INTRODUCTION
The International Agency for Research on Cancer has confirmed that ionizing radiation is associated with an increased risk for a wide range of cancers, including breast cancer, thyroid cancer, and leukemia [1]. The risk for carcinogenesis associated with radiation exposure is influenced by genetic background [2, 3]. Understanding gene–environment interactions in carcinogenesis has been a stated priority for the National Cancer Institute [4].
The ataxia telangiectasia-mutated (ATM) protein plays a central role in mediating the cellular response to radiation-induced DNA damage [5]. Germ-line inactivating mutations in the ATM gene cause ataxia-telangiectasia, a recessive genetic disorder with a high incidence of cancer [6]. Ataxia-telangiectasia heterozygotes appear to have a greater risk of developing cancer than the wild-type homozygotes, leading to the estimation that polymorphisms in the ATM gene may alter the risk of carcinogenesis [7]. In the past two decades, about 100 studies have been published to evaluate the associations of ATM genetic polymorphisms with cancer risk. Some of the polymorphisms have been reported by more than 10 studies, such as rs1801516, IVS10-6T > G, rs1800057, rs1800054, rs1800056, rs1800058, and rs4986761. Although most of the findings on these polymorphisms were inconsistent, a meta-analysis of 11120 participants showed a significant association between the rs1800057 polymorphism and breast cancer risk [8]. Recently, two meta-analyses demonstrated evidence for gene-environment interactions between the ATM gene and radiation exposure in the development of radiotherapy-induced adverse events [9, 10]. Taken together, these suggest a possible role of ATM genetic polymorphisms in carcinogenesis through gene–radiation interactions.
A number of studies have investigated the joint effect between the ATM gene and radiation exposure on cancer risk. The first study published in 2002 showed that polymorphisms in the ATM gene were not associated with an increased breast cancer risk in patients with Hodgkin’s disease after radiotherapy [11]. Subsequently, 5 studies have been conducted on this issue, with inconsistent results [12–16]. Given the uncertainty and the lack of a meta-analysis on this topic, we conducted two meta-analyses of ATM genetic polymorphisms and cancer risk in individuals in the presence or absence of radiation exposure to determine whether there was a joint effect between the ATM gene and radiation exposure in carcinogenesis.
RESULTS
Assessing quality of included studies
rs1801516 was the only ATM genetic polymorphism investigated by more than 3 studies of radiation exposure, and was eligible for the present study. A total of 29 studies were identified for the meta-analysis of individuals without radiation exposure [12, 17–44], and 6 studies for the meta-analysis of individuals with radiation exposure [11–16] (Figure 1). The ATM rs1801516 genotype distribution in controls was not in Hardy–Weinberg equilibrium (HWE) in 5 studies [12, 18–21], could not be assessed in 4 studies [11, 13, 25, 26], and was in HWE for the other studies [17, 22–24, 27–44]. As a result, 5 studies were identified with methodological errors and were excluded from a meta-analysis [12, 18–21]. The quality assessments according to Newcastle–Ottawa scale (NOS) [45] were described in Supplementary Table S1. The included studies had a relatively high quality with a median score of 7, ranging from 5 to 9. The quality was high for 22 studies (≥ 6) [11–16, 25, 26, 28–30, 32–40, 42, 43] and low for 8 studies (≤ 5) [17, 22–24, 27, 31, 41, 44].
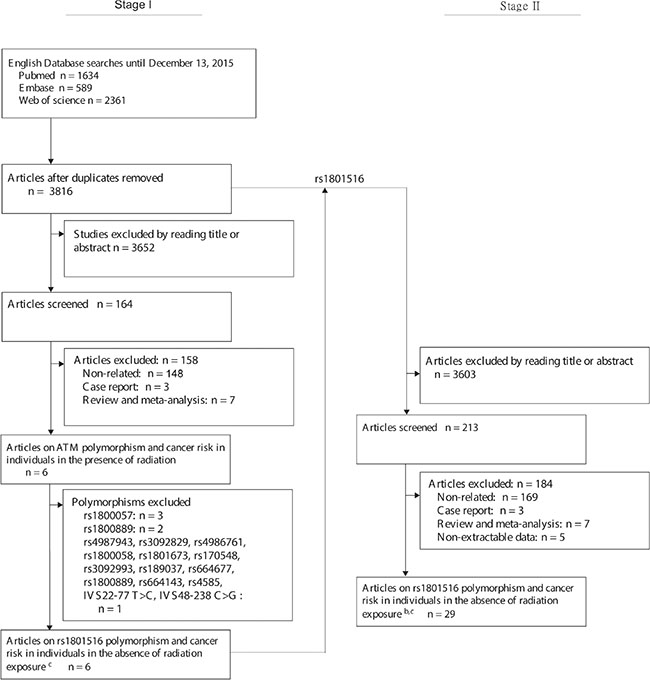
Figure 1: Flow chart for inclusion and exclusion of studies. aThe search on Chinese National Knowledge Infrastructure (CNKI) database identified no study of the ATM rs1801516 polymorphism and cancer risk. b5 studies were identified with methodological errors and were excluded from the present meta-analysis in the subsequent quality assessment procedure [12, 18–21]. cOne article reported data for radiation exposed as well as unexposed populations, the results for each group were considered as a separate study [12].
Meta-analysis for individuals in the absence of radiation exposure
This meta-analysis included 24 studies with 9858 cases and 13475 controls [17, 22–44] (Table 1). When all cancer types were considered, there was no significant association of the rs1801516 polymorphism with cancer risk (homozygous model: odds ratio [OR] = 0.84, 95% confidence interval [CI]: 0.68, 1.03, P = 0.074; heterozygous model: OR = 0.99, 95%CI: 0.91, 1.07, P = 0.784; recessive model: OR = 0.87, 95%CI: 0.69, 1.10, P = 0.231; dominant model: OR = 0.97, 95%CI: 0.89, 1.06, P = 0.632). There was little evidence of heterogeneity across studies (I2 ≤ 29.1%). Subgroup analyses were conducted in order to check whether the features of the included studies affected the results of this meta-analysis. For each genetic model, there was little variation in the effect sizes according to cancer site, ethnicity, study quality, and study size. Figures 2–3 showed the forest plot of the association under the homozygous and dominant models, and Tables 2–3 showed the subgroup analyses under the homozygous and dominant models. The results under the heterozygous and recessive models were similar to those under the dominant and homozygous models, and thus were not shown in figures and tables. For all the meta-analyses, sensitivity analyses did not identify any single study that markedly influenced the estimates, indicating that these results were reliable.
Table 1: Characteristics of studies included in the meta-analysis of individuals in the absence of radiation exposure
First author, year [Ref.] |
Ethnicity |
Region/Country |
Type of cancer |
Family history of cancer |
HWE in controls |
Minor allele frequency |
Cases/controls |
|---|---|---|---|---|---|---|---|
Maillet P, 2000, [44] |
Swiss |
Switzerland |
Colorectal cancer |
Yes |
Yes |
0.14 |
46/163 |
Buchholz TA, 2004, [43] |
Mixed population (75% Caucasian) |
USA |
Breast cancer |
No |
Yes |
0.14 |
58/528 |
Heikkinen K, 2005, [42] |
Finnish |
Finland |
Breast cancer |
Yes |
Yes |
0.25 |
121/306 |
Gonzalez-Hormazabal P, 2008, [41] |
Chilean |
Chile |
Breast cancer |
Yes |
Yes |
0.07 |
126/200 |
Angele S, 2003, [40] |
NR |
France |
Breast cancer |
No |
Yes |
0.13 |
254/312 |
Renwick A, 2006, [39] |
UK ethnic(whites) |
UK |
Breast cancer |
Yes |
Yes |
0.16 |
443/521 |
Angele S, 2004, [38] |
Caucasian |
UK |
Prostate cancer |
No |
Yes |
0.17 |
628/445 |
Yang H, 2007, [37] |
Caucasian |
USA |
Non-small cell lung cancer |
No |
Yes |
> 0.05 |
544/546 |
Tommiska J, 2006, [36] |
Finnish |
Finland |
Breast cancer |
Both |
Yes |
0.24 |
1581/702 |
Wu X, 2006, [35] |
Whites (89.3%) |
USA |
Bladder cancer |
No |
Yes |
0.14 |
608/592 |
Sommer SS, 2002, [34] |
Caucasian (> 80%) |
USA |
Breast cancer |
No |
Yes |
0.13 |
43/43 |
Xu L, 2012, [33] |
Non-hispanic whites; mixed population |
USA |
Thyroid carcinoma |
No |
Yes |
> 0.10 |
592/885 |
Margulis V, 2008, [32] |
NR |
USA |
Renal cancer |
No |
Yes |
0.14 |
323/337 |
Al-Hadyan KS, 2012, [31] |
NR |
Saudi Arabia |
Head and neck cancer |
No |
Yes |
0.07 |
156/251 |
Schrauder M, 2008, [30] |
NR |
German |
Breast cancer |
No |
Yes |
0.15 |
514/511 |
Dork T, 2001, [29] |
Caucasian |
Germany |
Breast cancer |
No |
Yes |
0.13 |
1000/325 |
Wojcicka A, 2014, [28] |
Caucasian |
Poland |
Thyroid cancer |
No |
Yes |
0.11 |
1603/1844 |
Kristensen AT, 2004, [27] |
NR |
Norway |
Rectal cancer |
No |
Yes |
0.17 |
151/3526 |
Hirsch AE, 2008, [26] |
African-American |
USA |
Breast cancer |
No |
NR |
> 0.05 |
37/95 |
Bretsky P, 2003, [25] |
African-American, Latina,Japanese, and Caucasian |
USA |
Breast cancer |
No |
NR |
> 0.03 |
428/426 |
Pereda CM, 2015, [24] |
mixed |
Cuban |
Thyroid cancer |
No |
Yes |
0.11 |
197/206 |
Tecza K, 2015, [23] |
Caucasian |
Poland |
Ovarian cancer |
No |
Yes |
0.13 |
223/335 |
Meier M, 2005, [22] |
Caucasian |
Germany |
T cell acute lymphoblastic leukemia |
No |
Yes |
0.13 |
103/96 |
Oliveira S, 2012, [17] |
Portuguese |
Portugal |
Cervical cancer |
No |
Yes |
0.17 |
79/280 |
Abbreviations: HWE, Hardy–Weinberg equilibrium.
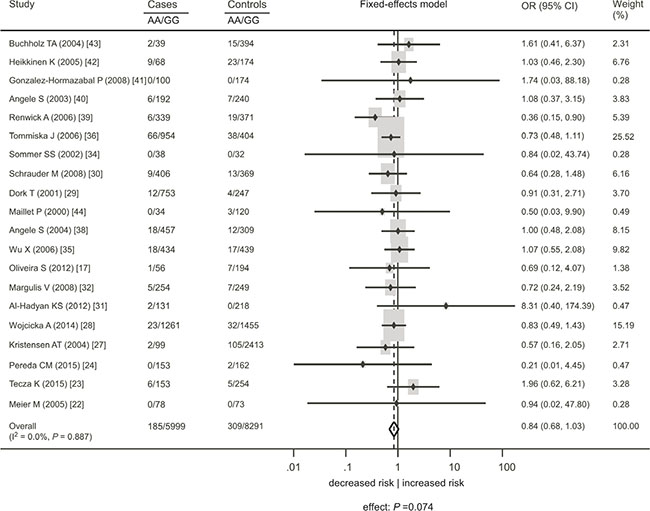
Figure 2: Association between the ATM rs1801516 polymorphism and cancer risk in individuals in the absence of radiation exposure under the homozygous model. AA represents the number of individuals who carry the AA alleles. GG represents the number of individuals who carry the GG alleles. ORs for each study are represented by the squares, and the horizontal line crossing the square represents the 95% CI. The diamond represents the estimated overall effect based on the meta-analysis. ORs and 95%CIs were computed by applying a continuity correction (addition of 0.5 in all the cells) in order to overcome problems resulted from cells containing zero values [69]. All statistical tests were two sided. Abbreviations: CI, confidence interval; OR, odds ratio.
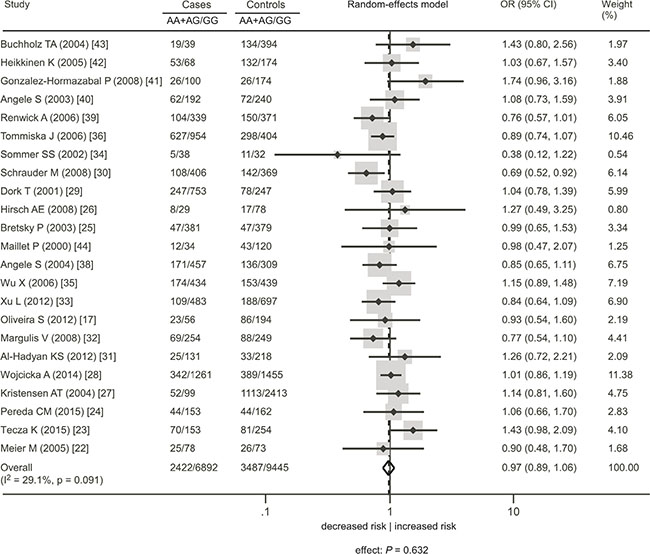
Figure 3: Association between the ATM rs1801516 polymorphism and cancer risk in individuals in the absence of radiation exposure under the dominant model. AA + AG represents the number of individuals who carry the AA or AG alleles. GG represents the number of individuals who carry the GG alleles. ORs for each study are represented by the squares, and the horizontal line crossing the square represents the 95% CI. The diamond represents the estimated overall effect based on the meta-analysis. All statistical tests were two sided. Abbreviations: CI, confidence interval; OR, odds ratio.
Table 2: Subgroup analyses for the association between the ATM rs1801516 polymorphism and cancer risk in individuals in the absence of radiation exposure under the homozygous model
Study selection |
Studies (n) |
Cases |
Controls |
Heterogeneity |
Effect |
||
|---|---|---|---|---|---|---|---|
AA/GG |
AA/GG |
I2 (%) |
P value |
OR (95%CI) |
P value |
||
Quality score |
|||||||
≥ 6 |
12 |
174/5195 |
187/4683 |
0.0 |
0.858 |
0.81 (0.65–1.01) |
0.060 |
≤ 5 |
8 |
11/804 |
122/3608 |
0.0 |
0.617 |
0.99 (0.53–1.83) |
0.976 |
Sample size |
|||||||
Large (> 500) |
12 |
173/5341 |
274/7144 |
0.0 |
0.675 |
0.82 (0.66–1.02) |
0.081 |
Small (< 500) |
8 |
12/658 |
35/1147 |
0.0 |
0.909 |
0.98 (0.52–1.86) |
0.962 |
Family history of casesb |
|||||||
Sporadic cancer |
16 |
137/4989 |
264/7452 |
0.0 |
0.960 |
0.88 (0.70–1.11) |
0.269 |
Family cancer |
5 |
48/1010 |
83/1243 |
4.1 |
0.947 |
0.71 (0.49–1.03) |
0.071 |
Ethnicity |
|||||||
Caucasion |
17 |
183/5615 |
307/7737 |
0.0 |
0.941 |
0.82 (0.67–1.02) |
0.066 |
Site |
|||||||
Breast |
9 |
110/2889 |
119/2404 |
0.0 |
0.704 |
0.76 (0.57–1.01) |
0.060 |
Sum |
20 |
185/5999 |
309/8291 |
0.0 |
0.887 |
0.84 (0.68–1.03) |
0.074 |
AA represents the number of individuals who carry the AA alleles. GG represents the number of individuals who carry the GG alleles. Abbreviations: CI, confidence interval; HWE, Hardy–Weinberg equilibrium; OR, odds ratio. aThe genotype distribution in controls was in HWE in all the studies. b The study by Tommiska et al. [36] reported the risks of both familial and sporadic cancer in comparison with the same controls, and the results for each were considered as a separate study.
Table 3: Subgroup analyses for the association between the ATM rs1801516 polymorphism and cancer risk in individuals in the absence of radiation exposure under the dominant model
Study selection |
Studies (n) |
Cases |
Controls |
Heterogeneity |
Effect |
||
|---|---|---|---|---|---|---|---|
AA + AG/GG |
AA + AG/GG |
I2 (%) |
P value |
OR (95% CI) |
P value |
||
Quality score |
|||||||
≥ 6 |
15 |
2145/6088 |
2035/5837 |
32.4 |
0.103 |
0.92 (0.85–1.00) |
0.054 |
≤ 5 |
8 |
277/804 |
1452/3608 |
0.0 |
0.705 |
1.18 (1.00–1.41) |
0.058 |
Sample size |
|||||||
Large (> 500) |
14 |
2201/6205 |
3069/8220 |
0.052 |
41.5 |
0.95 (0.86–1.05) |
0.325 |
Small (< 500) |
9 |
221/687 |
418/1225 |
0.573 |
0.0 |
1.06 (0.87–1.30) |
0.536 |
Family history of casesa |
|||||||
Familial cancer |
5 |
504/1010 |
649/1243 |
0.169 |
37.8 |
0.91 (0.79–1.06) |
0.214 |
Sporadic cancer |
19 |
1902/5665 |
3115/8428 |
0.170 |
23.6 |
0.97 (0.90–1.04) |
0.352 |
Ethnicity |
|||||||
Caucasian |
19 |
2279/6068 |
3320/8345 |
0.126 |
27.9 |
0.95 (0.88–1.02) |
0.114 |
Site |
|||||||
Breast |
11 |
1306/3299 |
1107/2862 |
0.085 |
39.5 |
0.95 (0.81–1.10) |
0.462 |
Thyroid |
3 |
495/1897 |
621/2314 |
0.304 |
16.1 |
0.96 (0.84–1.10) |
0.571 |
HWE in controls |
|||||||
Yes |
21 |
2367/6482 |
3423/8988 |
0.073 |
32.9 |
0.96 (0.89–1.03) |
0.640 |
Overall |
23 |
2422/6892 |
3487/9445 |
0.114 |
27.1 |
0.97 (0.89–1.06) |
0.632 |
AA + AG represents the number of individuals who carry the AA or AG alleles. GG represents the number of individuals who carry the GG alleles. Abbreviations: CI, confidence interval; HWE, Hardy–Weinberg equilibrium; OR, odds ratio. aThe study by Tommiska et al. [36] reported the risks of both familial and sporadic cancer in comparison with the same controls, and the results for each were considered as a separate study.
We examined if there was evidence of publication bias for each meta-analysis that included 10 or more studies. Asymmetry in the funnel plots was not observed under any comparisons, and significant asymmetry was not suggested by Egger’s linear regression test or Begg’s rank correlation test (Supplementary Figure S1).
Meta-analysis for individuals in the presence of radiation exposure
There were 6 studies with 1459 cases and 2328 controls eligible for this meta-analysis [11–16]. The main characteristics of these studies were presented in Table 4. 2 out of 6 studies investigated the association between the rs1801516 polymorphism and contralateral breast cancer risk in breast cancer patients after radiotherapy [13, 14], 1 study investigated the association between the rs1801516 polymorphism and breast cancer risk in patients with Hodgkin’s disease after radiotherapy [11], and 3 studies investigated the association between the rs1801516 polymorphism and papillary thyroid carcinoma risk in individuals who lived in the areas contaminated by radionuclides [12, 15, 16]. 5 out of 6 studies were conducted in Caucasians [11–15], and 1 in Polynesians [16]. All the included studies had used histologic analyses to confirm cancers.
Table 4: Characteristics of studies included in the meta-analysis of individuals in the presence of radiation exposure
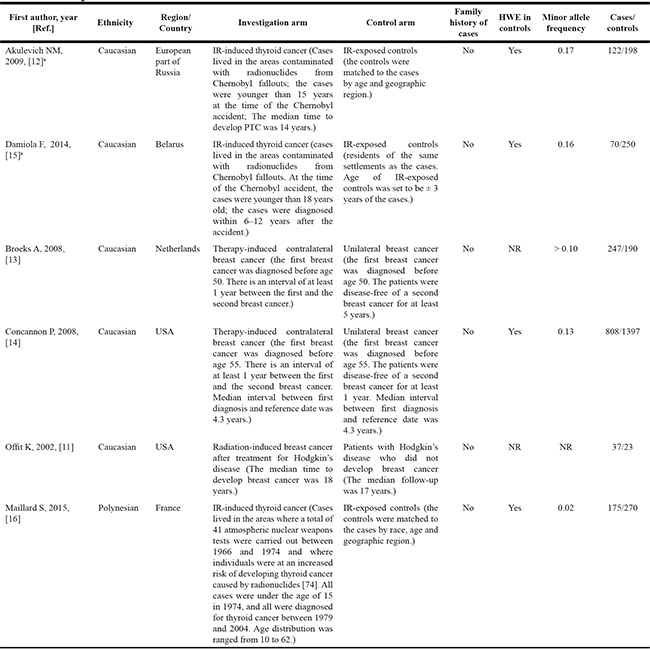
Abbreviations: HWE, Hardy–Weinberg equilibrium; IR, ionizing radiation.
aThere is no overlap in the participants between the two studies [12, 15].
To include all 6 studies for a summary OR estimate, the meta-analysis could only be conducted under the dominant model. The result showed a significant association between the rs1801516 polymorphism and a decreased risk of radiation-induced cancer (OR = 0.64, 95% CI: 0.41, 0.99; P = 0.044), with high between study heterogeneity (I2 = 71.4%, P = 0.004) (Figure 4). Sensitivity analyses identified that the study by Maillard et al. was the outlier, and the association was more significant after this study was excluded (OR = 0.55, 95% CI: 0.36, 0.83; P = 0.005) [16]. However, the heterogeneity remained significant (I2 = 66.9%, P = 0.017), indicating that other factors might contribute to the heterogeneity. Table 5 showed the results of the subgroup analyses. A significant association was shown among Caucasians (OR = 0.55, 95% CI: 0.36, 0.83; P = 0.005), whereas no association was shown among other subgroups. In addition, there was obvious evidence of heterogeneity in all subgroups (I2 ranged 66.9% to 81.8%), suggesting that the examined factors had a minimal influence on the variation of the estimates.
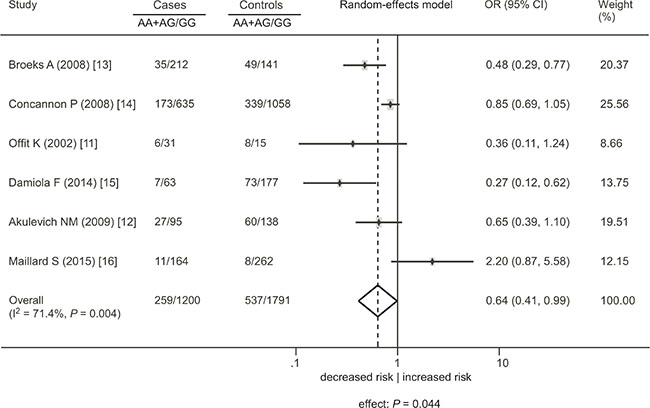
Figure 4: Association between the ATM rs1801516 polymorphism and cancer risk in individuals in the presence of radiation exposure under the dominant model. AA + AG represents the number of individuals who carry the AA or AG alleles. GG represents the number of individuals who carry the GG alleles. ORs for each study are represented by the squares, and the horizontal line crossing the square represents the 95% CI. The diamond represents the estimated overall effect based on the meta-analysis. All statistical tests were two sided. Abbreviations: CI, confidence interval; OR, odds ratio.
Table 5: Subgroup analyses for the association between the ATM rs1801516 polymorphism and cancer risk in individuals in the presence of radiation exposure under the dominant model
Study selection |
Studies (n) |
Cases |
Controls |
Heterogeneity |
Effect |
||
|---|---|---|---|---|---|---|---|
AA + AG/GG |
AA + AG/GG |
I2 (%) |
P value |
OR (95%CI) |
P value |
||
Sample size |
|||||||
Small (< 500) |
4 |
86/565 |
198/733 |
68.1 |
0.014 |
0.58 (0.33–1.03) |
0.065 |
HWE in controls |
|||||||
Yes |
4 |
218/857 |
480/1635 |
75.2 |
0.007 |
0.75 (0.43–1.30) |
0.300 |
Ethnicity |
|||||||
Caucasion |
5 |
248/1036 |
529/1529 |
66.9 |
0.017 |
0.55 (0.36–0.83) |
0.005 |
Site |
|||||||
Breast |
3 |
214/878 |
396/1214 |
67.5 |
0.046 |
0.61 (0.37–1.03) |
0.063 |
Thyroid |
3 |
45/322 |
141/577 |
81.8 |
0.004 |
0.71 (0.26–1.97) |
0.511 |
Sum |
6 |
259/1200 |
537/1791 |
71.4 |
0.004 |
0.64 (0.41–0.99) |
0.044 |
AA + AG represents the number of individuals who carry the AA or AG alleles. GG represents the number of individuals who carry the GG alleles.
Abbreviations: CI, confidence interval; HWE, Hardy–Weinberg equilibrium; OR, odds ratio.
aAll the studies included in these analyses were scored as high quality, and all the participants included were classified as sporadic groups.
Differences in the effect estimates between individuals in the presence or absence of radiation exposure
The effect estimates for individuals in the absence and presence of radiation exposure were compared to determine the relationship of the interaction (synergistic or antagonistic) between radiation exposure and the rs1801516 polymorphism in carcinogenesis. Figure 5 displayed the comparisons of the ORs between the main meta-analyses and between the subgroup analyses under the dominant model. The genetic effect for all participants in the presence of radiation exposure was borderline significantly larger than that for all participants in the absence of radiation exposure (radio of OR = 0.66, 95% CI: 0.42, 1.03; P = 0.066). The difference was statistically significant when only Caucasians were included (radio of OR = 0.58, 95% CI: 0.38, 0.88; P = 0.011).
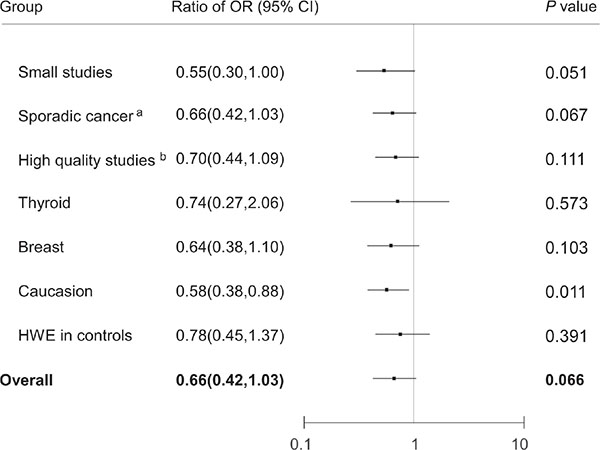
Figure 5: Odds ratios from the meta-analyses of individuals in the presence of radiation exposure were compared with odds ratios from the meta-analyses of individuals in the absence of radiation exposure (dominant model). ORs for each group are represented by the squares, and the horizontal line crossing the square represents the 95% CI. All statistical tests were two sided. Abbreviations: CI, confidence interval; OR, odds ratio. aAll the participants included in the meta-analysis of individuals in the presence of radiation exposure were classified as sporadic groups. bAll the studies included in the meta-analysis of individuals in the presence of radiation exposure were scored as high quality.
DISCUSSION
This work represents the first comprehensive assessment of the literature on the gene-environment interaction for polymorphisms in the ATM gene and radiation exposure in carcinogenesis. rs1801516, which was the only ATM genetic polymorphism investigated by more than 3 studies, was eligible for the present study. Our meta-analyses showed that the rs1801516 polymorphism interacted with radiation exposure, resulting in a synergistic effect in carcinogenesis. In addition, we showed convincing evidence of no association between the rs1801516 polymorphism and cancer risk for individuals in the absence of radiation exposure.
The present meta-analysis of 23333 participants in the absence of radiation exposure had a very large sample size, and was able to provide convincing evidence of no association between the rs1801516 polymorphism and cancer risk. Up to now, 5 meta-analyses have been performed for the role of the rs1801516 polymorphism on cancer risk: 4 on breast cancer [8, 46–48] and 1 on thyroid cancer [49]. One of the meta-analyses showed that homozygous carriers of the rs1801516 genotype had a lower breast cancer risk compared with carriers of the heterozygous and homozygous wild-type genotypes [48]. However, the other studies did not find a significant association between the rs1801516 polymorphism and cancer risk [8, 46, 47, 49]. Compared with the previous meta-analyses [8, 46–49], the present meta-analysis included more studies, and was able to employ rigorous methodology to estimate the genetic effect of the rs1801516 polymorphism on carcinogenesis. The overall meta-analyses of individuals in the absence of radiation exposure showed no association between the rs1801516 polymorphism and cancer risk under the four genetic models. We also conducted subgroup analyses based on cancer site, ethnicity, familial cancer history, study quality, and sample size. For each genetic model, we observed a small variability in the effect sizes between the subgroup analyses and the main meta-analysis. These suggested that the results of the main meta-analysis were independent on the features of the included studies. The extensive consistency provided optimal evidence of the credibility of no association between the rs1801516 polymorphism and cancer risk for individuals in the absence of radiation exposure.
Our meta-analysis of 3787 participants in the presence of radiation exposure provided evidence of an association between the rs1801516 polymorphism and a decreased cancer risk for individuals who exposed to radiation. This meta-analysis included 6 studies across two ethnicities: 1 study in Polynesians and 5 studies in Caucasions. The natures of the two populations are different: the Polynesians are geographically isolated from the rest of the world, and have a significant variation in allele frequencies (minor allele frequency [MAF] in Polynesians = 0.02) as compared to the Caucasians (MAF in Caucasians = 0.19) [16]. The study in Polynesians showed that the minor allele carriers of the rs1801516 polymorphism were associated with an increased cancer risk compared with the main allele carriers in the presence of radiation exposure [16]. On the contrary, all the other studies (Caucasians) showed a consistently decreased cancer risk of the minor allele carriers compared with the main allele carriers in the presence of radiation exposure (2 of 5 comparisons were individually significant [13, 15]). In addition, the test of interaction showed a significant difference in the effect estimates between Caucasions in the presence and absence of radiation exposure. Furthermore, two meta-analyses demonstrated convincing evidence of an association between the rs1801516 polymorphism and radiotherapy-induced adverse events [9, 10] . Taken together, these suggested a gene-environment interaction between the rs1801516 polymorphism and radiation exposure in carcinogenesis, and the interaction might be modified by ethnicity. However, we could not rule out the possibility that the observed association between the rs1801516 polymorphism and cancer risk of Polynesians in the presence of radiation exposure was a chance finding. It should be noted that there was a high variability across studies included in this meta-analysis. Our subgroup analyses failed to explain the heterogeneity, indicating that the study-level factors examined had little influence on the variation of the estimates.
The ATM rs1801516 polymorphism is a polymorphic G-to-A transition at nucleotide 5557 of exon 39, resulting in a change from aspartic acid to asparagine at amino acid position 1853 of the protein [50]. In vitro data showed that human fibroblasts carrying the minor alleles of the rs1801516 polymorphism increased cellular radiosensitivity compared with those carrying the major alleles [51, 52]. Some variants of the ATM gene, including the rs1801516 polymorphism, were reported to be associated with a decreased ATM expression and a reduced capacity of DNA damage recognition [42, 53]. Based on these data, it was difficult to figure out how this single polymorphism might be associated with a decreased cancer risk for individuals who were exposed to radiation. Instead, a gene-gene interaction of the ATM gene with BRCA1 has been reported [28, 52]. Therefore, it could be expected that the polygenic action of unidentified alleles or genes probably played a non-negligible role on the function of the rs1801516 polymorphism. The differences observed between Polynesians and Caucasians regarding the effect of the rs1801516 polymorphism on cancer risk following radiation exposure as well as the clinical heterogeneity were likely to be due to gene-gene interactions.
Our study has a number of possible limitations. 1) Due to fewer than 10 studies in the meta-analysis of individuals with radiation exposure, the publication bias was not tested by the funnel plot, for this method could not obtain enough power in the case [54]. However, based on the Venice criteria that assess cumulative evidence on genetic associations, an OR of > 0.85 or < 1.15 could be easily susceptible to biases, including phenotyping errors, genotyping errors, population stratification, and selective reporting biases [55–57]. This meta-analysis yielded an OR of 0.55, suggesting that this genetic effect was not so vulnerable to biases. 2) Except for the dominant model, other genetic models, such as recessive, heterozygous, and homozygous models, were not examined because of the limited information in the meta-analysis of individuals in the presence of radiation exposure. Therefore, the gene-environment interaction in other genetic models could not be determined. 3) Due to the lack of individual patient data, we were not able to conduct the present meta-analyses based on individual patient data, in which we can: (a) check each study to apply consistent conditions for inclusion and to standardize analysis techniques, and (b) adjust the analyses for covariates (radiation dose, gender, and age). It is especially so for the study by Broeks et al. that reported the significance of ATM variants on secondary breast cancer risk after treatment of primary breast cancer [13]. In this study, 32% patients included in the present meta-analysis did not receive radiotherapy [13]. Because the sensitivity analyses showed no difference in the effect estimates after exclusion of this study, we believed that the incomplete data might reduce the power of the analysis but did not bias it. Moreover, literature based meta-anlayses were considered to be often consistent with those based on individual patient data [58], and should not be viewed as “inferior” [59].
In conclusion, the present study gave a clear picture of gene-environment interaction for the ATM rs1801516 genotype and radiation exposure in carcinogenesis: there was convincing evidence of no association between the rs1801516 polymorphism and cancer risk of individuals in the absence of radiation exposure; there was evidence of a gene-environment interaction between the rs1801516 polymorphism and radiation exposure in carcinogenesis, and the heterogeneity observed across studies might be due to gender-ethnicity or gene-gene interactions. Further studies are needed to elucidate sources of the heterogeneity.
MATERIALS AND METHODS
Our meta-analyses were conducted according to the Preferred Reporting Items for Systematic Reviews and Meta-Analyses guidelines [60].
Selection criteria
To be eligible for inclusion in our meta-analyses, a study had to meet all the following criteria: (1) it should be a case-control, cross-sectional, or cohort study in humans; (2) it can be published in any language, but it must be a full-text paper in an international peer-reviewed journal before December 31, 2015; (3) there was no restriction on cancer type, but it must report adequate information on genotype frequencies to estimate ORs for the cancer type. Case reports, editorials, meta-analyses, and review articles were excluded.
A systematic literature search was conducted in Electronic databases, including PubMed, Web of Science, EMBASE, and Chinese National Knowledge Infrastructure (including China Doctoral/Master Dissertation Full-text Database, China Academic Journals Full-text Database, Century Journals Project, and China Proceedings of Conference Full-text Database), before December 31, 2015. We used the keywords: “(atm OR ataxia telangiectasia mutated) AND (polymorphism* OR variant* OR mutant* OR genotype*)”, in the searching process. This search yielded 3816 articles.
To achieve adequate statistical power for the meta-analysis on gene-environment interactions in carcinogenesis, eligible polymorphisms were those reported by more than three data sources of radiation exposure. For this purpose, we employed a two-stage screen strategy (Figure 1). First, we collected articles on the association between ATM genetic polymorphisms and cancer risk in individuals in the presence of radiation exposure. After screened by title, abstract, or full text if necessary, we identified 6 articles including 17 polymorphisms. References from the relevant articles or reviews were also searched for additional studies. This search yielded no extra articles. Finally, we found that rs1801516 was the only ATM polymorphism investigated by more than 3 articles. Second, we collected articles on the association between the rs1801516 polymorphism and cancer risk in individuals in the absence of radiation exposure. We included all surrogates of the rs1801516 polymorphism, including rs52821794, rs60879649, rs17503060 (http://www.ncbi.nlm.nih.gov/snp/), and rs4988023 [61]. Our search on Chinese National Knowledge Infrastructure database identified no article on the rs1801516 polymorphism and cancer risk (possibly due to a low MAF of < 0.05 in Asians [25, 62]. If different articles reported on the same sample, only the most complete information was included. If an article included multiple sources or study populations, data were extracted separately if possible. The article by Akulevich et al. studied radiation exposed populations as well as unexposed populations, the results for each group were considered as a separate study [12]. Finally, 29 studies without radiation exposure were identified to meet the inclusion criteria for subsequent quality assessment (Figure 1).
Data collection
Two authors independently extracted data based on a standardized form. The following information was collected from each study: first author, year of publication, country of origin, ethnicity, family history of cases (familial cancer or sporadic cancer), MAF in controls, controls in HWE, cancer site, and number of genotyped cases and controls. Ethnicity was classified as African-American, Amerindian, Asian, Caucasian, or others based on the ethnicity of at least 80% of the study population [63]. When a study did not state the included ethnic groups, we considered the ethnicity of the source population based on the country where the study was performed [63]. When an article reported data for different ethnic groups, the results for each group were considered as a separate study. If it was impossible to separate participants according to ethnicity, the participants were considered as “others”. Study authors were contacted when there was insufficient information. Disagreement was resolved by discussion between authors.
Quality assessment
Two authors independently evaluated the quality of each study, with discrepancies resolved during a consensus meeting. We performed two types of quality assessments. The first one was the assessment of methodological errors. Deviation from HWE in controls is an indication of a genotyping error or selection bias [64, 65], and was considered as a methodological error. Because including studies with methodological errors may lower the quality of evidence in a meta-analysis [66], these studies were excluded. However, it should be noted: (1) in case-only studies, HWE deviations may reflect an association with the disease, rather than poor genotyping [67]; (2) studies with insufficient information to determine whether the controls were in HWE were eligible for a meta-analysis, but the influence of these studies on the pooled result was examined in subgroup analyses. Second, the quality of each study was assessed according to the NOS specific to case-control study [45]. The NOS evaluates the quality of a study in three domains: selection, comparability, and exposure. For each study, a maximum score of 4 is assigned for selection, 2 for comparability, and 3 for exposure. A study is considered low (or high) quality if total NOS score is < 6 (or ≥ 6). Because the NOS score is a continuum, distinction between high and low quality is inevitably arbitrary. Due to the subjective nature, the NOS score was used as a stratification factor in the subgroup analysis to evaluate whether the results of the meta-analysis depended on the quality of the included studies [68].
Procedures of meta-analyses
To clarify whether there was a joint effect between the rs1801516 polymorphism and radiation exposure in carcinogenesis, we performed three steps: 1). meta-analysis of the rs1801516 polymorphism and cancer risk in individuals in the presence of radiation exposure; 2). meta-analysis of the rs1801516 polymorphism and cancer risk in individuals in the absence of radiation exposure; 3). comparison of the differences in the effect estimates of the rs1801516 polymorphism on cancer risk between the two groups.
Subgroup meta-analyses were conducted based on pre-specified interests, including cancer site, ethnicity, familial cancer history, study quality, sample size, and HWE in controls. The criteria for a subgroup analysis required at least 3 studies. We aimed at determining whether the result of the overall meta-analysis was stable or dependent on some features of the included studies. Sensitivity analysis was conducted by excluding 1 study at a time and analyzing the remaining ones to explore whether the result was influenced by a particular study.
Statistical analysis
ORs and 95% CIs were used to assess the strength of the association between cancer risk and the rs1801516 polymorphism. The ORs were calculated under four genetic models: (1) heterozygous model (AG versus GG), (2) homozygous model (AA versus GG), (3) dominant model (AA+AG versus GG), and (4) recessive model (AA versus AG+GG). The statistical significance of the ORs was evaluated by using the Z test. In case of zero cells, an appropriate continuity correction (addition of 0.5 in all the cells) was implemented [69]. Between-study heterogeneity was evaluated by using the Cochrane Q test and the I2 statistic. We used the random effects model (DerSimonian and Laird’s method [70]) to calculate the ORs when the P value of the Cochrane Q test was < 0.10 or the I2 value was > 50%; otherwise, the fixed effects model was applied. The test of interaction proposed by Altman et al. [71] was used to compare differences in effect estimates between subgroups. When there were more than 10 studies in a meta-analysis, we estimated publication bias by visualizing funnel plots and by Egger’s linear regression test [72] and Begg’s rank correlation test [73]. To assess deviation from HWE, we performed the appropriate goodness-of-fit χ2 test. The above statistical analyses were performed by using Stata, version 12, software (StataCorp LP, College Station, Texas) with 2-sided P values. Statistical significance was defined as P < 0.05.
CONFLICTS OF INTEREST
The authors have no conflicts of interest to disclose.
GRANT SUPPORT
This work was supported by the General Program of National Natural Science Foundation of China (Grant No. 81673090 to FJH), Natural Science Foundation of Science and Technology Agency of Jilin Province (Bethune special foundation), China (Grant No. 20160101098JC to FJH), Youth Scientific Project from National Science Foundation of China (Grant No. 81300724 to YGZ), Doctoral Program for New Teachers of China’s Ministry of Education (Grant No. 20120061120087 to YGZ). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
REFERENCES
1. Cogliano VJ, Baan R, Straif K, Grosse Y, Lauby-Secretan B, El Ghissassi F, Bouvard V, Benbrahim-Tallaa L, Guha N, Freeman C, Galichet L, Wild CP. Preventable exposures associated with human cancers. J Natl Cancer Inst. 2011; 103:1827–1839.
2. Brennan P. Gene-environment interaction and aetiology of cancer: what does it mean and how can we measure it? Carcinogenesis. 2002; 23:381–387.
3. Rothman N, Wacholder S, Caporaso NE, Garcia-Closas M, Buetow K, Fraumeni JF, Jr. The use of common genetic polymorphisms to enhance the epidemiologic study of environmental carcinogens. Biochim Biophys Acta. 2001; 1471:C1–10.
4. Ghazarian AA, Simonds NI, Bennett K, Pimentel CB, Ellison GL, Gillanders EM, Schully SD, Mechanic LE. A review of NCI’s extramural grant portfolio: identifying opportunities for future research in genes and environment in cancer. Cancer Epidemiol Biomarkers Prev. 2013; 22:501–507.
5. Khanna KK, Lavin MF, Jackson SP, Mulhern TD. ATM, a central controller of cellular responses to DNA damage. Cell Death Differ. 2001; 8:1052–1065.
6. Savitsky K, Bar-Shira A, Gilad S, Rotman G, Ziv Y, Vanagaite L, Tagle DA, Smith S, Uziel T, Sfez S, Ashkenazi M, Pecker I, Frydman M, et al. A single ataxia telangiectasia gene with a product similar to PI-3 kinase. Science. 1995; 268:1749–1753.
7. Athma P, Rappaport R, Swift M. Molecular genotyping shows that ataxia-telangiectasia heterozygotes are predisposed to breast cancer. Cancer Genet Cytogenet. 1996; 92:130–134.
8. Zhang B, Beeghly-Fadiel A, Long J, Zheng W. Genetic variants associated with breast-cancer risk: comprehensive research synopsis, meta-analysis, and epidemiological evidence. Lancet Oncol. 2011; 12:477–488.
9. Dong L, Cui J, Tang F, Cong X, Han F. Ataxia telangiectasia-mutated gene polymorphisms and acute normal tissue injuries in cancer patients after radiation therapy: a systematic review and meta-analysis. Int J Radiat Oncol Biol Phys. 2015; 91:1090–1098.
10. Zhang Y, Liu Z, Wang M, Tian H, Su K, Cui J, Dong L, Han F. Single Nucleotide Polymorphism rs1801516 in Ataxia Telangiectasia-Mutated Gene Predicts Late Fibrosis in Cancer Patients After Radiotherapy: A PRISMA-Compliant Systematic Review and Meta-Analysis. Medicine (Baltimore). 2016; 95:e3267.
11. Offit K, Gilad S, Paglin S, Kolachana P, Roisman LC, Nafa K, Yeugelewitz V, Gonzales M, Robson M, McDermott D, Pierce HH, Kauff ND, Einat P, et al. Rare variants of ATM and risk for Hodgkin’s disease and radiation-associated breast cancers. Clin Cancer Res. 2002; 8:3813–3819.
12. Akulevich NM, Saenko VA, Rogounovitch TI, Drozd VM, Lushnikov EF, Ivanov VK, Mitsutake N, Kominami R, Yamashita S. Polymorphisms of DNA damage response genes in radiation-related and sporadic papillary thyroid carcinoma. Endocr Relat Cancer. 2009; 16:491–503.
13. Broeks A, Braaf LM, Huseinovic A, Schmidt MK, Russell NS, van Leeuwen FE, Hogervorst FB, Van ‘t Veer LJ. The spectrum of ATM missense variants and their contribution to contralateral breast cancer. Breast Cancer Res Treat. 2008; 107:243–248.
14. Concannon P, Haile RW, Borresen-Dale AL, Rosenstein BS, Gatti RA, Teraoka SN, Diep TA, Jansen L, Atencio DP, Langholz B, Capanu M, Liang X, Begg CB, et al. Variants in the ATM gene associated with a reduced risk of contralateral breast cancer. Cancer Res. 2008; 68:6486–6491.
15. Damiola F, Byrnes G, Moissonnier M, Pertesi M, Deltour I, Fillon A, Le Calvez-Kelm F, Tenet V, McKay-Chopin S, McKay JD, Malakhova I, Masyakin V, Cardis E, et al. Contribution of ATM and FOXE1 (TTF2) to risk of papillary thyroid carcinoma in Belarusian children exposed to radiation. Int J Cancer. 2014; 134:1659–1668.
16. Maillard S, Damiola F, Clero E, Pertesi M, Robinot N, Rachedi F, Boissin JL, Sebbag J, Shan L, Bost-Bezeaud F, Petitdidier P, Doyon F, Xhaard C, et al. Common variants at 9q22.33, 14q13.3, and ATM loci, and risk of differentiated thyroid cancer in the French Polynesian population. PLoS One. 2015; 10:e0123700.
17. Oliveira S, Ribeiro J, Sousa H, Pinto D, Baldaque I, Medeiros R. Genetic polymorphisms and cervical cancer development: ATM G5557A and p53bp1 C1236G. Oncol Rep. 2012; 27:1188–1192.
18. Calderon-Zuniga Fdel C, Ocampo-Gomez G, Lopez-Marquez FC, Recio-Vega R, Serrano-Gallardo LB, Ruiz-Flores P. ATM polymorphisms IVS24-9delT, IVS38-8T > C, and 5557G > A in Mexican women with familial and/or early-onset breast cancer. Salud Publica Mex. 2014; 56: 206–212.
19. Tapia T, Sanchez A, Vallejos M, Alvarez C, Moraga M, Smalley S, Camus M, Alvarez M, Carvallo P. ATM allelic variants associated to hereditary breast cancer in 94 Chilean women: susceptibility or ethnic influences? Breast Cancer Res Treat. 2008; 107:281–288.
20. Pena-Chilet M, Blanquer-Maceiras M, Ibarrola-Villava M, Martinez-Cadenas C, Martin-Gonzalez M, Gomez-Fernandez C, Mayor M, Aviles JA, Lluch A, Ribas G. Genetic variants in PARP1 (rs3219090) and IRF4 (rs12203592) genes associated with melanoma susceptibility in a Spanish population. BMC Cancer. 2013; 13:160.
21. Li D, Suzuki H, Liu B, Morris J, Liu J, Okazaki T, Li Y, Chang P, Abbruzzese JL. DNA repair gene polymorphisms and risk of pancreatic cancer. Clin Cancer Res. 2009; 15:740–746.
22. Meier M, den Boer ML, Hall AG, Irving JA, Passier M, Minto L, van Wering ER, Janka-Schaub GE, Pieters R. Relation between genetic variants of the ataxia telangiectasia-mutated (ATM) gene, drug resistance, clinical outcome and predisposition to childhood T-lineage acute lymphoblastic leukaemia. Leukemia. 2005; 19:1887–1895.
23. Tecza K, Pamula-Pilat J, Kolosza Z, Radlak N, Grzybowska E. Genetic polymorphisms and gene-dosage effect in ovarian cancer risk and response to paclitaxel/cisplatin chemotherapy. J Exp Clin Cancer Res. 2015; 34:2.
24. Pereda CM, Lesueur F, Pertesi M, Robinot N, Lence-Anta JJ, Turcios S, Velasco M, Chappe M, Infante I, Bustillo M, Garcia A, Clero E, Xhaard C, et al. Common variants at the 9q22.33, 14q13.3 and ATM loci, and risk of differentiated thyroid cancer in the Cuban population. BMC Genet. 2015; 16:22.
25. Bretsky P, Haiman CA, Gilad S, Yahalom J, Grossman A, Paglin S, Van Den Berg D, Kolonel LN, Skaliter R, Henderson BE. The relationship between twenty missense ATM variants and breast cancer risk: the Multiethnic Cohort. Cancer Epidemiol Biomarkers Prev. 2003; 12:733–738.
26. Hirsch AE, Atencio DP, Rosenstein BS. Screening for ATM sequence alterations in African-American women diagnosed with breast cancer. Breast Cancer Res Treat. 2008; 107:139–144.
27. Kristensen AT, Bjorheim J, Wiig J, Giercksky KE, Ekstrom PO. DNA variants in the ATM gene are not associated with sporadic rectal cancer in a Norwegian population-based study. Int J Colorectal Dis. 2004; 19:49–54.
28. Wojcicka A, Czetwertynska M, Swierniak M, Dlugosinska J, Maciag M, Czajka A, Dymecka K, Kubiak A, Kot A, Ploski R, de la Chapelle A, Jazdzewski K. Variants in the ATM-CHEK2-BRCA1 axis determine genetic predisposition and clinical presentation of papillary thyroid carcinoma. Genes Chromosomes Cancer. 2014; 53:516–523.
29. Dork T, Bendix R, Bremer M, Rades D, Klopper K, Nicke M, Skawran B, Hector A, Yamini P, Steinmann D, Weise S, Stuhrmann M, Karstens JH. Spectrum of ATM gene mutations in a hospital-based series of unselected breast cancer patients. Cancer Res. 2001; 61:7608–7615.
30. Schrauder M, Frank S, Strissel PL, Lux MP, Bani MR, Rauh C, Sieber CC, Heusinger K, Hartmann A, Schulz-Wendtland R, Strick R, Beckmann MW, Fasching PA. Single nucleotide polymorphism D1853N of the ATM gene may alter the risk for breast cancer. J Cancer Res Clin Oncol. 2008; 134:873–882.
31. Al-Hadyan KS, Al-Harbi NM, Al-Qahtani SS, Alsbeih GA. Involvement of single-nucleotide polymorphisms in predisposition to head and neck cancer in Saudi Arabia. Genet Test Mol Biomarkers. 2012; 16:95–101.
32. Margulis V, Lin J, Yang H, Wang W, Wood CG, Wu X. Genetic susceptibility to renal cell carcinoma: the role of DNA double-strand break repair pathway. Cancer Epidemiol Biomarkers Prev. 2008; 16:2366–2373.
33. Xu L, Morari EC, Wei Q, Sturgis EM, Ward LS. Functional variations in the ATM gene and susceptibility to differentiated thyroid carcinoma. J Clin Endocrinol Metab. 2012; 97:1913–1921.
34. Sommer SS, Buzin CH, Jung M, Zheng J, Liu Q, Jeong SJ, Moulds J, Nguyen VQ, Feng J, Bennett WP, Dritschilo A. Elevated frequency of ATM gene missense mutations in breast cancer relative to ethnically matched controls. Cancer Genet Cytogenet. 2002; 134:25–32.
35. Wu X, Gu J, Grossman HB, Amos CI, Etzel C, Huang M, Zhang Q, Millikan RE, Lerner S, Dinney CP, Spitz MR. Bladder cancer predisposition: a multigenic approach to DNA-repair and cell-cycle-control genes. Am J Hum Genet. 2006; 78:464–479.
36. Tommiska J, Jansen L, Kilpivaara O, Edvardsen H, Kristensen V, Tamminen A, Aittomaki K, Blomqvist C, Borresen-Dale AL, Nevanlinna H. ATM variants and cancer risk in breast cancer patients from Southern Finland. BMC Cancer. 2006; 6:209.
37. Yang H, Spitz MR, Stewart DJ, Lu C, Gorlov IP, Wu X. ATM sequence variants associate with susceptibility to non-small cell lung cancer. Int J Cancer. 2007; 121:2254–2259.
38. Angele S, Falconer A, Edwards SM, Dork T, Bremer M, Moullan N, Chapot B, Muir K, Houlston R, Norman AR, Bullock S, Hope Q, Meitz J, et al. ATM polymorphisms as risk factors for prostate cancer development. Br J Cancer. 2004; 91:783–787.
39. Renwick A, Thompson D, Seal S, Kelly P, Chagtai T, Ahmed M, North B, Jayatilake H, Barfoot R, Spanova K, McGuffog L, Evans DG, Eccles D, et al. ATM mutations that cause ataxia-telangiectasia are breast cancer susceptibility alleles. Nat Genet. 2006; 38:873–875.
40. Angele S, Romestaing P, Moullan N, Vuillaume M, Chapot B, Friesen M, Jongmans W, Cox DG, Pisani P, Gerard JP, Hall J. ATM haplotypes and cellular response to DNA damage: association with breast cancer risk and clinical radiosensitivity. Cancer Res. 2003; 63:8717–8725.
41. Gonzalez-Hormazabal P, Bravo T, Blanco R, Valenzuela CY, Gomez F, Waugh E, Peralta O, Ortuzar W, Reyes JM, Jara L. Association of common ATM variants with familial breast cancer in a South American population. BMC Cancer. 2008; 8:117.
42. Heikkinen K, Rapakko K, Karppinen SM, Erkko H, Nieminen P, Winqvist R. Association of common ATM polymorphism with bilateral breast cancer. Int J Cancer. 2005; 116:69–72.
43. Buchholz TA, Weil MM, Ashorn CL, Strom EA, Sigurdson A, Bondy M, Chakraborty R, Cox JD, McNeese MD, Story MD. A Ser49Cys variant in the ataxia telangiectasia, mutated, gene that is more common in patients with breast carcinoma compared with population controls. Cancer. 2004; 100:1345–1351.
44. Maillet P, Chappuis PO, Vaudan G, Dobbie Z, Muller H, Hutter P, Sappino AP. A polymorphism in the ATM gene modulates the penetrance of hereditary non-polyposis colorectal cancer. Int J Cancer. 2000; 88:928–931.
45. Wells GA, Shea B, O’Connell D, Peterson J, Welch V, Losos M, Tugwell P. The Newcastle–Ottawa Scale (NOS) for Assessing the Quality of Nonrandomised Studies in Meta-Analyses. Interdiscip Perspect Infect Dis. 2014:2014:625670. http://www.ohri.ca/programs/clinical_epidemiology/oxford.htm.
46. Mao C, Chung VC, He BF, Luo RC, Tang JL. Association between ATM 5557G > A polymorphism and breast cancer risk: a meta-analysis. Mol Biol Rep. 2012; 39:1113–1118.
47. Gao LB, Pan XM, Sun H, Wang X, Rao L, Li LJ, Liang WB, Lv ML, Yang WZ, Zhang L. The association between ATM D1853N polymorphism and breast cancer susceptibility: a meta-analysis. J Exp Clin Cancer Res. 2010; 29:117.
48. Lu PH, Wei MX, Si SP, Liu X, Shen W, Tao GQ, Chen MB. Association between polymorphisms of the ataxia telangiectasia mutated gene and breast cancer risk: evidence from the current studies. Breast Cancer Res Treat. 2011; 126:141–148.
49. Kang J, Deng XZ, Fan YB, Wu B. Relationships of FOXE1 and ATM genetic polymorphisms with papillary thyroid carcinoma risk: a meta-analysis. Tumour Biol. 2014.
50. Andreassen CN, Overgaard J, Alsner J, Overgaard M, Herskind C, Cesaretti JA, Atencio DP, Green S, Formenti SC, Stock RG, Rosenstein BS. ATM sequence variants and risk of radiation-induced subcutaneous fibrosis after postmastectomy radiotherapy. Int J Radiat Oncol Biol Phys. 2006; 64:776–783.
51. Alsbeih G, El-Sebaie M, Al-Harbi N, Al-Buhairi M, Al-Hadyan K, Al-Rajhi N. Radiosensitivity of human fibroblasts is associated with amino acid substitution variants in susceptible genes and correlates with the number of risk alleles. Int J Radiat Oncol Biol Phys. 2007; 68:229–235.
52. Alsbeih G, Al-Meer RS, Al-Harbi N, Bin Judia S, Al-Buhairi M, Venturina NQ, Moftah B. Gender bias in individual radiosensitivity and the association with genetic polymorphic variations. Radiother Oncol. 2016; 119:236–243.
53. West CM, Elyan SA, Berry P, Cowan R, Scott D. A comparison of the radiosensitivity of lymphocytes from normal donors, cancer patients, individuals with ataxia-telangiectasia (A-T) and A-T heterozygotes. Int J Radiat Biol. 1995; 68:197–203.
54. Macaskill P, Walter SD, Irwig L. A comparison of methods to detect publication bias in meta-analysis. Stat Med. 2001; 20:641–654.
55. Vineis P, Manuguerra M, Kavvoura FK, Guarrera S, Allione A, Rosa F, Di Gregorio A, Polidoro S, Saletta F, Ioannidis JP, Matullo G. A field synopsis on low-penetrance variants in DNA repair genes and cancer susceptibility. J Natl Cancer Inst. 2009; 101:24–36.
56. Chatzinasiou F, Lill CM, Kypreou K, Stefanaki I, Nicolaou V, Spyrou G, Evangelou E, Roehr JT, Kodela E, Katsambas A, Tsao H, Ioannidis JP, Bertram L, et al. Comprehensive field synopsis and systematic meta-analyses of genetic association studies in cutaneous melanoma. J Natl Cancer Inst. 2011; 103:1227–1235.
57. Ioannidis JP, Boffetta P, Little J, O’Brien TR, Uitterlinden AG, Vineis P, Balding DJ, Chokkalingam A, Dolan SM, Flanders WD, Higgins JP, McCarthy MI, McDermott DH, et al. Assessment of cumulative evidence on genetic associations: interim guidelines. Int J Epidemiol. 2008; 37:120–132.
58. Steinberg KK, Smith SJ, Stroup DF, Olkin I, Lee NC, Williamson GD, Thacker SB. Comparison of effect estimates from a meta-analysis of summary data from published studies and from a meta-analysis using individual patient data for ovarian cancer studies. Am J Epidemiol. 1997; 145:917–925.
59. Huncharek M, Kupelnick B. In regards to Baujat et al. Chemotherapy in locally advanced nasopharyngeal carcinoma: An individual patient data meta-analysis of eight randomized trials and 1753 patients (Int J Radiat Oncol Biol Phys 2006;64:47–56). Int J Radiat Oncol Biol Phys. 2006; 65:958; author reply 958–959.
60. Liberati A, Altman DG, Tetzlaff J, Mulrow C, Gotzsche PC, Ioannidis JP, Clarke M, Devereaux PJ, Kleijnen J, Moher D. The PRISMA statement for reporting systematic reviews and meta-analyses of studies that evaluate health care interventions: explanation and elaboration. PLoS Med. 2009; 6:e1000100.
61. Barnett GC, Coles CE, Elliott RM, Baynes C, Luccarini C, Conroy D, Wilkinson JS, Tyrer J, Misra V, Platte R, Gulliford SL, Sydes MR, Hall E, et al. Independent validation of genes and polymorphisms reported to be associated with radiation toxicity: a prospective analysis study. Lancet Oncol. 2012; 13:65–77.
62. Suga T, Ishikawa A, Kohda M, Otsuka Y, Yamada S, Yamamoto N, Shibamoto Y, Ogawa Y, Nomura K, Sho K, Omura M, Sekiguchi K, Kikuchi Y, et al. Haplotype-based analysis of genes associated with risk of adverse skin reactions after radiotherapy in breast cancer patients. Int J Radiat Oncol Biol Phys. 2007; 69:685–693.
63. Ioannidis JP, Ntzani EE, Trikalinos TA. ‘Racial’ differences in genetic effects for complex diseases. Nat Genet. 2004; 36:1312–1318.
64. Trikalinos TA, Salanti G, Khoury MJ, Ioannidis JP. Impact of violations and deviations in Hardy-Weinberg equilibrium on postulated gene-disease associations. Am J Epidemiol. 2006; 163:300–309.
65. Hosking L, Lumsden S, Lewis K, Yeo A, McCarthy L, Bansal A, Riley J, Purvis I, Xu CF. Detection of genotyping errors by Hardy-Weinberg equilibrium testing. Eur J Hum Genet. 2004; 12:395–399.
66. Higgins JPT, Green S (editors). Cochrane Handbook for Systematic Reviews of Interventions Version 5.1.0 [updated March 2011]. The Cochrane Collaboration, 2011. Available from http://handbook.cochrane.org.
67. Tapper W, Hammond V, Gerty S, Ennis S, Simmonds P, Collins A, Eccles D. The influence of genetic variation in 30 selected genes on the clinical characteristics of early onset breast cancer. Breast Cancer Res. 2008; 10:R108.
68. Stang A. Critical evaluation of the Newcastle-Ottawa scale for the assessment of the quality of nonrandomized studies in meta-analyses. Eur J Epidemiol. 2010; 25:603–605.
69. Sweeting MJ, Sutton AJ, Lambert PC. What to add to nothing? Use and avoidance of continuity corrections in meta-analysis of sparse data. Stat Med. 2004; 23:1351–1375.
70. DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986; 7:177–188.
71. Altman DG, Bland JM. Interaction revisited: the difference between two estimates. BMJ. 2003; 326:219.
72. Egger M, Davey Smith G, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997; 315:629–634.
73. Begg CB, Mazumdar M. Operating characteristics of a rank correlation test for publication bias. Biometrics. 1994; 50:1088–1101.
74. de Vathaire F, Drozdovitch V, Brindel P, Rachedi F, Boissin JL, et al. Thyroid cancer following nuclear tests in French Polynesia. Br J Cancer. 2010; 103:1115–1121.

