INTRODUCTION
Head and neck squamous cell carcinomas (HNSCC) are one of the most common cancers in the world and are ranked sixth for men and eleventh for women in cancer frequency [1]. Although the term HNSCC is often used to describe tumours in different locations, this is not precisely correct. It would be more accurate to subdivide carcinomas by different anatomical and clinical subtypes such as OSCC (oral squamous cell carcinoma), OPSCC (oropharyngeal squamous cell carcinoma), LSSC (laryngeal squamous cell carcinoma) as well as nasopharyngeal squamous cell carcinoma (NPSCC). This analysis focuses solely on OSCC, the largest subgroup of HNSCC next to OPSCC. Significant surgical efforts, including tumour site reconstruction using free microvascular flap techniques and excellent tumour patient aftercare, have improved the long-term quality of life in tumour patients [2]. However, despite these advances, disease-specific survival rates (DSS) are still low for OSCC. Local recurrence and locoregional second carcinomas often appear within first 5 years after the primary diagnosis of OSCC, and the 5-year survival rate of advanced OSCC is approximately 30 percent [3]. TNM and UICC stages are generally still used to describe the size of the tumour and its invasion into the lymphatic system from a topographic point of view, although recently, these systems were shown to be poor at accurately predicting an outcome [4, 5]. The latter classifications of OSCC are used for outcome prediction and decision-making concerning additional radiotherapy in cases of positive nodal metastasis [6]. Therefore, the need to detect reliable predictive biomarkers, genetic alterations or epigenetic modifications in OSCC to estimate recurrence for therapy stratification and planning is significant, and it is also important to identify significant molecular pathways to precisely select patients who are suitable for additional therapy [7–9].
Recently, HPV has been discussed as a relevant predictive biomarker for OSCC [10, 11]. Higher rates of HPV+ tumours, better outcomes for HPV+ patients, and improved survival rates compared to HPV− groups have often been published in studies that did not differentiate between OSCC and OPSCC [12, 13] and did not examine other confounders significantly influencing DSS. Only few studies were published which differentiate sufficiently between the anatomical localisations [14]. In OPSCC in particular, lymphoid tissue and lymphoid precursor cells interact with HPV infection and also carcinoma cells through various molecular mechanisms [15]. Because of this, HPV exhibiting p16INK4a positivity is more often seen in OPSCC [16], requiring OPSCC to be judged separately from OSCC. HPV infection is often detected with immunohistochemistry (IHC) of p16INK4a. HPV associated oncoproteins as E7 are influencing the expression of p16INK4a (Figures 1 and 2) [17–19]. Therefore p16INK4a is used as an HPV-related surrogate marker in many study protocols because the procedure is well-established and time saving [20, 21]. It has a distinct role in cell cycle regulation [22], acting as a tumour suppressor protein. It is well known that detection of HPV infection with p16INK4a staining often does not give accurate results [23, 24]. Another pitfall making survival comparisons even more difficult is the comparison of non-homogeneous groups. Bias by age and corresponding comorbidities as well as different tumour stages influence survival rates. Often, false conclusions are drawn based on different survival rates without regarding the diverse ages of the HPV+ and −groups.
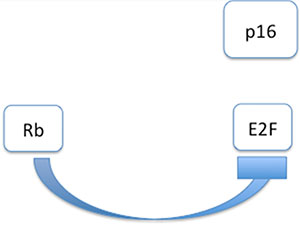
Figure 1: Interaction modell of p16INK4a in HPV negative cases: Tumoursuppressor Retinoblastoma gene (Rb) is inhibiting transcription factor E2F. As a consequence, p16INK4a is not affectd by E2F.
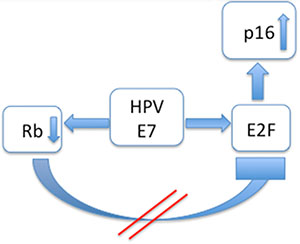
Figure 2: Interaction modell of p16INK4a in HPV positive cases: The HPV associated oncoprotein E7 acts in a double way: Rb is inhibited and E2F is promoted. As a result E2F is upregulated and p16INK4a is overexpressed.
To date, only a few OSCC studies have examined homogenous groups in terms of patient age and tumour stage. The aim of the current analysis was I) to compare a completely homogenous group and assess two of the most commonly used HPV detection methods with respect to HPV expression rates and II) to evaluate HPV status for relevance to patient survival and clinical outcome.
RESULTS
p16INK4a
12 (5.94%) patients tested positive for p16INK4a (Table 1). These results were clear with a strong intensity staining in all 12 cases (Figure 3), and cells with no detectable staining were considered negative (Figure 3). No intermediate staining was evaluated. All IHC results were evaluated under a light microscope (Zeiss, Jena, Germany). Samples of HPV+ and − cervical cancer were used as positive and negative controls. No difference of p16INK4a staining was given by the different tumour localisations as the centre of the tumour, the invasion front and the lymph nodes. If positivity/negativity of p16INK4a was given, every localisation was positive/negative for p16INK4a.
Table 1: Clinicopathological features
Clinical Parameters |
HPV − * (n = 195) |
HPV + * (n = 7) |
|---|---|---|
Median age in years (range) |
57.6 (29.5–85.8) |
57.4 (44.3–65.8) |
Gender |
||
Male/Female |
141/54 |
4/3 |
UICC Stage |
||
I |
48 |
× |
II |
35 |
× |
III |
38 |
3 |
IVa |
74 |
4 |
Tumor Size |
||
T1 |
72 |
2 |
T2 |
73 |
1 |
T3 |
21 |
2 |
T4a/b |
29 |
2 |
N Stage |
||
N0 |
100 |
3 |
N1 |
35 |
2 |
N2 |
60 |
2 |
Extracapsular spread |
21 |
2 |
Grading |
||
G1 |
12 |
× |
G2 |
130 |
5 |
G3 |
2 |
2 |
p16 |
||
Positive |
8 |
4 |
Negative |
187 |
3 |
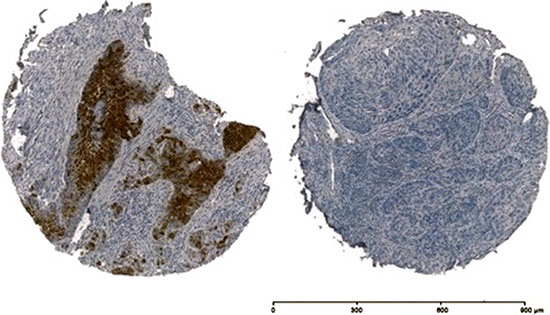
Figure 3: Immunohistochemistry of p16INK4a of two OSCC samples, on the left side p16INK4a + and on the right side p16INK4a.
HPV DNA PCR
7 (3.47%) patients were scored as HPV+ based on the PCR results (Figure 4.1). A sample from an HPV+ cervical carcinoma was used as a positive control (+C) to confirm HPV infection. Control PCR (Figure 4.2) was performed in every case to ensure sufficient amounts of DNA. Only in 4 (1.98%) cases did the PCR result correspond to positive p16INK4a staining. In 8 (3.96%) cases, p16INK4a staining was positive, but the PCR result was negative. In 3 (1.49%) cases, p16INK4a staining was negative but the PCR for HPV was positive. In conclusion, p16INK4a staining for the detection of HPV is insufficiently sensitive and specific.
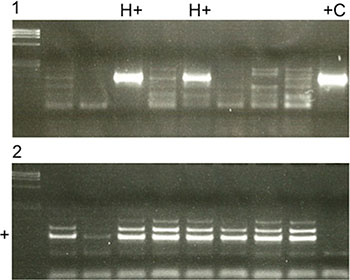
Figure 4: (1) PCR results for detecting HPV positive (H+) and negative cases (H−). A HPV positive cervix carcinoma was used as a positive control (+C). (2) A control PCR was performed in every case to detect sufficient amounts of DNA. If DNA amounts were insufficient as shown in row 2, DNA was newly extracted from FFPE. Otherwise a clear signal was seen in the specific row (+).
Detection of HPV subtypes
To identify the HPV subtype, each patient was scored with the HPV LCD Array Kit (Zytomed Systems, Berlin, Germany). Furthermore, it served as proof of the reliability of the PCR method, as there were no discordant results between the PCR compared to the HPV LCD Array Kit, indicating that every OSCC identified as HPV+ by PCR was also positive with the LCD kit. In 5 OSCC cases, HPV subtype 16 was present, in one case HPV subtypes 16 and 52 were present, and in one case HPV subtypes 16 and 53 were present.
Clinical data
Relevant clinical data from the 202 included patients with an OSCC diagnosis are listed in Table 1. Risk factors such as smoking and alcohol consumption were evaluated in comparable equal numbers from both groups. The majority of both sub-groups smoked, and the combination of smoking and regular alcohol consumption was observed in 10 percent of both sub-groups. Karnofsky performance status was assessed for every tumor patient. All patients of the current study had a Karnofsky status between 90–100 per cent, independent of HPV positivity. Due to high Karnofsky stages, the status was not significantly associated with the survival rates.
The survival rates from both groups were not significantly different according to univariate and multivariate analyses (p > 0.05). HPV+ patients with OSCC had an average overall survival (OS) of 33.17 months (sd 5.21; 95% Confidence Interval (CI): 22.96−43.38), and HPV− patients showed an average OS of 78.34 months (sd 4.27; 95% CI: 69.99–86.70).
Survival rates were also not significantly different between p16INK4a+ cases (average OS 40.08 months, sd 4.28; 95% CI: 69.99−86.77) and p16 INK4a− cases (average OS 78.38 months, sd 6.55; 95% CI: 27.24−52.92). HPV+ / p16INK4a+ cases had also no better survival rates (average OS 31.56 months, sd 16.62; 95% CI: 14,94−48.18).
Both local recurrence and lymph node recurrence played a major role in survival (p = 0.001) and was evaluated to the same degree in 52 patients. Three of these patients were HPV+, four were positive for p16INK4a, and two of the p16INK4a+ patients were also HPV+.
Recurrence free survival (RFS) of HPV+ patients was an average of 8.56 months (sd 1.56; 95% CI: 5.51−11.61); HPV− patients showed an average RFS of 18.66 months (sd 2.57; 95% CI: 13.63–23.69).
Recurrence free survival (RFS) of p16INK4a+ patients was an average of 22.05 months (sd 5.78; 95% CI: 10.73−33.38), and p16INK4a- patients had an average RFS of 17.70 months (sd 2.54; 95% CI: 12.73−22.67).
The UICC stage (p = 0.031), patients’ ages (p = 0.012) and lymph node metastasis (p = 0.003) at the time of primary diagnosis had a significant influence on overall survival rates independent of HPV status or p16INK4a.
In contrast, patient gender, T category, extra capsular spread and tumour grading were not significantly associated with overall tumour related survival or DSS (p > 0.05) independent of HPV or p16INK4a status. Resection margins were assessed and were R0, tumour free, in every case.
DISCUSSION
There is an ongoing discussion about the impact of an HPV infection on the prognosis and therapy regimes for HNSCC. To date, there are many ambiguities in the field. Because of unknowns in the literature, this study was performed to evaluate the HPV infection rate in a large homogenous collection of OSCC patients, examining different HPV detection methods and patients’ overall and recurrence free survival. To our knowledge, this is the first large study that does not show HPV+ status improves the survival rates of OSCC patients. Furthermore, this study demonstrated that HPV infection only occurs in a relatively small number of OSCC cases. In the literature, the rate of HPV positivity in HNSCC is provided with a wide range and often substantially differs between from one study to another [32, 33]. Further examination of published data shows that the wide range of HPV+ status in SCC is the result of poor differentiation between OSCC and OPSCC [34, 35]. Studies that only included OSCC in their evaluation show a comparably smaller number of HPV+ SCC, similar to this study [36, 37]. However, studies focusing on OPSCC, especially tonsil SCC, have higher HPV positivity rates [38] because the virus interacts with lymphoid tissue. The HPV has a selective tropism for the epithelium lining the tonsillar crypts. This interaction makes the difference to sites where lymphoid tissue is not the dominant tissue, such as the oral cavity. Some similarities in morphology and function are given in OPSCC and nasopharyngeal squamous cell carcinoma (NPSCC). The numbers of HPV positivity in NPSCC have also a big variety and are depending of the collectives and detection methods. Some experts, also written in the guidelines from the College of American Pathologists, promote HPV testing in EBV-negative NPSCC [39]. Further studies could give helpful knowlegde for patients with HPV+ NPSCC. A topic of interest for further research is the interaction between HPV and different tissue types, especially lymphoid tissue, in OPSCC [40]. Better outcomes of HPV+ HNSCC are mostly described for HPV+ OPSCC regarding adjuvant therapy strategies as adjuvant radiation [41] or antibody specific therapy [42].
Evaluation of the HPV detection methods that have been used in several studies provides an additional explanation for the wide range of HPV+ SCC reported. Various methods are available for HPV detection. As often discussed, an insufficient method for HPV detection that is still often applied is p16INK4a immunostaining [43]. Because this protein interacts with the cell cycle in multiple ways, it is an insensitive predictor of HPV status with a low predictive value for HPV infection [24] and should be viewed as an independent marker influencing the cell cycle of carcinoma cells. This supports our findings that p16INK4a immunostaining and overexpression is an insufficient HPV detection method, because we observed a high failure rate in cases that were positive for p16INK4a but were actually HPV−. Using the latter method, the number of false-positive HPV+ patients would be 50 percent higher. Studies that assessed p16INK4a immunostaining without other detection methods have a higher positive rate of HPV infection [44]. Remarkably, a strong correlation between p16INK4a overexpression and OPSCC, mostly tonsil SCC, is seen [45]. However, in OPSCC as well as in OSCC, the correlation between p16INK4a overexpression and HPV requires further research.
We used RT-PCR of extracted DNA from the FFPE of OSCC as the standard method for HPV detection and collected valuable results, as described previously [44]. One advantage is the possibility to use FFPE for RT-PCR, because this allowed a retrospective investigation of a large group OSCC patients as a screening method. Additionally, the HPV subtypes were detected with the LCD Array using DNA extracted from FFPE. The use of RT-PCR and the latter LCD array gives more advantages as relaiable results and better economical aspects than the use of the Sanger sequence analysis [46]. With the additional method of the LCD Array, we obtained more information about the oncogenic risk potential of underlying HPV subtypes in addition to confirmation of the RT-PCR results, as a positive result for the LCD Array was only observed if the OSCC DNA was HPV+. HPV subtype 16 was detected in all HPV+ cases, as the most common oncogenic HPV subtype described for OSCC in literature before [47] promote better OS rates. Further evaluated subtypes were 52 and 53, the latter one is ranked as potentially oncogenic. These results show the relevance of HPV driven carcinogenesis in the HPV+ cases of the current study. False positive results for HPV infection and an overestimation of the number of HPV+ patients could therefore be excluded in view of the results presented here. The low incidence of HPV in OSCC patients in the literature was clearly confirmed with the results of this study. p16INK4a overexpression was not relevant to differences in age, gender, tumour stage or the survival rates of p16INK4a+ patients, suggesting that p16INK4a plays a less important role in OSCC. Recent studies focusing on p16INK4a expression have described improved survival rates in p16INK4a+ patients without differentiating between the two main localisation subgroups of OSCC and OPSCC. Only a few OSCC studies focused on p16INK4a, making a comparison of the literature with this data difficult. Tumour patients treated over a certain period of time were collected, and homogeneous groups of tumour patients are sometimes rare but are a strong requirement for any serious comparison. Our study group showed relatively homogenous characteristics since both in the HPV+ and HPV− collective advanced tumor stages were occuring in high numbers/percentage of the total amount of included HPV+ and HPV− patients and allowed comparisons between HPV+ and − without bias. In the literature, HPV+ patients are primarily younger, which could certainly influence OS and DSS and promote better OS rates [48]. Our collective also showed homogenous results for all of the evaluated variables (Table 1). The patient specific data was completely independent of the presence of an HPV infection. The limitations of this study are, as discussed before, the lack of knowledge about the specific molecular pathways of p16INK4a and HPV. Further, multiple ways of interactions of HPV and different types of tissues of the oral cavity and pharynx could yet not be explained sufficiently. Another aim for prospect studies could be the distinct evaluation of these interactions.
MATERIALS AND METHODS
Patients
Two hundred two patients treated between 2009 and 2011 at our maxillofacial surgery department with the intention of curative treatment were included in our study. The inclusion criteria were the availability of relevant data (Table 1) from patients diagnosed with OSCC for statistical evaluation and of formalin fixed and paraffin embedded tissue (FFPE) for laboratory use. Follow up data were available for every patient of the study. All patients attended regular follow up examinations at our departement. Tumours of the tongue base, and tumours with involvement of the latter anatomical side were excluded in the current study to avoid bias of HPV interacting with lymphoid tissue. To the present knowledge these tumours should be classified as OPSCC. The therapy regimes of the patients included were primary surgery, with intraoperative margin control with the help of frozen sections and with neck dissection. Due to the german guidelines of oral cancer surgery [25] all patients underwent an ipsilateral selective neck dissection with clearance of the nodes of the ispsilateral levels I-III. In case of metastatic nodal disease (pN1 or pN2) that was intraoperatively confirmed by frozen section, a modified neck dissection with extended nodal clearance down to level IV and V and contra lateral neck dissection of level I-III was done. Postoperative adjuvant cisplatin-based chemoradiation, with a dose depending from 50 to 66 Gy, was performed in case of pN1, pN2 or tumor infiltration of the jaw or locally infiltrating tumor growth of the oral cavity (T4a/b), positive microscopic resection margins and/or extracapsular spread, according to the German guidelines for oral cancer.
Exclusion criteria were death resulting from a cause other than OSCC, distant metastasis at primary diagnosis and the use of primary radiochemotherapy before operation. The methods were approved by the local ethics committee (no. 212108) and are in accordance with the Declaration of Helsinki. All patients gave their consent.
Tissue microarray construction
Two independent pathologists defined the centre of the tumour and the invasion front and the lymph nodes of every study patient. The tissue was formalin fixed and paraffin embedded in blocks. The pathologists then marked the areas to be represented in the tissue microarray (TMA). A minimum of two tumour cores from the centre of the tumour, the invasion front and the corresponding lymph nodes with a 6 mm core size were assembled into the TMA using a Tissue Microarrayer (Beecher Instruments, Sun Praierie, USA) as described before [26].
IHC of p16INK4a
Four-micron TMA sections were stained with p16INK4a antibody (CIN tec; Ventana Medical Systems, Tucson, AZ, USA) according to the manufacturer’s recommendations. Visualisation of staining was performed with the chromogenic substrate 3.3′-diaminobenzidine (DakoCytomation; 10 min, room temperature). Finally, nuclei were counterstained with Mayer’s acid haematoxylin, and the slides were covered with Pertex mounting medium (MEDITE, Burgdorf, Germany) as described previously [27]. Positivity of p16INK4a was evaluated as described before with a methodological standardization to ensure result reproducibility [28].
DNA detection with PCR
FFPE full-face slides of the central tumour area from every patient were cut in 15-micron sections to extract DNA [29, 30]. Extraction of HPV DNA was performed with a QIAamp DNA FFPE Tissue Kit (Qiagen GmbH, Hilden Germany), and PCR was performed with prime Mix A and Mix B (Mix My 11/09 and Mix 125) provided by the LCD-Array HPV 3.5 kit (Zytomed Systems, Berlin, Germany).
Detection of HPV subtypes
After exhibiting a positive HPV signal by PCR, the HPV 3.5 LCD Array Kit (Zytomed Systems, Berlin, Germany) was used to distinguish which of the 32 HPV subtypes were present, including high and low risk types, as described previously [31]. After amplification of HPV DNA, hybridization was performed on the DNA LCD Array chip. Specific HPV+ probes were immobilised on the LCD chip surface. After washing the chips and drying via centrifugation, the visualisation of bound amplicons was possible with an enzyme-substrate reaction. Stained arrays were analysed with a scanning device and software.
Statistics
Data and figures were analysed with the ‘Statistical Package for the Social Sciences’ (SPSS for Windows, release 22.0.0, 2013, SPSS Inc., Chicago, IL, USA). Cox regression was used for survival analysis. Categories were tested for associations using cross tabs (Chi-squared-test). To compare groups, the non-parametric Mann Whitney U-test was used. Probabilities of less than 0.05 were considered significant.
CONCLUSIONS
p16INK4a immunostaining should not be used for HPV detection in OSCC. Furthermore, OSCC as a distinct localisation with principally squamous cell tissue should be clearly differentiated from OPSCC, which mainly contains lymphoid tissue. Considering the reliability of current data from the literature and the results presented here, there is no justification to promote therapy de-escalation in OSCC cases based on an HPV infection. HPV infection in OSCC is of low incidence and does not seem to be a predictive biomarker.
ACKNOWLEDGMENTS AND FUNDING
The authors would like to thank Daniela Angermeier and Daniela Hellmann for their excellent technical assistance.
CONFLICTS OF INTEREST
None.
REFERENCES
1. Ang KK, Harris J, Wheeler R, Weber R, Rosenthal DI, Nguyen-tan PF, Westra WH, Chung CH, Jordan RC, Lu C, Kim H, Axelrod R, Silverman CC. Human papillomavirus and survival of patients with oropharyngeal cancer. N Engl J Med. 2010; 363:24–35.
2. Annertz K, Rosenquist K, Andersson G, Jacobsson H, Hansson BG, Wennerberg J. High-risk HPV and survival in patients with oral and oropharyngeal squamous cell carcinoma - 5-year follow up of a population-based study. Acta Otolaryngol. 2014; 134:843–51.
3. Argiris A. EGFR inhibition for recurrent or metastatic HNSCC. Lancet Oncol. 2015; 16:488–9.
4. Benson E, LI R, Eisele D, Fakhry C. The clinical impact of HPV tumor status upon head and neck squamous cell carcinomas. Oral Oncol. 2014; 50:565–74.
5. Coppock JD, Vermeer PD, Vermeer DW, Lee KM, Miskimins WK, Spanos WC, LEE JH. mTOR inhibition as an adjuvant therapy in a metastatic model of HPV+ HNSCC. Oncotarget. 2016; 7:24228–41. doi: 10.18632/oncotarget.8286.
6. Dasgupta T, Nweze EI, YueH, Wang L, Jin J, Ghosh SK, Kawsar HI, Zender C, Androphy EJ, Weinberg A, Mccormick TS, Jin G. Human papillomavirus oncogenic E6 protein regulates human beta-defensin 3 (hBD3) expression via the tumor suppressor protein p53. Oncotarget. 2016; 7:27430–44. doi: 10.18632/oncotarget.8443.
7. Dok R, Kalev P, Van Limbergen EJ, Asbagh LA, Vazquez I, Hauben E, Sablina A, Nuyts S. p16INK4a impairs homologous recombination-mediated DNA repair in human papillomavirus-positive head and neck tumors. Cancer Res. 2014; 74:1739–51.
8. Duray A, Descamps G, Decaestecker C, Remmelink M, Sirtaine N, Lechien J, Ernoux-Neufcoeur P, Bletard N, Somja J, Depuydt CE, Delvenne P, Saussez S. Human papillomavirus DNA strongly correlates with a poorer prognosis in oral cavity carcinoma. Laryngoscope. 2012; 122:1558–65.
9. Gillison ML, Chaturvedi AK, Anderson WF, Fakhry C. Epidemiology of Human Papillomavirus-Positive Head and Neck Squamous Cell Carcinoma. J Clin Oncol. 2015; 33: 3235–42.
10. Jemal A, Siegel R, Ward E, Hao Y, Xu J, Thun MJ. Cancer statistics, 2009. CA Cancer J Clin. 2009; 59:225–49.
11. Kansy K, Thiele O, Freier K. The role of human papillomavirus in oral squamous cell carcinoma: myth and reality. Oral Maxillofac Surg. 2014; 18:165–72.
12. Keane FK, Chen YH, Neville BA, Tishler RB, Schoenfeld JD, Catalano PJ, Margalit DN. Changing prognostic significance of tumor stage and nodal stage in patients with squamous cell carcinoma of the oropharynx in the human papillomavirus era. Cancer. 2015; 121:2594–602.
13. Kolk A, Jubitz N, Mengele K, Mantwill K, Bissinger O, Schmitt M, Kremer M, Holm PS. Expression of Y-box-binding protein YB-1 allows stratification into long- and short-term survivors of head and neck cancer patients. Br J Cancer. 2011; 105:1864–73.
14. Krul EJ, Van De Vijver MJ, Schuuring E, Van Kanten RW, Peters AA, Fleuren GJ. Human papillomavirus in malignant cervical lesions in Surinam, a high-risk country, compared to the Netherlands, a low-risk country. Int J Gynecol Cancer. 1999; 9:206–211.
15. Larsen SR, Johansen J, Sorensen JA, Krogdahl A. The prognostic significance of histological features in oral squamous cell carcinoma. J Oral Pathol Med. 2009; 38: 657–62.
16. Lassen P, Primdahl H, Johansen J, Kristensen CA, Andersen E, Andersen LJ, Evensen JF, Eriksen JG, Overgaard J. Impact of HPV-associated p16-expression on radiotherapy outcome in advanced oropharynx and non-oropharynx cancer. Radiother Oncol. 2014; 113:310–6.
17. Lee CC, Ho HC, Su YC, Yu CH, Yang CC. Modified Tumor Classification With Inclusion of Tumor Characteristics Improves Discrimination and Prediction Accuracy in Oral and Hypopharyngeal Cancer Patients Who Underwent Surgery. Medicine (Baltimore). 2015; 94:e1114.
18. Liang C, Marsit CJ, Mcclean MD, Nelson HH, Christensen BC, Haddad RI, Clark JR, Wein RO, Grillone GA, Houseman EA, Halec G, Waterboer T, Pawlita M. Biomarkers of HPV in head and neck squamous cell carcinoma. Cancer Res. 2012; 72:5004–13.
19. Liggett WH, JR., Sidransky D. Role of the p16 tumor suppressor gene in cancer. J Clin Oncol. 1998; 16:1197–206.
20. Lingen MW, Xiao W, Schmitt A, Jiang B, Pickard R, Kreinbrink P, Perez-Ordonez B, Jordan RC, Gillison ML. Low etiologic fraction for high-risk human papillomavirus in oral cavity squamous cell carcinomas. Oral Oncol. 2013; 49:1–8.
21. Malinowsky K, Wolff C, Ergin B, Berg D, Becker KF. Deciphering signaling pathways in clinical tissues for personalized medicine using protein microarrays. J Cell Physiol. 2010a; 225:364–370.
22. Malinowsky K, Wolff C, Gundisch S, Berg D, Becker K. Targeted therapies in cancer-challenges and chances offered by newly developed techniques for protein analysis in clinical tissues. J Cancer. 2010b; 2:26–35.
23. Marur S, D‘souza G, Westra WH, Forastiere AA. HPV-associated head and neck cancer: a virus-related cancer epidemic. Lancet Oncol. 2010; 11:781–9.
24. Mirghani H, Amen F, Moreau F, Lacau ST, Guily J. Do high-risk human papillomaviruses cause oral cavity squamous cell carcinoma? Oral Oncol. 2015; 51:229–36.
25. Mucke T, Koschinski J, Wolff KD, Kanatas A, Mitchell DA, Loeffelbein DJ, Deppe H, Rau A. Quality of life after different oncologic interventions in head and neck cancer patients. J Craniomaxillofac Surg. 2015; 43:1895–8.
26. Nagai MA. Genetic alterations in head and neck squamous cell carcinomas. Braz J Med Biol Res. 1999; 32:897–904.
27. Nakao Y, Yang X, Yokoyama M, Ferenczy A, Tang SC, Pater MM, Pater A. Induction of p16 during immortalization by HPV 16 and 18 and not during malignant transformation. Br J Cancer. 1997; 75:1410–6.
28. Ndiaye C, Mena M, Alemany L, Arbyn M, Castellsague X, Laporte L, Bosch FX, De Sanjose S, Trottier H. HPV DNA, E6/E7 mRNA, and p16INK4a detection in head and neck cancers: a systematic review and meta-analysis. Lancet Oncol. 2014; 15:1319–31.
29. O‘rorke MA, Ellison MV, Murray LJ, Moran M, James J, Anderson LA. Human papillomavirus related head and neck cancer survival: a systematic review and meta-analysis. Oral Oncol. 2012; 48:1191–201.
30. Oguejiofor KK, Hall JS, Mani N, Douglas C, Slevin NJ, Homer J, Hall G, West CM. The prognostic significance of the biomarker p16 in oropharyngeal squamous cell carcinoma. Clin Oncol (R Coll Radiol). 2013; 25:630–8.
31. Okami K, Reed AL, Cairns P, Koch WM, Westra WH, Wehage S, Jen J, Sidransky D. Cyclin D1 amplification is independent of p16 inactivation in head and neck squamous cell carcinoma. Oncogene. 1999; 18:3541–5.
32. Ozalp SS, Us T, Arslan E, Oge T, Kasifoglu N. HPV DNA and Pap smear test results in cases with and without cervical pathology. J Turk Ger Gynecol Assoc. 2012; 13:8–14.
33. Rampias T, Sasaki C, Weinberger P, Psyrri A. E6 and e7 gene silencing and transformed phenotype of human papillomavirus 16-positive oropharyngeal cancer cells. J Natl Cancer Inst. 2009; 101:412–23.
34. Rettig EM, Wentz A, Posner MR, Gross ND, Haddad RI, Gillison ML, Fakhry C, Quon H, Sikora AG, Stott WJ, Lorch JH, Gourin CG, Guo Y. Prognostic Implication of Persistent Human Papillomavirus Type 16 DNA Detection in Oral Rinses for Human Papillomavirus-Related Oropharyngeal Carcinoma. JAMA Oncol. 2015; 1:907–15.
35. Robinson M, Suh YE, Paleri V, Devlin D, Ayaz B, Pertl L, Thavaraj S. Oncogenic human papillomavirus-associated nasopharyngeal carcinoma: an observational study of correlation with ethnicity, histological subtype and outcome in a UK population. Infect Agent Cancer. 2013; 8:30.
36. Rosenquist K, Wennerberg J, Annertz K, Schildt EB, Hansson BG, Bladstrom A, Andersson G. Recurrence in patients with oral and oropharyngeal squamous cell carcinoma: human papillomavirus and other risk factors. Acta Otolaryngol. 2007; 127:980–7.
37. Schlecht NF, Brandwein-Gensler M, Nuovo GJ, Li M, Dunne A, Kawachi N, Smith RV, Burk RD, Prystowsky MB. A comparison of clinically utilized human papillomavirus detection methods in head and neck cancer. Mod Pathol. 2011; 24:1295–305.
38. Shin DM, Khuri FR. Advances in the management of recurrent or metastatic squamous cell carcinoma of the head and neck. Head Neck. 2013; 35:443–53.
39. Smeets SJ, Hesselink AT, Speel EJ, Haesevoets A, Snijders PJ, Pawlita M, Meijer CJ, Braakhuis BJ, Leemans CR, Brakenhoff RH. A novel algorithm for reliable detection of human papillomavirus in paraffin embedded head and neck cancer specimen. Int J Cancer. 2007; 121:2465–72.
40. Straub M, Drecoll E, Pfarr N, Weichert W, Langer R, Hapfelmeier A, Gotz C, Wolff KD, Kolk A, Specht K. CD274/PD-L1 gene amplification and PD-L1 protein expression are common events in squamous cell carcinoma of the oral cavity. Oncotarget. 2016; 7:12024–34. doi: 10.18632/oncotarget.7593.
41. Syrjanen S. HPV infections and tonsillar carcinoma. J Clin Pathol. 2004; 57:449–55.
42. Walline HM, Komarck C, Mchugh JB, Byrd SA, Spector ME, Hauff SJ, Graham MP, Bellile E, Moyer JS, Prince ME, Wolf GT, Chepeha DB, Worden FP. High-risk human papillomavirus detection in oropharyngeal, nasopharyngeal, and oral cavity cancers: comparison of multiple methods. JAMA Otolaryngol Head Neck Surg. 2013; 139:1320–7.
43. Westra WH. Detection of human papillomavirus (HPV) in clinical samples: evolving methods and strategies for the accurate determination of HPV status of head and neck carcinomas. Oral Oncol. 2014; 50:771–9.
44. Wirth LJ. Cetuximab in Human Papillomavirus-Positive Oropharynx Carcinoma. J Clin Oncol. 2016.
45. Wolff KD, Follmann M, Nast A. The diagnosis and treatment of oral cavity cancer. Dtsch Arztebl Int. 2012; 109:829–35.
46. Young RJ, Urban D, Angel C, Corry J, Lyons B, Vallance N, Kleid S, Iseli TA, Solomon B, Rischin D. Frequency and prognostic significance of p16 protein overexpression and transcriptionally active human papillomavirus infection in laryngeal squamous cell carcinoma. Br J Cancer. 2015; 112:1098–1104.
47. Zhao N, Ang MK, Yin XY, Patel MR, Fritchie K, Thorne L, Muldrew KL, Hayward MC, Sun W, Wilkerson MD, Chera BS, Hackman T, Zanation AM. Different cellular p16(INK4a) localisation may signal different survival outcomes in head and neck cancer. Br J Cancer. 2012; 107:482–90.
48. Ziemann F, Arenz A, Preising S, Wittekindt C, Klussmann JP, Engenhart-Cabillic R, Wittig A. Increased sensitivity of HPV-positive head and neck cancer cell lines to x-irradiation +/− Cisplatin due to decreased expression of E6 and E7 oncoproteins and enhanced apoptosis. Am J Cancer Res. 2015; 5:1017–31.

