and measuring the lncRNA levels in saliva may provide a convenient method to rapidly and non-invasively diagnose OSCC [90]. HOTAIR expression in OSCC tissues is also correlated with a larger tumor size and advanced clinical stages, suggesting that HOTAIR may be used as a biomarker for diagnosing OSCC [29].
Diagnostic biomarkers in LSCC
The combination of the serum levels of the HOTAIR and miR-21 exosomes is valuable for diagnosing LSCC. The serum levels of HOTAIR and miR-21 exosomes were higher in 52 LSCC patients than in 49 patients with vocal cord polyps, and high expression levels of miR-21 and HOTAIR in LSCC patients are significantly associated with advanced clinical stages and LNM. Moreover, the area under the receiver-operating characteristic (ROC) curve of the combination of HOTAIR and miR-21 is 87.6%, which is greater than the value observed for serum exosomes of miR-21 (80.1%) or HOTAIR (72.7%). The combination of HOTAIR and miR-21 might be a promising molecular marker to screen for LSCC [91].
Diagnostic biomarkers in NPC
LINC00312 is distinctly down-regulated in NPC, and positive LINC00312 expression is negatively correlated to the clinical classification and tumor size. LINC00312 expression is used to distinguish healthy persons from NPC patients. Down-regulation of LINC00312 may contribute to NPC oncogenesis and may be used to diagnose early-stage NPC. The dysregulation of lncRNAs in HNC may be a promising indicator for use in providing an exact and early cancer diagnosis [92].
Tumor prognosis
The dysregulation of many lncRNAs is significantly related to HNC pathogenesis and prognosis. These molecules may be used as potential prognostic predictors, and their expression levels may predict patients’ disease outcomes [17].
Prognostic biomarkers in OSCC
HOTTIP, MEG3, and HOTAIR could be used as independent prognostic parameters in TSCC. High HOTTIP expression has been observed in TSCC tissues, and patients with T3-4-grade tumors, distant metastasis or patients in clinical stages III-IV exhibit higher HOTTIP expression than patients with T1-2-grade tumors and no distant metastasis or patients in clinical stages I-II. HOTTIP over-expression is strongly associated with poor overall survival (OS) [85]. MEG3 expression is much lower in TSCC tumor tissues than in nearby nonmalignant tissues, and patients with low MEG3 expression always have short survival times [51]. HOTAIR expression is higher in OSCC tumor tissue than in paired normal tissue, and high HOTAIR expression always indicates a relatively poor OS or disease-free survival (DFS) [29].
Prognostic biomarkers in NPC
The lncRNAs HOTAIR, AFAP1-AS1 and ENST00000438550 could be used as prognostic biomarkers to predict disease outcomes in NPC patients [23, 73, 86]. Excessive AFAP1-AS1 levels have been detected in NPC samples compared with normal nasopharyngeal epithelium samples. AFAP1-AS1 up-regulation is related to distant tumor metastasis and poor OS and relapse-free survival (RFS) [73]. HOTAIR expression is also higher in paraffin-embedded NPC biopsies than in non-tumor tissue samples, and high HOTAIR levels are associated with advanced clinical stages and a poor prognosis [23]. The ENST00000438550 expression level is positively correlated with NPC progression, and high levels of ENST00000438550 are an indicator of NPC progression. Some down-regulated lncRNAs are also used as molecule tools to predict NPC outcomes [86]. The Kaplan-Meier method and log-rank test have been used to analyze the predictive potential of lncRNAs. Down-regulation of LET in NPC is related to poor RFS and OS, and low LET expression may predict a poor prognosis of NPC patients [77]. Positive LINC00312 expression in NPC patients without LNM is significantly correlated with good DFS and OS; however, high LINC00312 expression in patients with LNM is correlated with poor DFS and OS. LINC00312 may be a powerful independent indicator of NPC patient survival [92].
Prognostic biomarkers in LSCC
The lncRNA HOTAIR might be a diagnostic marker for LSCC and an independent prognostic factor for LSCC [28]. HOTAIR levels are higher in LSCC tumor tissues than in adjacent non-neoplastic tissues and are significantly associated with shorter patient survival. HOTAIR has a major effect on LSCC progression and is an independent prognostic indicator [28]. Low expression levels of AC026166.2-001 and RP11-169D4.1-001 are statistically linked to OS in LSCC patients and are independent indicators of a poor prognosis [93]. Thus, lncRNAs may be used as prognostic factors to forecast cancer outcomes.
Table 1: Dysregulation of lncRNAs in HNC
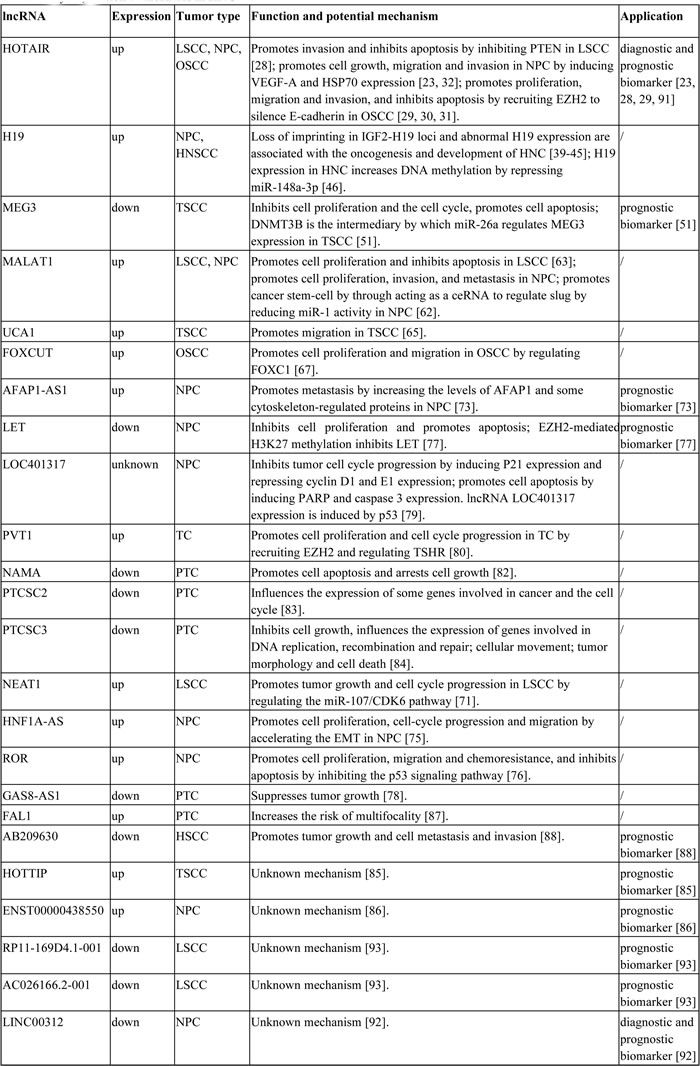
Abbreviations: LSCC, laryngeal squamous-cell carcinoma; NPC, nasopharyngeal carcinoma; OSCC, oral squamous cell carcinoma; HNSCC, head and neck squamous cell carcinoma; TSCC, tongue squamous cell carcinoma; TC, thyroid cancer; PTC, papillary thyroid carcinoma; HSCC, hypopharyngeal squamous cell carcinoma.
Tumor treatment
The considerable importance of abnormal lncRNAs expression in HNC development and progression suggests that lncRNAs are potential therapeutic targets. Some tumorigenic lncRNAs are up-regulated in HNC and promote cancer growth. However, their cancer-promoting effects could be weakened by interference by siRNAs [28-31]. Reduction of the HOTAIR or MALAT1 levels in LSCC inhibits cell proliferation and induces apoptosis [28, 63]. In addition, tumor-suppressing lncRNAs are expressed at low levels in cancer tissues, and transfection of an lncRNA expression vector in cancer cells to increase their expression inhibits tumor growth [50, 51]. MEG3 expression is substantially reduced in OSCC tumor cell lines. Increased MEG3 levels in OSCC cells suppress cell proliferation, disrupt the cell cycle and stimulate apoptosis [51]. Thus, restoration of the expression of these lncRNAs in HNC may be a feasible therapeutic approach.
Some medical treatments for cancer target dysregulated lncRNAs and reduce the levels of aberrantly expressed lncRNAs in tumor tissues [93, 94]. The lncRNAs CDKN2B-AS1, HOTAIR and MALAT1 are up-regulated in LSCC. The levels of CDKN2B-AS1, HOTAIR and MALAT1 in LSCC cells are substantially reduced by cisplatin and paclitaxel in a concentration- and time-dependent manner [94]. lncRNA-MBL2-4:3 is significantly up-regulated and lncRNA-AL355149.1-1 is significantly down-regulated in TSCC tumor tissues. The expression levels of these lncRNAs are restored by the injection of 5-fluorouracil and paclitaxel in HN21B TSCC cells in a dose-dependent manner [93]. Thus, dysregulated lncRNAs are potential targets of HNC treatments.
lncRNAs are related to HNC radioresistance [95]. According to some analyses, the plasma levels of the lncRNA GAS5 in HNC patients respond to radical chemoradiotherapy. The pre- and post-radiotherapy GAS5 levels are significantly higher in HNC patients with a partial response to radiotherapy than in HNC patients exhibiting complete responses. Plasma GAS5 levels are a potential molecule biomarker for predicting radiotherapy responses in HNC patients [95]. MALAT1 expression is substantially increased in NPC tissues. Increased MALAT1 expression reduces NPC radiation sensitivity by improving the self-renewal of CSCs. MALAT1 knockdown in NPC sensitizes the tumor cells to ionizing radiation and reduces the percentage of ALDH1-postive cells displaying properties associated with CSCs [64]. Differences in lncRNAs expression have been observed between NPC cells treated with ionizing radiation and NPC cells treated without ionizing radiation. Curcumin alters lncRNA expression in NPC cells while increasing the susceptibility of NPC cells to ionizing radiation [96]. lncRNAs play a significant role in cancer radioresistance. lncRNA-based therapeutics might represent a new direction for cancer treatment, although numerous experiments must be performed.
Conclusions
Whole-genome sequencing studies have revealed that protein-coding genes constitute only a small portion, approximately 1.5%, of the genome. Most genes in the genome are transcribed into noncoding RNAs (ncRNAs), including lncRNAs, by RNA polymerase II [7]. An increasing number of reports indicate that lncRNAs significantly contribute to cancer biology, including HNC biology [15]. Dysregulated lncRNAs are significantly correlated with cancer development and progression in HNC [16]. The successful identification and characterization of lncRNAs may be a feasible strategy for studying lncRNAs in cancer. Microarray technology is one prospective method for screening irregular lncRNAs expression in malignant cancers. Knockdown or over-expression of certain lncRNAs in vitro and in vivo may reveal their functions and mechanisms in cancer biology. Some lncRNAs modulate gene expression at the transcriptional or post-transcriptional levels [61]. Some lncRNAs interact with DNA, RNA or proteins to exert their gene-regulating functions and affect the behavior of cancer cells. However, lncRNAs have multiple intricate functions and molecular mechanisms, and only a small fraction of lncRNAs and their functions in cancer biology have been discovered and investigated. In addition, the functions of the few known lncRNAs remain to be investigated in detail and additional functional lncRNAs remain to be identified.
Differential lncRNA expression between HNC tumors and normal tissues suggests that lncRNAs have potential for use as sensitive biomarkers in HNC detection and prognosis [16]. For example, high HOTAIR expression levels are related to HNC development, and lower levels of EMG3 are related to cancer metastasis [31, 51]. The correlations between lncRNA expression in patient saliva and the clinical stages of OSCC provide markers for a noninvasive and rapid diagnosis [90]. Over-expression of some tumor-promoting lncRNAs and reduced expression of some tumor-suppressing lncRNAs are related to poor OS and DFS. Cancer-related lncRNAs could be used as independent prognostic parameters in HNC [51, 91]. Although some reports have identified lncRNAs in body fluids [90, 91], the stability of peripheral lncRNAs remains largely unknown and lncRNA degradation may restrict the use of lncRNAs as biomarkers. Although some lncRNAs are negatively or positively correlated with the outcomes of HNC patients, the analysis of large samples is required to assess the utility of lncRNAs in clinical predictions.
As reported in numerous studies, dysregulation of lncRNAs plays a major role in HNC development, and lncRNAs are promising novel therapeutic targets for cancer intervention [4] because oncogenic lncRNAs are up-regulated and tumor-suppressing lncRNAs are down-regulated in HNC. Thus, the modulation of ectopic lncRNA expression is one direction for the development of lncRNA-based cancer therapies. For example, knockdown of HOTAIR or MALAT1 with siRNAs inhibits cancer-cell proliferation and increases apoptosis [28, 63]. Although success has been achieved in cell-based experiments in which lncRNAs served as therapeutic targets, lncRNA-targeted cancer therapies must overcome many obstacles before its implementation, and a tremendous amount of research is required to elucidate lncRNA structures, folding, functions, mechanisms and interactions with other molecules. The development of lncRNA-targeted therapies will require a detailed knowledge of the functions of lncRNAs in HNC.
In summary, dysregulated lncRNAs play an essential role in HNC tumorigenesis and progression. They may serve as oncogenes or tumor inhibitors, and further research is needed to determine their precise, detailed mechanisms and functions. The importance of lncRNAs in cancer suggests the possibility of developing lncRNA-oriented clinical applications. Although this problem presents some difficulties, there is also the strong possibility of significant advancements in the near future.
Abbreviations
HNC: head and neck cancer; HNSCC: head and neck squamous cell carcinoma; lncRNAs: long noncoding RNAs; HOTAIR: HOX antisense intergenic RNA; PRC2: polycomb repressive complex 2; LSD1: lysine-specific demethylase 1A; EZH2: histone methylase enhancer of Zeste homolog 2; LSCC: laryngeal squamous cell carcinoma; NPC: nasopharyngeal carcinoma; OSCC oral squamous cell carcinoma; LNM: lymph node metastasis; PTEN: tensin homolog deleted on chromosome ten; PI3K: phosphatidylinositol 3-kinase; VEGF-A: vascular endothelial growth factor-A; GRP78: glucose-regulated protein 78; HSP70: heat shock protein 70; IGF2: insulin-like growth factor 2; LOI: loss of imprinting; LOH: loss of heterozygosity; DNMT1: DNA methyltransferase enzyme 1; TSCC: tongue squamous cell carcinoma; GDF15: growth differentiation factor 15; DNMT3B: DNA methyltransferase 3B; MALAT1: metastasis-associated lung adenocarcinoma transcript 1; NSCLC: non-small cell lung cancer; NEAT2: nuclear-enriched abundant transcript 2; SR: serine/arginine; CSC: cancer stem cells; UCA1: urothelial cancer-associated 1; FOXC1: forkhead box C1; FOXCUT: FOXC1 upstream transcript; EMT: epithelial-mesenchymal transition; NEAT1: nuclear paraspeckle assembly transcript1; AFAP1-AS1: actin filament associated protein 1-antisense RNA1; HNF1A-AS: hepatocyte nuclear factor 1A-antisense RNA; GAS8-AS1: growth arrest-specific 8-antisense RNA 1; PARP: poly (ADP-ribose) polymerase; PVT1: plasmacytoma variant translocation 1; TSHR: thyroid-stimulating hormone receptor; TSH: thyroid-stimulating hormone; NAMA: non-protein coding RNA, associated with MAP kinase pathway and growth arrest; TC: thyroid cancer; PTC: papillary thyroid carcinoma; PTCSC2: papillary thyroid cancer susceptibility candidate 2; PTCSC3: papillary thyroid cancer susceptibility candidate 3; HOTTIP: HOXA transcript at the distal tip; FAL1: focally amplified lncRNA on chromosome 1; HSCC: hypopharyngeal squamous cell carcinoma; ROC: receiver-operating characteristic; OS: overall survival; DFS: disease-free survival; RFS: relapse-free survival; ncRNAs: noncoding RNAs.
Acknowledgments
This work was supported by the Natural Science Foundation of Shanghai Province, China (15ZR1413000).
Conflicts of interests
No potential conflicts of interest exist.
References
1. Rettig EM, D’Souza G. Epidemiology of head and neck cancer. Surg Ocol Clin N Am. 2015; 24: 379-396. doi: 10.1016/j.soc.2015.03.001.
2. Petersson F. Nasopharyngeal carcinoma: a review. Semin Diagn Pathol. 2015; 32: 54-73. doi: 10.1053/j.semdp.2015.02.021.
3. Metcalfe CW, Lowe D, Rogers SN. What patients consider important: temporal variations by early and late stage oral, oropharyngeal and laryngeal subsites. J Craniomaxillofac Surg. 2014; 42: 641-647. doi: 10.1016/j.jcms.2013.09.008.
4. Spence T, Bruce J, Yip KW, Liu FF. HPV associated head and neck cancer. Cancers. 2016; 8: 75. doi: 10.3390/cancers8080075.
5. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F. A J, Yu X, He J. Cancer Statistics in China, 2015. CA Cancer J Clin. 2016; 66: 115-132. doi: 10.3322/caac.21338.
6. Rahimy E, Kuo SZ, Ongkeko MW. Evaluation of non-coding RNAs as potential targets in head and neck squamous cell carcinoma cancer stem cells. Curr Drug Targets. 2014; 15: 1247-1260. doi: 10.2174/1389450115666141024113446.
7. Zecha JA, Raber-Durlacher JE, Nair RG, Epstein JB, Sonis ST, Elad S, Hamblin MR, Barasch A, Migliorati CA, Milstein DM, Genot MT, Lansaat L, van der Brink R, et al. Low level laser therapy/photobiomodulation in the management of side effects of chemoradiation therapy in head and neck cancer: part 1: mechanisms of action, dosimetric, and safety considerations. Support Care Cancer. 2016; 24: 2781-2792. doi: 10.1007/s00520-016-3153-y.
8. Zecha JA, Raber-Durlacher JE, Nair RG, Epstein JB, Elad S, Hamblin MR, Barasch A, Migliorati CA, Milstein DM, Genot MT, Lansaat L, van der Brink R, Arnabat-Dominguez J, et al. Low-level laser therapy/photobiomodulation in the management of side effects of chemoradiation therapy in head and neck cancer: part 2: proposed applications and treatment protocols. Support Care Cancer. 2016; 24: 2793-2805. doi: 10.1007/s00520-016-3153-y.
9. Dunne S, Mooney O, Coffey L, Sharp L, Desmond D, Timon C, O’Sullivan E, Gallagher P. Psychological variables associated with quality of life following primary treatment for head and neck cancer: a systematic review of the literature from 2004 to 2015. Psychooncology. 2016. doi: 10.1002/pon.4109.
10. Schmitz SU, Grote P, Herrmann BG. Mechanisms of long noncoding RNA function in development and disease. Cell Mol Life Sci. 2016; 73: 2491-2509. doi: 10.1007/s00018-016-2174-5.
11. Ratajczak MZ. Igf2-H19, an imprinted tandem gene, is an important regulator of embryonic development, a guardian of proliferation of adult pluripotent stem cells, a regulator of longevity, and a‘passkey’to cancerogenesis. Folia Histochem Cytobiol. 2012; 50: 171-179. doi: 10.5603/FHC.2012.0026.
12. Audas TE, Lee S. Stressing out over long noncoding RNA. Biochim Biophys Acta. 2016; 1859: 184-191. doi: 10.1016/j.bbagrm.2015.06.010.
13. Martens-Uzunova ES, Böttcher R, Croce CM, Jenster G, Visakorpi T, Calin GA. Long noncoding RNA in prostate, bladder, and kidney cancer. Eur Urol. 2014; 65: 1140-1151. doi: 10.1016/j.eururo.2013.12.003.
14. Peng Z, Zhang C, Duan C. Functions and mechanisms of long noncoding RNAs in lung cancer. Onco Targets Ther. 2016; 9: 4411-4424. doi: 10.2147/OTT.S109549.
15. Li CH, Chen Y. Targeting long non-coding RNAs in cancers: progress and prospects. Int J Biochem Cell Biol. 2013; 45: 1895-1910. doi: 10.1016/j.biocel.2013.05.030.
16. Zou AE, Ku J, Honda TK, Yu V, Kuo SZ, Zheng H, Xuan Y, Saad MA, Hinton A, Brumund KT, Lin JH, Wang-Rodriguez J, Ongkeko WM. Transcriptome sequencing uncovers novel long noncoding and small nucleolar RNAs dysregulated in head and neck squamous cell carcinoma. RNA. 2015; 21: 1122-1134. doi: 10.1261/rna.049262.114.
17. Yang QQ, Deng YF. Long non-coding RNAs as novel biomarkers and therapeutic targets in head and neck cancers. Int J Clin Exp Pathol. 2014; 7: 1286-1292.
18. González-Ramírez I, Soto-Reyes E, Sánchez-Pérez Y, Herrera LA, García-Cuellar C. Histones and long non-coding RNAs: the new insights of epigenetic deregulation involved in oral cancer. Oral Oncol. 2014; 50: 691-695. doi: 10.1016/j.oraloncology.2014.04.006.
19. Wu Y, Huang C, Meng X, Li J. Long noncoding RNA MALAT1: insights into its biogenesis and implications in human disease. Curr Pharm Des. 2015; 21: 5017-5028. doi: 10.2174/1381612821666150724115625.
20. Zeng S, Xiao YF, Tang B, Hu CJ, Xie R, Yang SM, Li BS. Long noncoding RNA in digestive tract cancers: function, mechanism, and potential biomarker. Oncologist. 2015; 20: 898-906. doi: 10.1634/theoncologist.2014-0475.
21. Hajjari M, Salavaty A. HOTAIR: an oncogenic long non-coding RNA in different cancers. Cancer Biol Med. 2015; 12: 1-9. doi: 10.7497/j.issn.2095-3941.2015.0006.
22. Bhan A, Mandal SS. LncRNA HOTAIR: a master regulator of chromatin dynamics and cancer. Biochim Biophys Acta. 2015; 1856: 151-164. doi: 10.1016/j.bbcan.2015.07.001.
23. Nie Y, Liu X, Qu S, Song E, Zou H, Gong C. Long non-coding RNA HOTAIR is an independent prognostic marker for nasopharyngeal carcinoma progression and survival. Cancer Sci. 2013; 104: 458-464. doi: 10.1111/cas.12092.
24. Lee M, Kim HJ, Kim SW, Park SA, Chun KH, Cho NH, Song YS, Kim YT. The long non-coding RNA HOTAIR increases tumour growth and invasion in cervical cancer by targeting the notch pathway. Oncotarget. 2016; 7: 44558-44571. doi: 10.18632/oncotarget.10065.
25. Luo ZF, Zhao D, Li XQ, Cui YX, Ma N, Lu CX, Liu MY, Zhou Y. Clinical significance of HOTAIR expression in colon cancer. World J Gastroenterol. 2016; 22: 5254-5259. doi: 10.3748/wjg.v22.i22.5254.
26. Zhou X, Chen J, Tang W. 2014. The molecular mechanism of HOTAIR in tumorigenesis, metastasis, and drug resistance. Acta Biochim Biophys Sin (Shanghai). 2014; 46: 1011-1015. doi: 10.1093/ abbs gmu104.
27. Zhang J, Zhang P, Wang L, Piao HL, Ma L. 2014. Long non-coding RNA HOTAIR in carcinogenesis and metastasis. Acta Biochim Biophys Sin (Shanghai). 2014; 46: 1-5. doi: 10.1093/abbs/gmt117.
28. Li D, Feng J, Wu T, Wang Y, Sun Y, Ren J, Liu M. Long intergenic noncoding RNA HOTAIR is overexpressed and regulates PTEN methylation in laryngeal squamous cell carcinoma. Am J Pathol. 2013; 182: 64-70. doi: 10.1016/j.ajpath.2012.08.042.
29. Wu J, Xie H. Expression of long noncoding RNA-HOX transcript antisense intergenic RNA in oral squamous cell carcinoma and effect on cell growth. Tumour Biol. 2015; 36: 8573-8578. doi: 10.1007/s13277-015-3598-5.
30. Wu Y, Zhang L, Zhang L, Wang Y, Li H, Ren X, Wei F, Yu W, Liu T, Wang X, Zhou X, Yu J, Hao X. Long non-coding RNA HOTAIR promotes tumor cell invasion and metastasis by recruiting EZH2 and repressing E-cadherin in oral squamous cell carcinoma. Int J Oncol. 2015; 46: 2586-2594. doi: 10.3892/ijo.2015.2976.
31. Kong L, Zhou X, Wu Y, Wang Y, Chen L, Li P, Liu S, Sun S, Ren Y, Mei M, Wang X, Zhang L. Targeting HOTAIR induces mitochondria related apoptosis and inhibits tumor growth in head and neck squamous cell carcinoma in vitro and in vivo. Curr Mol Med. 2015; 15: 952-960. doi: 10.2174/1566524016666151123112716.
32. Fu WM, Lu YF, Hu BG, Liang WC, Zhu X, Yang HD, Li G, Zhang JF. Long noncoding RNA hotair mediated angiogenesis in nasopharyngeal carcinoma by direct and indirect signaling pathways. Oncotarget. 2016; 7: 4712-4723. doi: 10.18632/oncotarget.6731.
33. Stambolic V, Suzuki A, De La Pompa JL, Brothers GM, Mirtsos C, Sasaki T, Ruland J, Penninger JM, Siderovski DP, Mak TW. Negative regulation of PKB/Akt-dependent cell survival by the tumor suppressor PTEN. Cell. 1998; 95: 29-39. doi: 10.1016/S0092-8674(00)81780-8.
34. Kuo LJ, Hung CS, Chen WY, Chang YJ, Wei PL. Glucose-regulated protein 78 silencing down-regulates vascular endothelial growth factor/vascular endothelial growth factor receptor 2 pathway to suppress human colon cancer tumor growth. J Surg Res. 2013; 185: 264-272. doi: 10.1016/j.jss.2013.05.020.
35. Dong D, Stapleton C, Luo B, Xiong S, Ye W, Zhang Y, Jhaveri N, Zhu G, Ye R, Liu Z, Bruhn KW, Craft N, Groshen S, et al. A critical role for GRP78/BiP in the tumor microenvironment for neovascularization during tumor growth and metastasis. Cancer Res. 2011; 71: 2848-2857. doi: 10.1158/0008-5472.CAN-10-3151.
36. Livingstone C. IGF2 and cancer. Endocr Relat Cancer. 2013; 20: R321-R339. doi: 10.1530/ERC-13-0231.
37. Nordin M, Bergman D, Halje M, Engström W, Ward A. Epigenetic regulation of the Igf2/H19 gene cluster. Cell Prolif. 2014; 47: 189-199. doi: 10.1111/cpr.12106.
38. Kallen AN, Zhou XB, Xu J, Qiao C, Ma J, Yan L, Lu L, Liu C, Yi JS, Zhang H, Min W, Bennett AM, Gregory RI, et al. The imprinted h19 lncrna antagonizes let-7 micrornas. Mol Cell. 2013; 52: 101-112. doi: 10.1016/j.molcel.2013.08.027.
39. Ng A, Tang JP, Goh CH, Hui KM. Regulation of the H19 imprinting gene expression in human nasopharyngeal carcinoma by methylation. Int J Cancer. 2003; 104: 179-187. doi: 10.1002/ijc.10926.
40. Gabory A, Jammes H, Dandolo L. The H19 locus: role of an imprinted non-coding RNA in growth and development. BioEssays. 2010; 32: 473-480. doi: 10.1002/bies.200900170.
41. Barsyte-Lovejoy D, Lau SK, Boutros PC, Khosravi F, Jurisica I, Andrulis IL, Tsao MS, Penn LZ. The c-Myc oncogene directly induces the H19 noncoding RNA by allele-specific binding to potentiate tumorigenesis. Cancer Res. 2006; 66: 5330-5337. doi: 10.1158/0008-5472.CAN-06-0037
42. Rainho CA, Kowalski LP, Rogatto SR. Loss of imprinting and loss of heterozygosity on 11p15. 5 in head and neck squamous cell carcinomas. Head Neck. 2001; 23: 851-859. doi: 10.1002/hed.1124.
43. Coutinho-Camillo CM, Brentani MM, Butugan O, Torloni H, Nagai MA. Relaxation of imprinting of IGFII gene in juvenile nasopharyngeal angiofibromas. Diagn Mol Pathol. 2003; 12: 57-62.
44. El-Naggar AK, Lai S, Tucker SA, Clayman GL, Goepfert H, Hong WK, Huff V. Frequent loss of imprinting at the IGF2 and H19 genes in head and neck squamous carcinoma. Oncogene. 1999; 18: 7063-7069. doi: 10.1038/sj.onc.1203192.
45. Esteves LI, Javaroni AC, Nishimoto IN, Magrin J, Squire JA, Kowalski LP, Rainho CA, Rogatto SR. DNA methylation in the CTCF-binding site I and the expression pattern of the H19 gene: Does positive expression predict poor prognosis in early stage head and neck carcinomas? Mol Carcinog. 2005; 44: 102-110. doi: 10.1002/mc.20126.
46. Wu T, Qu L, He G, Tian L, Li L, Zhou H, Jin Q, Ren J, Wang Y, Wang J, Kan X, Liu M, Shen J, et al. Regulation of laryngeal squamous cell cancer progression by the lncRNAH19/miR-148a-3p/DNMT1axis. Oncotarget. 2016; 7: 11553-11566. doi: 10.18632/oncotarget.7270.
47. Sun L, Li Y, Yang B. Downregulated long non-coding RNA MEG3 in breast cancer regulates proliferation, migration and invasion by depending on p53’s transcriptional activity. Biochem Biophys Res Commun. 2016; 478: 323-329. doi: 10.1016/j.bbrc.2016.05.031.
48. Chen RP, Huang ZL, Liu LX, Xiang MQ, Li GP, Feng JL, Liu B, Wu LF. Involvement of endoplasmic reticulum stress and p53 in lncRNA MEG3-induced human hepatoma HepG2 cell apoptosis. Oncol Rep. 2016; 36: 1649-1657. doi: 10.3892/or.2016.4919.
49. Hu D, Su C, Jiang M, Shen Y, Shi A, Zhao F, Chen R, Shen Z, Bao J, Tang W. Fenofibrate inhibited pancreatic cancer cells proliferation via activation of p53 mediated by upregulation of LncRNA MEG3. Biochem Biophys Res Commun. 2016; 471: 290-295. doi: 10.1016/j.bbrc.2016.01.169.
50. Zhou Y, Zhang X, Klibanski A. MEG3 noncoding RNA: a tumor suppressor. J Mol Endocrinol. 2012; 48: R45-R53. doi: 10.1530/JME-12-0008.
51. Jia LF, Wei SB, Gan YH, Guo Y, Gong K, Mitchelson K, Cheng J, Yu GY. Expression, regulation and roles of miR-26a and MEG3 in tongue squamous cell carcinoma. Int J Cancer. 2014; 135: 2282-2293. doi: 10.1002/ijc.28667.
52. Benetatos L, Vartholomatos G, Hatzimichael E. Meg3 imprinted gene contribution intumorigenesis. Int J Cancer. 2011; 129: 773-779. doi: 10.1002/ijc.26052.
53. Zhou Y, Zhong Y, Wang Y, Zhang X, Batista DL, Gejman R, Ansell PJ, Zhao J, Weng C, Klibanski A. Activation of p53 by MEG3 non-coding RNA. J Biol Chem. 2007; 282: 24731-24742. doi: 10.1074/jbc.M702029200.
54. Sandhu R, Rivenbark AG, Coleman WB. Loss of post-transcriptional regulation of DNMT3b by microRNAs: a possible molecular mechanism for the hypermethylation defect observed in a subset of breast cancer cell lines. Int J Oncol. 2012; 41: 721-732. doi: 10.3892/ijo.2012.1505.
55. Ji P, Diederichs S, Wang W, Böing S, Metzger R, Schneider PM, Tidow N, Brandt B, Buerger H, Bulk E, Thomas M, Berdel WE, Serve H, et al. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene. 2003; 22: 8031-8041. doi: 10.1038/sj.onc.1206928.
56. Gutschner T, Hämmerle M, Diederichs S. MALAT1—a paradigm for long noncoding RNA function in cancer. J Mol Med (Berl). 2013; 91: 791-801. doi: 10.1007/s00109-013-1028-y.
57. Chen Q, Su Y, He X, Zhao W, Wu C, Zhang W, Si X, Dong B, Zhao L, Gao Y, Yang X, Chen J, Lu J, et al. Plasma long non-coding RNA MALAT1 is associated with distant metastasis in patients with epithelial ovarian cancer. Oncol Lett. 2016; 12: 1361-1366. doi: 10.3892/ol.2016.4800.
58. Xia H, Chen Q, Chen Y, Ge X, Leng W, Tang Q, Ren M, Chen L, Yuan D, Zhang Y, Liu M, Gong Q, Bi F. The lncRNA MALAT1 is a novel biomarker for gastric cancer metastasis. Oncotarget. 2016; 7: 56209-56218. doi: 10.18632/oncotarget.10941.
59. Jin C, Yan B, Lu Q, Lin Y, Ma L. Reciprocal regulation of Hsa-miR-1 and long noncoding RNA MALAT1 promotes triple-negative breast cancer development. Tumour Biol. 2016; 37: 7383-7394. doi: 10.1007/s13277-015-4605-6.
60. Gutschner T, Hämmerle M, Eißmann M, Hsu J, Kim Y, Hung G, Revenko A, Arun G, Stentrup M, Gross M, Zörnig M, MacLeod AR, Spector DL, et al. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 2013; 73: 1180-1189. doi: 10.1158/0008-5472.CAN-12-2850.
61. Tripathi V, Ellis JD, Shen Z, Song DY, Pan Q, Watt AT, Freier SM,Bennett CF, Sharma A, Bubulya PA, Blencowe BJ, Prasanth SG, Prasanth KV. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol Cell. 2010; 39: 925-938. doi: 10.1016/j.molcel.2010.08.011.
62. Xie L, Hu Z, Wang X, Li Z. 2013. [Expression of long noncoding RNA MALAT1 gene in human nasopharyngeal carcinoma cell lines and its biological significance]. Nan Fang Yi Ke Da Xue Xue Bao. 2013; 33: 692-697. doi: 10.3969/j.issn.1673-4254.2013.05.13.
63. Feng J, Tian L, Sun Y, Li D, Wu T, Wang Y, Liu M. Expression of long non-coding ribonucleic acid metastasis-associated lung adenocarcinoma transcript-1 is correlated with progress and apoptosis of laryngeal squamous cell carcinoma. Head Neck Oncol. 2012; 4: 46-53.
64. Jin C, Yan B, Lu Q, Lin Y, Ma L. The role of MALAT1/miR-1/slug axis on radioresistance in nasopharyngeal carcinoma. Tumour Biol. 2016; 37: 4025-4033. doi: 10.1007/s13277-015-4227-z.
65. Fang Z, Wu L, Wang L, Yang Y, Meng Y, Yang H. Increased expression of the long non-coding RNA UCA1 in tongue squamous cell carcinomas: a possible correlation with cancer metastasis. Oral Surg Oral Med Oral Pathol Oral Radiol. 2014; 117: 89-95. doi: 10.1016/j.oooo.2013.09.007.
66. Xu ZY, Ding SM, Zhou L, Xie HY, Chen KJ, Zhang W, Xing CY, Guo HJ, Zheng SS. FOXC1 contributes to microvascular invasion in primary hepatocellular carcinoma via regulating epithelial-mesenchymal transition. Int J Biol Sci. 2012; 8: 1130-1141. doi: 10.7150/ijbs.4769.
67. Kong XP, Yao J, Luo W, Feng FK, Ma JT, Ren YP, Wang DL, Bu RF. The expression and functional role of a FOXC1 related mRNA-lncRNA pair in oral squamous cell carcinoma. Mol Cell Biochem. 2014; 394: 177-186. doi: 10.1007/s11010-014-2093-4.
68. Zeng C, Xu Y, Xu L, Yu X, Cheng J, Yang L, Chen S, Li Y. Inhibition of long non-coding RNA NEAT1 impairs myeloid differentiation in acute promyelocytic leukemia cells. BMC Cancer. 2014; 14: 693. doi: 10.1186/1471-2407-14-693.
69. Guo S, Chen W, Luo Y, Ren F, Zhong T, Rong M, Dang Y, Feng Z, Chen G. Clinical implication of long non-coding RNA NEAT1 expression in hepatocellular carcinoma patients. Int J Clin Exp Pathol. 2015; 8: 5395 - 5402.
70. Pan LJ, Zhong TF, Tang RX, Li P, Dang YW, Huang SN, Chen G. Upregulation and clinicopathological significance of long non-coding NEAT1 RNA in NSCLC tissues. Asian Pac J Cancer Prev. 2015; 16: 2851-2855.
71. Wang P, Wu T, Zhou H, Jin Q, He G, Yu H, Xuan L, Wang X, Tian L, Sun Y, Liu M, Qu L. Long noncoding RNA NEAT1 promotes laryngeal squamous cell cancer through regulating miR-107/CDK6 pathway. J Exp Clin Cancer Res. 2016; 35: 22. doi: 10.1186/s13046-016-0297-z.
72. Poomsawat S, Sanguansin S, Punyasingh J, Vejchapipat P, Punyarit P. Expression of cdk6 in head and neck squamous cell carcinoma. Clin Oral Investig. 2016; 20: 57-63. doi: 10.1007/s00784-015-1482-8.
73. Bo H, Gong Z, Zhang W, Li X, Zeng Y, Liao Q, Chen P, Shi L, Lian Y, Jing Y, Tang K, Li Z, Zhou Y, et al. Upregulated long non-coding RNA AFAP1-AS1 expression is associated with progression and poor prognosis of nasopharyngeal carcinoma. Oncotarget. 2015; 6: 20404-20418. doi: 10.18632/oncotarget.4057.
74. Baisden JM, Qian Y, Zot HM, Flynn DC. The actin filament-associated protein AFAP-110 is an adaptor protein that modulates changes in actin filament integrity. Oncogene. 2001; 20: 6435-6447. doi: 10.1038/sj.onc.1204784.
75. Zhuang K, Wu Q, Jin CS, Yuan HJ, Cheng JZ. Long non-coding RNA HNF1A-AS is upregulated and promotes cell proliferation and metastasis in nasopharyngeal carcinoma. Cancer Biomark. 2016; 16: 291-300. doi: 10.3233/CBM-150567.
76. Li L, Gu M, You B, Shi S, Shan Y, Bao L, You Y. Long non-coding RNA ROR promotes proliferation, migration and chemoresistance of nasopharyngeal carcinoma. Cancer Sci. 2016; 107: 1215-1222. doi: 10.1111/cas.12989. doi: 10.1111/cas.12989.
77. Sun Q, Liu H, Li L, Zhang S, Liu K, Liu Y, Yang C. Long noncoding RNA-LET, which is repressed by EZH2, inhibits cell proliferation and induces apoptosis of nasopharyngeal carcinoma cell. Med Oncol. 2015; 32: 226. doi: 10.1007/s12032-015-0673-0.
78. Pan W, Zhou L, Ge M, Zhang B, Yang X, Xiong X, Fu G, Zhang J, Nie X, Li H, Tang X, Wei J, Shao M, et al. Whole exome sequencing identifies lncRNA GAS8-AS1 and LPAR4 as novel papillary thyroid carcinoma driver alternations. Hum Mol Genet. 2016; 25: 1875-1884. doi: 10.1093/hmg/ddw056.
79. Gong Z, Zhang S, Zeng Z, Wu H, Yang Q, Xiong F, Shi L, Yang J, Zhang W, Zhou Y, Zeng Y, Li X, Xiang B, et al. LOC401317, a p53-regulated long non-coding RNA, inhibits cell proliferation and induces apoptosis in the nasopharyngeal carcinoma cell line HNE2. PLOS ONE. 2014; 9: e110674. doi: 10.1371/journal.pone.0110674.
80. Zhou Q, Chen J, Feng J, Wang J. Long noncoding RNA PVT1 modulates thyroid cancer cell proliferation by recruiting EZH2 and regulating thyroid-stimulating hormone receptor (TSHR). Tumour Biol. 2016; 37: 3105-3113. doi: 10.1007/s13277-015-4149-9.
81. Kimura T, Van Keymeulen A, Golstein J, Fusco A, Dumont JE, Roger PP. Regulation of thyroid cell proliferation by TSH and other factors: a critical evaluation of in vitro models. Endocr Rev. 2001; 22: 631-656. doi: 10.1210/edrv.22.5.0444.
82. Yoon H, He H, Nagy R, Davuluri R, Suster S, Schoenberg D, Pellegata N, Chapelle Ade L. Identification of a novel noncoding RNA gene, NAMA, that is downregulated in papillary thyroid carcinoma with braf mutation and associated with growth arrest. Int J Cancer. 2007; 121: 767-775. doi: 10.1002/ijc.22701.
83. He H, Li W, Liyanarachchi S, Jendrzejewski J, Srinivas M, Davuluri RV, Nagy R, de la Chapelle A. Genetic predisposition to papillary thyroid carcinoma: involvement of FOXE1, TSHR, and a novel lincRNA gene, PTCSC2. J Clin Endocrinol Metab. 2015; 100: E164-E172. doi: 10.1210/jc.2014-2147.
84. Jendrzejewski J, He H, Radomska HS, Li W, Tomsic J, Liyanarachchi S, Davuluri RV, Nagy R, de la Chapelle A. The polymorphism rs944289 predisposes to papillary thyroid carcinoma through a large intergenic noncoding RNA gene of tumor suppressor type. Proc Natl Acad Sci U S A. 2012; 109: 8646-8651. doi: 10.1073/pnas.1205654109.
85. Zhang H, Zhao L, Wang YX, Xi M, Liu SL, Luo LL. Long non-coding RNA HOTTIP is correlated with progression and prognosis in tongue squamous cell carcinoma. Tumour Biol. 2015; 36: 8805-8809. doi: 10.1007/s13277-015-3645-2.
86. Zhang W, Wang L, Zheng F, Zou R, Xie C, Guo Q, Hu Q, Chen J, Yang X, Yao H, Song E, Xiang Y. Long noncoding RNA expression signatures of metastatic nasopharyngeal carcinoma and their prognostic value. BioMed Res Int. 2015; 2015: 618924. doi: 10.1155/2015/618924.
87. Jeong S, Lee J, Kim D, Seol MY, Lee WK, Jeong JJ, Nam KH, Jung SG, Shin DY, Lee EJ, Chung WY, Jo YS. Relationship of focally amplified long noncoding on chromosome 1 (FAL1) lncRNA with E2F transcription factors in thyroid cancer. Medicine. 2016; 95: e2592. doi: 10.1097/MD.0000000000002592.
88. Zhou J, Li M, Yu W, Li W, Wang J, Xiang X, Li G, Pan X, Lei D. AB209630, a long non-coding RNA decreased expression in hypopharyngeal squamous cell carcinoma, influences proliferation, invasion, metastasis, and survival. Oncotarget. 2016; 7: 14628-14638. doi: 10.18632/oncotarget.7403.
89. Schwarzenbach H, Hoon DS, Pantel K. Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer. 2011; 11: 426-437. doi: 10.1038/nrc3066.
90. Tang H, Wu Z, Zhang J, Su B. Salivary lncRNA as a potential marker for oral squamous cell carcinoma diagnosis. Mol Med Rep. 2013; 7: 761-766. doi: 10.3892/mmr.2012.1254.
91. Wang J, Zhou Y, Lu J, Sun Y, Xiao H, Liu M, Tian L. Combined detection of serum exosomal miR-21 and HOTAIR as diagnostic and prognostic biomarkers for laryngeal squamous cell carcinoma. Med Oncol. 2014; 31: 148. doi: 10.1007/s12032-014-0148-8.
92. Zhang W, Huang C, Gong Z, Zhao Y, Tang K, Li X, Fan S, Shi L, Li X, Zhang P, Zhou Y, Huang D, Liang F, et al. Expression of LINC00312, a long intergenic non-coding RNA, is negatively correlated with tumor size but positively correlated with lymph node metastasis in nasopharyngeal carcinoma. J Mol Histol. 2013; 44: 545-554. doi: 10.1007/s10735-013-9503-x.
93. Shen Z, Li Q, Deng H, Lu D, Song H, Guo J. Long non-coding RNA profiling in laryngeal squamous cell carcinoma and its clinical significance: potential biomarkers for LSCC. PLOS ONE. 2014; 9: e108237. doi: 10.1371/journal.pone.0108237.
94. Chen H, Xin Y, Zhou L, Huang JM, Tao L, Cheng L, Tian J. Cisplatin and paclitaxel target significant long noncoding RNAs in laryngeal squamous cell carcinoma. Med Oncol. 2014; 31: 246. doi: 10.1007/s12032-014-0246-7.
95. Fayda M, Isin M, Tambas M, Guveli M, Meral R, Altun M, Sahin D, Ozkan G, Sanli Y, Isin H, Ozgur E, Gezer U. Do circulating long non-coding RNAs (lncRNAs)(LincRNA-p21, GAS 5, HOTAIR) predict the treatment response in patients with head and neck cancer treated with chemoradiotherapy? Tumour Biol. 2016; 37: 3969-3978. doi: 10.1007/s13277-015-4189-1.
96. Wang Q, Fan H, Liu Y, Yin Z, Cai H, Liu J, Wang Z, Shao M, Sun X, Diao J, Liu Y, Tong L, Fan Q. Curcumin enhances the radiosensitivity in nasopharyngeal carcinoma cells involving the reversal of differentially expressed long non-coding RNAs. Int J Oncol. 2014; 44: 858-864. doi: 10.3892/ijo.2013.2237.

