INTRODUCTION
Human cytomegalovirus, also known as human herpesvirus-5 (HHV-5), is a ubiquitous opportunistic herpesvirus that infects between 60% and 70% of adults in developed countries and almost 100% in developing countries. As one of the pathogens, HCMV is able to induce the symptoms characterized by sore throat, prolonged fever and mild hepatitis [1]. Of course, a latent infection without any symptom is a more common situation. HCMV remains life-long infection and can be reactivated at any time, eventually causing significant morbidity and even death, especially in immunocompromised hosts [2]. Currently, increasing evidence has highlighted the relationship between HCMV infection and human cancers. HCMV genomes or gene products have been detected in several cancers including malignant glioma, cervical carcinoma, Kaposi’s sarcoma and breast cancer [3–6]. In vitro studies have verified the capacity of HCMV for transforming cells and increasing tumourigenicity [7]. However, unlike other DNA viruses playing a role in human malignancies by specific viral genes, HCMV often presents no specific and even undetectable DNA sequence in transformed cells, which can be explained by “hit and run” mechanism [8, 9]. In recent years, the oncomodulatory potential of HCMV has been noted according to the data from studies with tumor cell lines infected by HCMV. Oncomodulation allows HCMV to increase malignancy of infected tumor cells by regulating signal pathways, transcription factors and tumor suppressor proteins [10, 11].
Colorectal cancer is globally one of the leading causes of cancer-related mortality. It shows a five-year survival rate less than 60% in Europe [12]. The pathogenesis of CRC has been long focused on environmental factors such as insufficient activity, high-fat diets, smoking and living in a developed country [13]. Recently increasing attention has been paid to the uncertain but significant effect the infection of HCMV as a microorganism exerts on CRC. Since HCMV nucleic acid was first detected in tumor tissues of CRC patients, the similar results had been confirmed by other studies [14]. Some controlled studies have supposed that the prevalence of HCMV DNA or proteins differs significantly between tumor tissues and matched normal tissues [15, 16]. The presence of human cytomegalovirus in tumor specimens is even related to prognosis of patients with CRC [17].
Nevertheless, detection of HCMV in CRC remains controversial. Negative results indicate that HCMV may present lack of association with CRC regardless of type of tissues or detection methods [18]. Inadequate data has been obtained to confirm the causal role of HCMV in carcinogenesis processes, tumor progression and metastasis [19]. Although HCMV infection is considered to have relationship with CRC, the clinical and pathological features of patients are also various [20, 21].
We pooled the original literature concerning detection of HCMV in CRC tissues and performed a meta-analysis to investigate the relationship between HCMV infection and CRC patients and their clinicopathological characteristics.
RESULTS
The literature search resulted in 192 relevant articles after removing the duplicate ones. By carefully screening the title of these articles, 166 were excluded for not focusing on the desired topic. The criteria of inclusion and exclusion further excluded 16 articles (4 reviews, 2 not in English and 10 without detecting DNA by PCR) thus 10 qualified articles were included (Figure 1) [15–17, 19–25]. One of these articles had separately studied two different regions (Sweden and Vietnam) of patients [16], so it was regarded as two individual articles and labelled as “Dimberg, J. 1” and “Dimberg, J. 2” respectively. Finally, the number of all trials involved was 11. The year of publication ranged from 2004 to 2016. Among these studies, 3 studies were performed in European patients and 6 in Asian patients. 1 study investigated patients in USA, while 1 in South America. For all 11 studies, 4 of them brought into analysis matched normal tissues from the same patient with CRC as a control group. Characteristics of all included studies were summarized in Table 1.
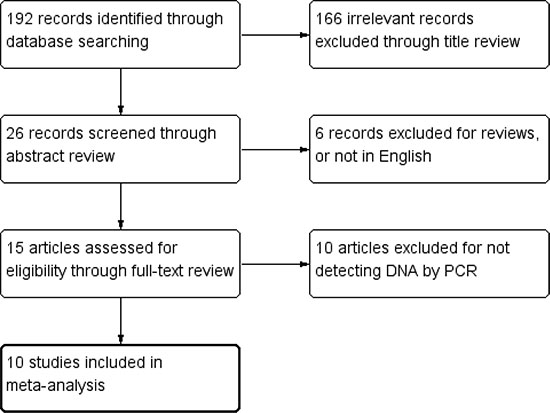
Figure 1: Flow diagram of search strategy.
Table 1: Detailed information of included studies
Study No. |
Publication year |
First author |
Regions |
Type of samples |
No. of cases |
No. of controls |
|---|---|---|---|---|---|---|
1 |
2013 |
Dimberg, J. 1 |
Sweden |
fresh/frozen |
119 |
119 |
2 |
2013 |
Dimberg, J. 2 |
Vietnam |
fresh/frozen |
83 |
83 |
3 |
2014 |
Tafvizi, F. |
Iran |
formalin-fixed |
42 |
- |
4 |
2015 |
Chen, H.P. |
Taiwan |
fresh/frozen |
115 |
115 |
5 |
2012 |
Chen, H.P. |
Taiwan |
fresh/frozen |
163 |
163 |
6 |
2016 |
Chen, H.P. |
Taiwan |
fresh/frozen |
83 |
- |
7 |
2013 |
Chen, H.P. |
Taiwan |
fresh/frozen |
95 |
- |
8 |
2009 |
Bender, C. |
Italy |
formalin-fixed |
36 |
- |
9 |
2008 |
Mariguela, V.C. |
Brazil |
fresh/frozen |
14 |
- |
10 |
2004 |
Knösel, T. |
Germany |
fresh/frozen |
57 |
- |
11 |
2005 |
Akintola-Ogunremi, O. |
USA |
formalin-fixed |
13 |
- |
The prevalence of HCMV infected CRC
Meta-analysis revealed considerable heterogeneity in positive rate of HCMV infection in patients diagnosed with colorectal cancer with the overall prevalence 27.5% (95% CI = 17.2%–37.8%) (Figure 2). The HCMV infection in cases with different features of patients and specimens were present (Table 2). The lowest prevalence of HCMV-infected CRC was in patients of USA, about 3.6% (95% CI = −6.1%–13.3%). And it was much higher in Asia subgroup 42.0% (95% CI = 37.9%−46.0%). Considering Europe and USA as a developed region, Asia and Brazil as a developing region, the former demonstrated a significantly lower positive proportion than the latter (aOR = 0.257, 95% CI = 0.159–0.414). As for the year of publication, the studies reported before 2010 presented a low prevalence of HCMV 7.9% (95% CI = 3.1%−12.7%), which was significantly lower than the studies published after 2010 (aOR = 0.488, 95% CI = 0.237–1.005). Compared to the fresh and frozen specimen, formalin-fixed specimen presented a markedly lower prevalence of HCMV DNA (aOR = 0.451, 95% CI = 0.265–0.766).
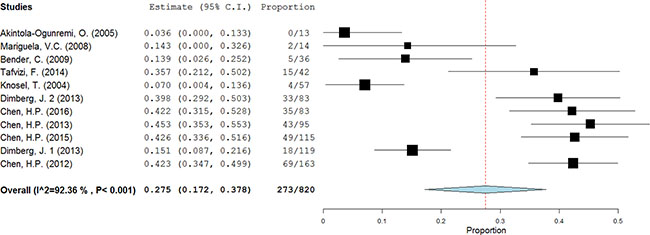
Figure 2: The prevalence of HCMV in CRC tissues. A random-effects model was performed.
Table 2: The comparison of HCMV prevalence in CRC tissues
Category |
Subcategory |
No. of studies |
Prevalence (%) (95% CI) |
Adjusted ORa |
P value |
|---|---|---|---|---|---|
Overall |
Total |
11 |
27.5 (17.2−37.8) |
− |
|
Region |
Developing region |
7 |
40.0 (34.8−45.1) |
Ref |
|
Developed region |
4 |
10.0 (4.6−15.5) |
0.257 (0.159−0.414) |
< 0.01 |
|
Year of publication |
After 2010 |
7 |
37.4 (27.3−47.4) |
Ref |
|
Before 2010 |
4 |
7.9(3.1−12.7) |
0.488 (0.237−1.005) |
< 0.01 |
|
type of samples |
Fresh and frozen |
8 |
31.2 (19.2−43.3) |
Ref |
|
Formalin-fixed |
3 |
17.0 (−0.2−34.2) |
0.451 (0.265−0.766) |
< 0.01 |
95% CI: confidence interval, OR: odds ratio.
a: Adjusted by region, year of publication and type of samples.
HCMV infection and CRC risk
Four articles studying tumor tissues with adjacent normal tissues from the same patients were selected for the meta-analysis. All of the 4 reports provided sufficient data to estimate the OR and 95%CI. The forest plot showed the effect size for each study and overall value (Figure 3). There was no significant heterogeneity across these studies (I2 = 0%, P = 0.71), so a fixed-effects model was used to estimate the pooled OR. The result revealed that the tumor tissues had a significantly higher rate of HCMV infection (OR = 6.59, 95% CI = 4.48–9.69). No evident asymmetry of plots was seen in the funnel plots suggesting no evidence of publication bias (Figure 4).
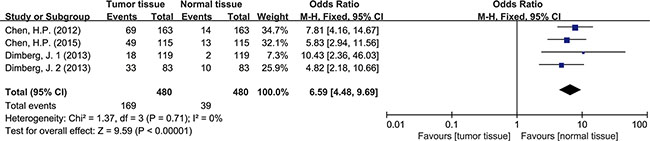
Figure 3: Forest plot of ORs for the association between HCMV infection and colorectal cancer risk.
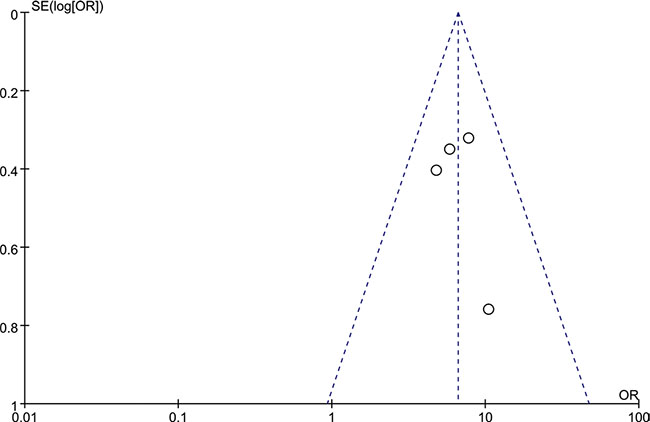
Figure 4: The funnel plots of the included studies. OR, odds ratio; SE, standard error.
HCMV infection in tissues with different features
Meta-analysis was used to investigate more detailed data. Colorectal cancer was classified into 4 stages according to the American Joint Committee on Cancer (AJCC) classification system. There was no clear evidence indicating the difference of HCMV infection between different stages of cancer, such as subgroups of stage I+II and stage III+IV (OR = 0.89, 95% CI = 0.56–1.42) (Figure 5). In addition, HCMV infection was not related to tumor cells with different degree of differentiation. We compared the subgroups of well differentiated cells with moderately differentiated cells, also well with poorly and moderately with poorly. None of them revealed a significant difference in pooled data (OR = 2.00, 95% CI = 0.80–5.01; OR = 2.00, 95%CI = 0.33–11.99; OR = 1.32, 95% CI = 0.32–5.47, respectively) (Figures 6, 7, 8). There were two articles containing the data of HCMV detection in polyps and adenocarcinomas. The HCMV infection didn’t statistically differ in tissues of polyp and adenocarcinoma (OR = 0.19, 95% CI = 0.03–1.10) (Figure 9).
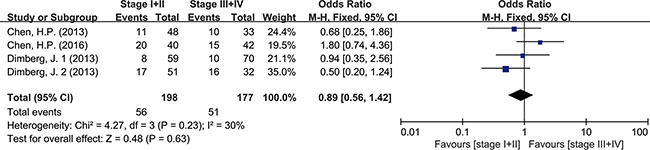
Figure 5: Difference in the tumor staging (stage I+II and stage III+IV).
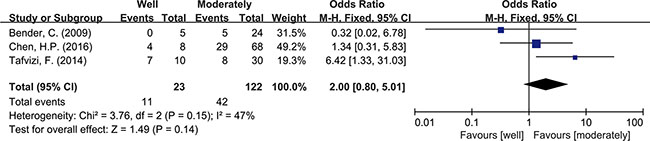
Figure 6: Difference in the grade of cell differentiation (well and moderately).
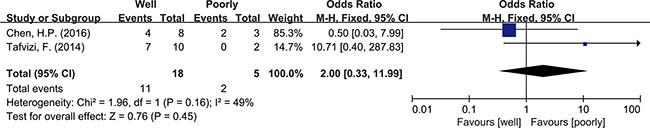
Figure 7: Difference in the grade of cell differentiation (well and poorly).
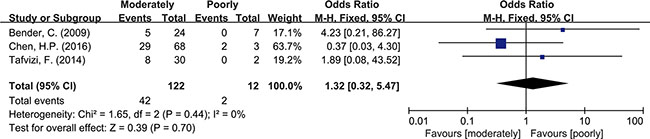
Figure 8: Difference in the grade of cell differentiation (moderately and poorly).
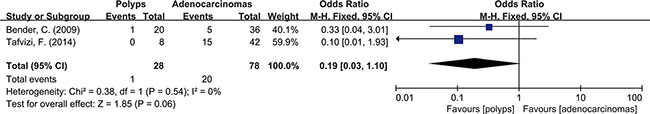
Figure 9: Difference in the tissues of polyp and adenocarcinoma.
DISCUSSION
Over the past few decades, great attention had been paid to the relationship between HCMV infection and colorectal cancer. However the exact role of HCMV in patients with CRC still remained unclear. Among the majority of researches, the rate of HCMV detection in tumor samples represented the rate of HCMV-positive cancer, thus the influence of sample types and detection techniques being unneglectable. One single study with insufficient cases inevitably leaded to bias, which made a meta-analysis necessary.
Viral DNA and gene products including encoded RNAs and proteins were two general targets for viral detection. In situ hybridization (ISH) and immunohistochemistry (IHC) were available for the detection of gene products. However, as a latent virus, HCMV might not consistently express products during the whole period of infection, which would cause false-negative results and affect the accuracy of analysis [26, 27]. The PCR method was an important technique for DNA detection. PCR showed higher positive rate than ICH when a few studies combined both techniques of them. [19, 24] This controversy might result from the different detection techniques. Considering the bias of different methods, we chose studies which used PCR for HCMV DNA detection.
We found that there was a certain prevalence of HCMV DNA (28.7%) in CRC tissues, compared to a high rate of HCMV detection (over 80%) in patients’ plasma [16, 21]. Some reports attributed low or negative detection to the type of sample. Tissues processed by formalin would fail to do a detection of viral nucleic acids because of its effect on fragmentation of DNA. On the contrary, fresh and frozen sample was suitable for DNA preservation and detection [28, 29]. Here the data supported this opinion. DNA detection in fresh and frozen tissue showed more positive results than in formalin-fixed tissue (P < 0.01).
It had been reported that racial and geographic distribution of HCMV infection was existing [30, 31]. Here western and non-western countries shared a distribution of HCMV infection in CRC tissues (aOR = 0.257, 95% CI = 0.159–0.414), which revealed that there was less association between HCMV infection and CRC patients in developed than developing countries. This heterogeneity might be explained by the rate of HCMV infection and use of testing techniques and diagnostic criteria for HCMV infection in different regions. Epidemiological data suggested that sporadic CRC was more common in developed countries. Nevertheless the incidence in developing countries was increasing and it had been reported that the incidence of early onset CRC in developing was much higher [32, 33]. Whether HCMV played a role in these phenomena needed explaining by more data and further research. The prevalence of HCMV in colorectal cancer was significantly lower in the publication period before 2010 than after 2010 (aOR = 0.488, 95% CI = 0.237–1.005). The technical improvements and a larger sample size (700 cases versus 120 cases) might be a reason for the difference and made the results reported after 2010 more convincing. However negative results tended not to be published especially in the condition that negative results had already been published. Although the funnel plots indicated no publication bias, this could be an explanation, too.
Detection of HCMV DNA in CRC tissues by itself didn’t provide adequate evidence to prove the causal role of HCMV in CRC. So data from 4 studies with total 480 tumor tissues and 480 matched normal tissues were pooled to give a more reliable evidence. Significant difference was seen in HCMV DNA prevalence between cancer and non-cancer tissue (OR = 6.59, 95% CI = 4.48–9.69). This observation supported the association of HCMV with colorectal tumor formation. The result refuted the conclusion that HCMV was not related to CRC risk, as some studies had drawn [19, 25]. For further analysis, we also pooled the data of different pathological features of tumor. The results indicated that HCMV seemed not to affect tumor staging and grade of differentiation since the detection of HCMV DNA showed no difference between stage I+II and stage III+IV, also no difference among high, moderate and low differentiations of tumor cell. As known to all, intestinal polyp, a precancerous lesion, could increase the risk of progression to colorectal cancer [34]. Our results showed no difference of HCMV infection in these two kinds of tissues. The results could be explained by one suggestion that HCMV infection might have influence on the every stage of tumor formation, not only on a single part. The detection of HCMV DNA might be a signal for malignant change of colorectal tissues but might not be an independent factor to predict the level of malignancy. However it had been reported that the distribution of HCMV proteins showed dramatic difference in tumor tissues (CRC or glioma) depending on the tumor histology [35, 36]. Viral non-coding RNAs regulatory proteins determined HCMV-induced oncomodulation thus affecting differentiation of tumor cells [37]. A further study about gene products of HCMV needed carrying out to provide a more specific explanation on tumor progression. Chen, et al. also reported that despite of TNM stage, HCMV in tumors were related to shorter disease-free survival in aged individuals with CRC [17]. It reminded us of other mechanisms that the virus had to affect the survival rate of patients.
It was important to realize the potential relationship between HCMV and CRC. This relationship could not only provide a deeper insight into the carcinogenic mechanism of CRC, but also offer a novel therapeutic option [38]. This was the first meta-analysis to investigate the association between HCMV infection and colorectal cancer. Moreover the parameters of geographical region, year of publication, type of samples, tumor staging, and histologic grade were evaluated in our study. These parameters could affect the detection of HCMV in CRC tissues and provide a basic knowledge for further analysis between HCMV or other viruses and CRC.
There were limitations in the present study. Our study focused only on researches related to DNA detection and didn’t analyze the data concerning viral proteins detection. 7 articles were absent of control groups. In addition, none of included studies were random trials.
In conclusion, the present study indicated that a HCMV infection was statistically associated with an increased CRC risk and it played a role in the initiation and progression of CRC. Much further researches with larger scale of samples and more abundant date should be conducted to further confirm these results.
MATERIALS AND METHODS
Searching method and selection criteria
Literatures published up to January 2016 were searched in the electronic database of PubMed, Medline and EMBASE using the keywords “cytomegalovirus”, or “human herpesvirus-5” and “colorectal cancer” or “colorectal carcinoma”. The retrieved articles and their additional relevant references were reviewed meeting the following criteria: (1) Studies had to use polymerase chain reaction (PCR) technique to detect the presence of HCMV DNA in tumor tissues. (2) The articles should be with full text and published in English. In vitro studies based on cell lines or cells and review articles without original data were excluded.
Data extraction
Data was extracted from the selected and qualified articles including the first author’s name, the year of publication, geographic regions. The pathological characteristics of CRC specimen were also collected.
Statistical analysis
The two main parameters analyzed for these studies were prevalence rate and odds ratio (OR). The statistical software OpenMeta-analyst (Center for Evidence-based Medicine, Brown University, Providence, R.I., USA) was used to analyze the proportions of samples positive for HCMV. The data from patient and control groups were pooled using Review Manager 5.3 (Cochrane Collaboration, Nordic Cochrane Centre, Copenhagen, Denmark). The meta-analysis was performed based on Mantel-Haenszel and DerSimonian-Laird methods according to heterogeneity which was tested by the Q statistic (significance level at P < 0.1) and the I2 statistic (significance level at I2 > 50%). If the Q or I2 value was statistically significant, a random-effects model was adopted, otherwise a fixed-effects model was selected. Binary logistic regression was performed to compare and adjust the prevalence of HCMV by following factors: regions, the year of publication, detection techniques and specimen types with the adjusted OR (aOR) as an effect size. P < 0.05 were considered statistically significant.
ACKNOWLEDGMENTS AND FUNDING
This article represents partial fulfillment of the requirements for a Master degree for Bingjun Bai. This work was supported by grants from the National Natural Science Foundation of China (81272681).
CONFLICTS OF INTEREST
All authors listed in the manuscript have concurred with the content of the manuscript and this submission. No conflict of interest was excited between us authors.
REFERENCES
1. Mattes FM, McLaughlin JE, Emery VC, Clark DA, Griffiths PD. Histopathological detection of owl’s eye inclusions is still specific for cytomegalovirus in the era of human herpesviruses 6 and 7. J Clin Pathol. 2000; 53:612–614.
2. Fulop T, Larbi A, Pawelec G. Human t cell aging and the impact of persistent viral infections. Front Immunol 2013; 4:271.
3. Richardson AK, Currie MJ, Robinson BA, Morrin H, Phung Y, Pearson JF, Anderson TP, Potter JD, Walker LC. Cytomegalovirus and epstein-barr virus in breast cancer. Plos One. 2015; 10:e118989.
4. Cobbs CS, Harkins L, Samanta M, Gillespie GY, Bharara S, King PH, Nabors LB, Cobbs CG, Britt WJ. Human cytomegalovirus infection and expression in human malignant glioma. Cancer Res. 2002; 62:3347–3350.
5. Giraldo G, Beth E, Huang ES: Kaposi’s sarcoma and its relationship to cytomegalovirus (cmnv). Iii. Cmv dna and cmv early antigens in kaposi’s sarcoma. Int J Cancer. 1980; 26:23–29.
6. Pacsa AS, Kummerlander L, Pejtsik B, Pali K. Herpesvirus antibodies and antigens in patients with cervical anaplasia and in controls. J Natl Cancer Inst. 1975; 55:775–781.
7. Geder KM, Lausch R, O’Neill F, Rapp F. Oncogenic transformation of human embryo lung cells by human cytomegalovirus. Science. 1976; 192:1134–1137.
8. Galloway DA, McDougall JK. The oncogenic potential of herpes simplex viruses: evidence for a ‘hit-and-run’ mechanism. Nature. 1983; 302:21–24.
9. Cinatl JJ, Cinatl J, Vogel JU, Rabenau H, Kornhuber B, Doerr HW. Modulatory effects of human cytomegalovirus infection on malignant properties of cancer cells. Intervirology. 1996; 39:259–269.
10. Cinatl J, Scholz M, Kotchetkov R, Vogel JU, Doerr HW. Molecular mechanisms of the modulatory effects of hcmv infection in tumor cell biology. Trends Mol Med. 2004; 10:19–23.
11. Michaelis M, Doerr HW, Cinatl J. The story of human cytomegalovirus and cancer: increasing evidence and open questions. Neoplasia. 2009; 11:1–9.
12. Cunningham D, Atkin W, Lenz HJ, Lynch HT, Minsky B, Nordlinger B, Starling N. Colorectal cancer. Lancet. 2010; 375:1030–1047.
13. Watson AJ, Collins PD. Colon cancer: a civilization disorder. Dig Dis. 2011; 29:222–228.
14. Huang ES, Roche JK. Cytomegalovirus d.n.a. And adenocarcinoma of the colon: evidence for latent viral infection. LANCET 1978; 1:957–960.
15. Chen H, Jiang J, Chen C, Chou T, Chen Y, Chang Y, Lin S, Chan C, Yang C, Lin C, Lin J, Cho W, Chan Y. Human cytomegalovirus preferentially infects the neoplastic epithelium of colorectal cancer: a quantitative and histological analysis. J Clin Virol. 2012; 54:240–244.
16. Dimberg J, Hong TT, Skarstedt M, Lofgren S, Zar N, Matussek A. Detection of cytomegalovirus dna in colorectal tissue from swedish and vietnamese patients with colorectal cancer. Anticancer Res. 2013; 33:4947–4950.
17. Chen HP, Jiang JK, Lai PY, Chen CY, Chou TY, Chen YC, Chan CH, Lin SF, Yang CY, Chen CY, Lin CH, Lin JK, Ho DM, et al. Tumoral presence of human cytomegalovirus is associated with shorter disease-free survival in elderly patients with colorectal cancer and higher levels of intratumoral interleukin-17. Clin Microbiol Infect. 2014; 20:664–671.
18. Hart H, Neill WA, Norval M. Lack of association of cytomegalovirus with adenocarcinoma of the colon. Gut. 1982; 23:21–30.
19. Knösel T, Schewe C, Dietel M, Petersen I. Cytomegalovirus is not associated with progression and metastasis of colorectal cancer. Cancer Lett. 2004; 211:243–247.
20. Tafvizi F, Fard ZT. Detection of human cytomegalovirus in patients with colorectal cancer by nested-pcr. Asian Pac J Cancer P. 2014; 15:1453–1457.
21. Chen HP, Jiang JK, Lai PY, Teo WH, Yang CY, Chou TY, Lin CH, Chan YJ. The serologic and viraemic status of human cytomegalovirus infection in patients with colorectal cancer is not correlated with viral replication and transcription in tumours. J Gen Virol. 2015.
22. Chen HP, Jiang JK, Chan CH, Teo WH, Yang CY, Chen YC, Chou TY, Lin CH, Chan YJ. Genetic polymorphisms of the human cytomegalovirus ul144 gene in colorectal cancer and its association with clinical outcome. J Gen Virol. 2015.
23. Bender C, Zipeto D, Bidoia C, Costantini S, Zamo A, Menestrina F, Bertazzoni U. Analysis of colorectal cancers for human cytomegalovirus presence. Infect Agent Cancer 2009; 4:6.
24. Mariguela VC, Chacha SG, Cunha AA, Troncon LE, Zucoloto S, Figueiredo LT. Cytomegalovirus in colorectal cancer and idiopathic ulcerative colitis. Rev Inst Med Trop Sao Paulo 2008; 50:83–87.
25. Akintola-Ogunremi O, Luo Q, He TC, Wang HL: Is cytomegalovirus associated with human colorectal tumorigenesis? Am J Clin Pathol. 2005; 123:244–249.
26. Sandford GR, Schumacher U, Ettinger J, Brune W, Hayward GS, Burns WH, Voigt S. Deletion of the rat cytomegalovirus immediate-early 1 gene results in a virus capable of establishing latency, but with lower levels of acute virus replication and latency that compromise reactivation efficiency. J Gen Virol. 2010; 91:616–621.
27. Penkert RR, Kalejta RF. Nuclear localization of tegument-delivered pp71 in human cytomegalovirus-infected cells is facilitated by one or more factors present in terminally differentiated fibroblasts. J Virol. 2010; 84:9853–9863.
28. Rogers BB, Alpert LC, Hine EA, Buffone GJ. Analysis of dna in fresh and fixed tissue by the polymerase chain reaction. Am J Pathol. 1990; 136:541–548.
29. Koshiba M, Ogawa K, Hamazaki S, Sugiyama T, Ogawa O, Kitajima T. The effect of formalin fixation on dna and the extraction of high-molecular-weight dna from fixed and embedded tissues. Pathol Res Pract. 1993; 189:66–72.
30. Jansen MA, van den Heuvel D, Bouthoorn SH, Jaddoe VW, Hooijkaas H, Raat H, Fraaij PL, van Zelm MC, Moll HA. Determinants of ethnic differences in cytomegalovirus, epstein-barr virus, and herpes simplex virus type 1 seroprevalence in childhood. J Pediatr. 2016; 170:126–134.
31. Lantos PM, Permar SR, Hoffman K, Swamy GK. The excess burden of cytomegalovirus in african american communities: a geospatial analysis. Open Forum Infect Dis. 2015; 2:v180.
32. Merika E, Saif MW, Katz A, Syrigos K, Morse M: Review. Colon cancer vaccines: an update. In Vivo. 2010; 24:607–628.
33. Zahir MN, Azhar EM, Rafiq S, Ghias K, Shabbir-Moosajee M. Clinical features and outcome of sporadic colorectal carcinoma in young patients: a cross-sectional analysis from a developing country. Isrn Oncol. 2014; 2014:461570.
34. Winawer S, Fletcher R, Rex D, Bond J, Burt R, Ferrucci J, Ganiats T, Levin T, Woolf S, Johnson D, Kirk L, Litin S, Simmang C. Colorectal cancer screening and surveillance: clinical guidelines and rationale-update based on new evidence. Gastroenterology0. 2003; 124:544–560.
35. Scheurer ME, Bondy ML, Aldape KD, Albrecht T, El-Zein R. Detection of human cytomegalovirus in different histological types of gliomas. Acta Neuropathol. 2008; 116:79–86.
36. Cai Z, Xu J, Zhou Y, Zheng J, Lin K, Zheng S, Ye M, He Y, Liu C, Xue Z. Human cytomegalovirus-encoded us28 may act as a tumor promoter in colorectal cancer. World J Gastroentero. 2016; 22:2789.
37. Michaelis M, Doerr HW, Cinatl J. The story of human cytomegalovirus and cancer: increasing evidence and open questions. Neoplasia. 2009; 11:1–9.
38. Prins RM, Cloughesy TF, Liau LM. Cytomegalovirus immunity after vaccination with autologous glioblastoma lysate. N Engl J Med. 2008; 359:539–541.

