INTRODUCTION
Laryngeal cancer (LC) is a common type of malignant head and neck tumor, and the incidence is increasing yearly [1]. However, the etiology of LC remains unclear and the prognosis is poor. LC can result from both environmental and genetic factors [2, 3]. While the majority of LC patients have a history of smoking and alcohol consumption [4], only a small percentage of individuals with similar histories eventually develop LC. This suggests that genetic susceptibility underlies LC [5].
Host genetic factors may influence the prognosis of cancer patients. Recently, various genetic polymorphisms were associated with a risk of LC [6–9]. Polymorphisms may contribute to cancer susceptibility, progression, and response to therapy. Previous studies have primarily assessed associations between single nucleotide polymorphisms (SNPs) and LC risk using case-control models. Rare host genetic factors that influence the prognosis of advanced LC patients have been reported. Long-term longitudinal studies are required to evaluate the impact of SNPs on disease progression, treatment response, and patient survival.
In this study, we investigated 37 SNPs in 27 genes that were previously associated with head and neck cancers to determine whether they were associated with the prognosis of LC patients.
RESULTS
The demographic and clinical characteristics of the LC patients are shown in Table 1. The median age of the patients was 60 years (range, 32–82). All of the patients were men who were metastasis-free. The mean follow-up period was 38 months (range: 3–122). There were 100 deaths at the time of the last observation. Overall, the median survival time was 48 months.
Table 1: Characteristics of patients included in this study
Variables |
N (%) |
p value |
|---|---|---|
Total number of patients enrolled |
170 |
|
Age |
> 0.05 |
|
< 60 |
80 (47.06) |
|
≥ 60 |
90 (52.94) |
|
Tumor differentiation |
> 0.05 |
|
Well |
31 (18.24) |
|
Moderate |
125 (73.53) |
|
Poor |
14 (8.23) |
|
pT |
< 0.001 |
|
T1 |
40 (23.53) |
|
T2 |
62 (36.47) |
|
T3 |
50 (29.41) |
|
T4 |
18 (10.59) |
|
pN |
< 0.001 |
|
N0 |
116 (68.23) |
|
N1 |
30 (17.65) |
|
N2 |
24 (14.12) |
|
WHO grade |
< 0.001 |
|
I |
37 (21.76) |
|
II |
36 (21.18) |
|
III |
61 (35.88) |
|
IV |
36 (21.18) |
|
Surgery method |
< 0.001 |
|
Partial laryngectomy |
104 (61.18) |
|
Total laryngectomy |
66 (38.82) |
|
Cervical lymph node dissection |
> 0.05 |
|
Yes |
37 (21.76) |
|
No |
133 (78.24) |
|
No. patients with follow-up information available |
170 |
|
Median follow-up time, months |
38 |
|
Median survival time, months |
48 |
|
Status at last observation |
||
Alive |
70 (41.18) |
|
Death |
100 (58.82) |
T, pathologic tumor stage; N, pathologic nodal stage.
Clinical factors including age, pT, pN, WHO grade, degree of tumor differentiation, surgical method, and whether the patient underwent cervical lymph node dissection were assessed in a univariate analysis (Table 2). The distribution of the studied SNPs in the WHO grade was listed in Supplementary Table S1. We identified significant associations between clinical factors including the degree of tumor differentiation, pT, pN, WHO grade, and surgical method and LC patient prognosis. All of these factors increased the risk of mortality. Compared to patients with T1–T2 stage disease, N0, I–II grade, and who underwent partial laryngectomy, patients with T3–T4 stage, N1–N2, III–IV grade, and who underwent total laryngectomy had elevated risks of death, with HRs and 95% CIs of 2.17 (1.448–3.253), 2.394 (1.582–3.623), 3.298 (2.100–5.180) and 2.346 (1.576–3.492), respectively (Figure 1).
Table 2: Univariate analysis of the impact of clinical factors on prognosis for LC patients
Variables |
Overall survival |
|||
|---|---|---|---|---|
Event/total |
MST (months) |
HR (95% CI) |
Log-rank p |
|
Age |
||||
< 60 |
47/80 |
59.0 |
Ref |
|
≥ 60 |
53/90 |
48.0 |
1.161 (0.782–1.722) |
0.456 |
Tumor differentiation |
||||
Well |
18/31 |
71.0 |
Ref |
|
Moderate |
70/125 |
59.0 |
0.933 (0.556–1.567) |
0.008 |
Poor |
12/14 |
15.0 |
2.397 (1.150–4.997) |
|
pT |
||||
T1–T2 |
52/102 |
77.0 |
Ref |
|
T3–T4 |
48/68 |
32.0 |
2.170 (1.448–3.253) |
< 0.001 |
pN |
||||
N0 |
61/116 |
71.0 |
Ref |
|
N1-N2 |
39/54 |
26.0 |
2.394 (1.582–3.623) |
< 0.001 |
WHO grade |
||||
I–II |
30/73 |
98.0 |
Ref |
|
III–IV |
70/97 |
32.0 |
3.298 (2.100–5.180) |
< 0.001 |
Surgery method |
||||
Partial laryngectomy |
50/104 |
73.0 |
Ref |
|
Total laryngectomy |
50/66 |
30.0 |
2.346 (1.576–3.492) |
< 0.001 |
Cervical lymph node dissection |
||||
Yes |
19/37 |
36.0 |
Ref |
|
No |
81/133 |
56.0 |
0.711 (0.425–1.189) |
0.188 |
T, pathologic tumor stage; N, pathologic nodal stage;
MST, median survival time; HR, hazard ratio; 95% CI, 95% confidence interval.
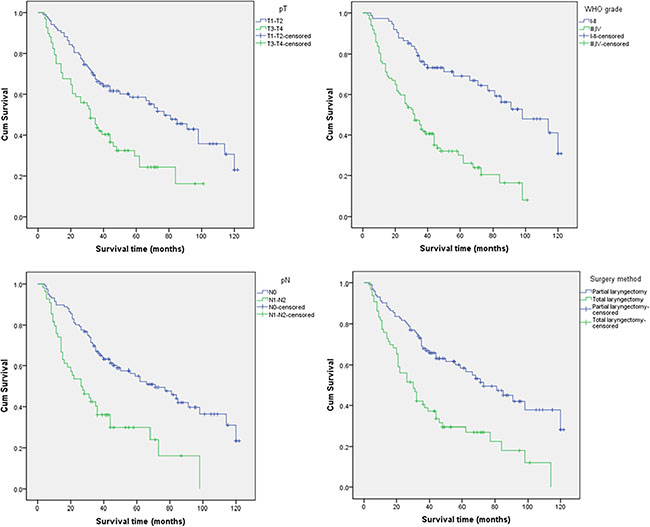
Figure 1: Kaplan-Meier analysis of LC patient overall survival according to the pT, pN, WHO grade, and surgical method.
The basic characteristics of all candidate SNPs that were analyzed in the study, including chromosome, position, band, alleles A/B, gene(s), and role(s), are shown in Table 3. Two of the 37 candidate SNPs evaluated showed statistically significantly correlations with overall survival (Table 4) according to Log-rank tests and Cox regression analysis. The A/G genotype of alcohol dehydrogenase 1B (ADH1B) rs1042026 (HR, 0.538; 95% CI, 0.345–0.839) and G/A genotype of rs1229984 (HR, 0.659; 95% CI, 0.438–0.991) were associated with increased overall survival. Kaplan-Meier curves of overall survival for the different genotypes of rs1042026 and rs1229984 are shown in Figure 2.
Table 3: Candidate SNP data
SNP ID |
Chr |
Position |
Band |
Alleles A/B |
Gene(s) |
Role |
|---|---|---|---|---|---|---|
rs13130787 |
4 |
94887031 |
4q22.2 |
T/C |
||
rs3805322 |
4 |
100056998 |
4q23 |
G/A |
ADH4 |
Intron |
rs1042026 |
4 |
100228466 |
4q23 |
A/G |
ADH1B |
3′ UTR |
rs1229984 |
4 |
100239319 |
4q23 |
G/A |
ADH1B |
Coding exon |
rs1789924 |
4 |
100274286 |
4q23 |
T/C |
ADH1C |
Promoter |
rs971074 |
4 |
100341861 |
4q23 |
A/G |
ADH7 |
Coding exon |
rs1000589 |
13 |
64141913 |
13q21.31 |
G/T |
||
rs1585440 |
13 |
66481815 |
13q21.32 |
A/C |
||
rs9573163 |
13 |
73908846 |
13q22.1 |
C/G |
||
rs9543325 |
13 |
73916628 |
13q22.1 |
T/C |
||
rs1886449 |
13 |
73932114 |
13q22.1 |
T/C |
||
rs2039553 |
13 |
80299722 |
13q31.1 |
G/A |
||
rs944289 |
14 |
36649246 |
14q13.3 |
T/C |
||
rs4444235 |
14 |
54410919 |
14q22.2 |
C/T |
BMP4 |
Downstream |
rs4779584 |
15 |
32994756 |
15q13.3 |
C/T |
SCG5 |
Downstream |
rs4785204 |
16 |
50103734 |
16q12.1 |
T/C |
HEATR3 |
Intron |
rs9929218 |
16 |
68820946 |
16q22.1 |
A/G |
CDH1 |
Intron |
rs17761864 |
17 |
2171637 |
17p13.3 |
A/C |
SMG6 |
Intron |
rs4924935 |
17 |
18753870 |
17p11.2 |
C/T |
PRPSAP2 |
Promoter |
rs225190 |
17 |
30877658 |
17q11.2 |
G/A |
MYO1D |
Intron |
rs6503659 |
17 |
39897264 |
17q21.2 |
A/T |
HAP1 |
Promoter |
rs2257205 |
17 |
56448297 |
17q22 |
A/G |
RNF43 |
Coding exon |
rs2847281 |
18 |
12821593 |
18p11.21 |
C/T |
PTPN2 |
Intron |
rs12456874 |
18 |
13366862 |
18p11.21 |
G/A |
C18orf1 |
Intron |
rs4939827 |
18 |
46453463 |
18q21.1 |
T/C |
SMAD7 |
Intron |
rs7504990 |
18 |
50517776 |
18q21.2 |
T/C |
DCC |
Intron |
rs961253 |
20 |
6404281 |
20p12.3 |
A/C |
||
rs2423279 |
20 |
7812350 |
20p12.3 |
C/T |
||
rs4925386 |
20 |
60921044 |
20q13.33 |
T/C |
LAMA5 |
Intron (boundary) |
rs372883 |
21 |
30717737 |
21q21.3 |
G/A |
BACH1 |
Intron |
rs455804 |
21 |
31146169 |
21q21.3 |
T/G |
NCRNA00110 |
Downstream |
rs2014300 |
21 |
36357861 |
21q22.12 |
A/G |
RUNX1 |
Intron |
rs1547374 |
21 |
43778895 |
21q22.3 |
G/A |
TFF1 |
Downstream |
rs4822983 |
22 |
29115066 |
22q12.1 |
T/C |
CHEK2 |
Intron |
rs738722 |
22 |
29130012 |
22q12.1 |
T/C |
HSCB |
Promoter |
rs2239815 |
22 |
29192670 |
22q12.1 |
T/C |
XBP1 |
Intron |
rs5768709 |
22 |
48929569 |
22q13.32 |
A/G |
FAM19A5 |
Intron |
A/B, minor/major alleles; Chr, chromosome.
Table 4: Univariate analysis of the associations between the candidate SNPs and LC patient survival
SNP ID |
Genotype |
Event/total |
MST (months) |
HR (95% CI) |
Log-rank p |
|---|---|---|---|---|---|
rs13130787 |
|||||
C/C |
19/38 |
98 |
Ref |
||
T/C |
71/113 |
39 |
1.638 (0.986–2.722) |
0.152 |
|
T/T |
10/19 |
62 |
1.414 (0.656–3.049) |
||
rs3805322 |
|||||
A/A |
22/39 |
73 |
Ref |
||
G/A |
59/98 |
44 |
0.982 (0.601–1.604) |
0.997 |
|
G/G |
19/33 |
50 |
0.981 (0.529–1.818) |
||
rs1042026 |
|||||
G/G |
30/40 |
31 |
Ref |
||
A/G |
58/114 |
81 |
0.538 (0.345–0.839) |
0.001 |
|
A/A |
3/4 |
8 |
2.344 (0.709–7.741) |
||
rs1229984 |
|||||
A/A |
39/54 |
36 |
Ref |
||
G/A |
57/104 |
48 |
0.659 (0.438–0.991) |
0.043 |
|
G/G |
2/9 |
0.291(0.070-1.206) |
|||
rs1789924 |
|||||
C/C |
95/158 |
46 |
Ref |
||
T/C |
4/11 |
84 |
0.542 (0.198–1.481) |
0.222 |
|
T/T |
|||||
rs971074 |
|||||
G/G |
73/125 |
50 |
Ref |
||
A/G |
26/44 |
46 |
1.118 (0.714–1.751) |
0.624 |
|
A/A |
|||||
rs1000589 |
|||||
T/T |
42/66 |
38 |
Ref |
||
G/T |
38/77 |
68 |
0.691 (0.444–1.075) |
0.003 |
|
G/G |
20/26 |
22 |
1.711 (0.997-2.937) |
||
rs1585440 |
|||||
C/C |
23/33 |
62 |
Ref |
||
A/C |
75/134 |
48 |
0.803 (0.502–1.286) |
0.595 |
|
A/A |
1/2 |
11 |
1.299 (0.174–9.683) |
||
rs9573163 |
|||||
G/G |
30/59 |
62 |
Ref |
||
C/G |
56/92 |
40 |
1.397 (0.896–2.180) |
0.113 |
|
C/C |
14/19 |
36 |
1.882 (0.996–3.558) |
||
rs9543325 |
|||||
C/C |
33/47 |
35 |
Ref |
||
T/C |
47/86 |
66 |
0.691 (0.443–1.080) |
0.200 |
|
T/T |
20/37 |
59 |
0.674 (0.386–1.176) |
||
rs1886449 |
|||||
C/C |
6/11 |
77 |
Ref |
||
T/C |
75/131 |
48 |
1.193 (0.518–2.746) |
0.898 |
|
T/T |
2/2 |
20 |
1.365 (0.266–6.991) |
||
rs2039553 |
|||||
A/A |
32/58 |
81 |
Ref |
||
G/A |
19/32 |
32 |
1.474 (0.832–2.612) |
0.389 |
|
G/G |
19/36 |
62 |
1.075 (0.607–1.902) |
||
rs944289 |
|||||
C/C |
16/26 |
62 |
Ref |
||
T/C |
79/137 |
50 |
0.712 (0.414–1.223) |
0.068 |
|
T/T |
2/3 |
6 |
2.792 (0.638–12.221) |
||
rs4444235 |
|||||
T/T |
25/34 |
40 |
Ref |
||
C/T |
63/115 |
59 |
0.762 (0.479–1.211) |
0.506 |
|
C/C |
6/11 |
84 |
0.859 (0.352–2.098) |
||
rs4779584 |
|||||
T/T |
65/103 |
48 |
Ref |
||
C/T |
30/57 |
62 |
0.805 (0.521–1.242) |
0.611 |
|
C/C |
5/10 |
44 |
0.905 (0.363–2.256) |
||
rs4785204 |
|||||
C/C |
51/87 |
46 |
Ref |
||
T/C |
32/54 |
59 |
1.002 (0.643–1.560) |
0.803 |
|
T/T |
10/16 |
37 |
1.248 (0.631–2.467) |
||
rs9929218 |
|||||
G/G |
70/118 |
48 |
Ref |
||
A/G |
27/48 |
73 |
0.855 (0.548–1.334) |
0.081 |
|
A/A |
2/2 |
14 |
3.931 (0.946–16.324) |
||
rs17761864 |
|||||
C/C |
75/120 |
46 |
Ref |
||
A/C |
16/37 |
0.660 (0.384–1.133) |
0.087 |
||
A/A |
3/4 |
7 |
2.282 (0.714–7.297) |
||
rs4924935 |
|||||
T/T |
80/130 |
44 |
Ref |
||
C/T |
13/28 |
98 |
0.596 (0.331–1.074) |
0.110 |
|
C/C |
5/7 |
32 |
1.578 (0.635–3.920) |
||
rs225190 |
|||||
A/A |
54/96 |
62 |
Ref |
||
G/A |
40/65 |
44 |
1.137 (0.754–1.715) |
0.634 |
|
G/G |
4/6 |
18 |
1.528 (0.551–4.238) |
||
rs6503659 |
|||||
T/T |
73/123 |
56 |
Ref |
||
A/T |
23/42 |
36 |
0.981 (0.614–1.569) |
0.634 |
|
A/A |
2/3 |
22 |
1.935 (0.473–7.918) |
||
rs2257205 |
|||||
G/G |
26/41 |
44 |
Ref |
||
A/G |
55/102 |
62 |
0.647 (0.403–1.039) |
0.152 |
|
A/A |
16/24 |
48 |
0.891 (0.477–1.664) |
||
rs2847281 |
|||||
T/T |
75/129 |
56 |
Ref |
||
C/T |
24/38 |
44 |
1.095 (0.690–1.739) |
0.698 |
|
C/C |
|||||
rs12456874 |
|||||
A/A |
91/158 |
48 |
Ref |
||
G/A |
9/12 |
26 |
1.327 (0.667–2.638) |
0.415 |
|
G/G |
|||||
rs4939827 |
|||||
C/C |
72/118 |
48 |
Ref |
||
T/C |
23/46 |
91 |
0.920 (0.575–1.473) |
0.772 |
|
T/T |
1/1 |
36 |
1.818 (0.251–13.145) |
||
rs7504990 |
|||||
C/C |
60/99 |
44 |
Ref |
||
T/C |
36/62 |
62 |
0.889 (0.587–1.347) |
0.729 |
|
T/T |
4/9 |
71 |
0.720 (0.261–1.982) |
||
rs961253 |
|||||
C/C |
87/146 |
44 |
Ref |
||
A/C |
13/24 |
68 |
0.928 (0.518–1.663) |
0.801 |
|
A/A |
|||||
rs2423279 |
|||||
T/T |
65/117 |
48 |
Ref |
||
C/T |
30/47 |
59 |
0.976 (0.632–1.508) |
0.912 |
|
C/C |
0/1 |
||||
rs4925386 |
|||||
C/C |
59/107 |
62 |
Ref |
||
T/C |
17/25 |
40 |
1.160 (0.676–1.991) |
0.608 |
|
T/T |
2/5 |
0.574 (0.139–2.370) |
|||
rs372883 |
|||||
A/A |
7/15 |
Ref |
|||
G/A |
82/137 |
56 |
0.993 (0.457–2.159) |
0.958 |
|
G/G |
1/2 |
11 |
1.331 (0.163–10.863) |
||
rs455804 |
|||||
G/G |
47/84 |
62 |
Ref |
||
T/G |
42/69 |
44 |
1.228 (0.809–1.863) |
0.126 |
|
T/T |
8/12 |
22 |
2.115 (0.989–4.520) |
||
rs2014300 |
|||||
G/G |
73/127 |
48 |
Ref |
0.522 |
|
A/G |
25/38 |
39 |
1.113 (0.706–1.755) |
||
A/A |
1/1 |
26 |
2.793 (0.385–20.276) |
||
rs1547374 |
|||||
A/A |
22/38 |
56 |
Ref |
||
G/A |
61/106 |
59 |
0.994 (0.610–1.623) |
0.330 |
|
G/G |
16/24 |
35 |
1.493 (0.777–2.866) |
||
rs4822983 |
|||||
C/C |
66/114 |
50 |
Ref |
||
T/C |
29/45 |
44 |
1.051 (0.678–1.628) |
0.976 |
|
T/T |
3/5 |
32 |
1.008 (0.312–3.257) |
||
rs738722 |
|||||
C/C |
17/27 |
66 |
Ref |
||
T/C |
65/119 |
59 |
0.713 (0.417–1.218) |
0.370 |
|
T/T |
3/4 |
5 |
1.113 (0.315–3.933) |
||
rs2239815 |
|||||
C/C |
34/65 |
56 |
Ref |
||
T/C |
50/77 |
44 |
1.300 (0.840–2.012) |
0.325 |
|
T/T |
6/11 |
114 |
1.113 (0.332–1.915) |
||
rs5768709 |
|||||
G/G |
18/31 |
59 |
Ref |
||
A/G |
77/132 |
48 |
1.025 (0.613–1.715) |
0.951 |
|
A/A |
5/7 |
35 |
1.173 (0.430–3.200) |
MST, median survival time; HR, hazard ratio; 95% CI, 95% confidence interval;
Long-rank p values were calculated using the Chi-Square test.
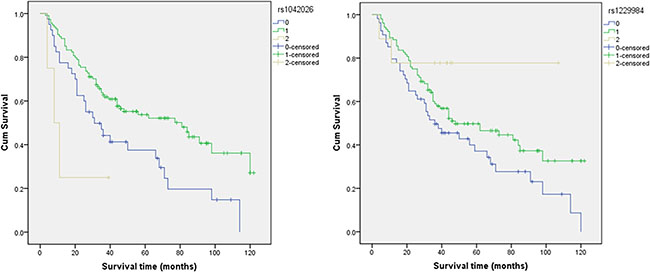
Figure 2: The individual effects of rs1042026 and rs1229984 on overall survival.
After adjusting for the various clinical factors, multivariate Cox regression analysis demonstrated that SNP genotype was an independent prognostic factor for overall survival. We identified significant correlations between two SNPs (ADH1B rs1229984 and CDH1 rs9929218) and the prognosis of LC patients (Table 5). The G/A genotype of ADH1B rs1229984 was associated with increased overall survival (HR, 0.537; 95% CI, 0.340–0.848; p = 0.008), and the A/A genotype of CDH1 rs9929218 with reduced overall survival (HR, 6.074; 95% CI, 1.426–25.870; p = 0.0015).
Table 5: Multivariate analysis of the associations between candidate SNPs and LC patient survival
SNP ID |
Genotype |
HR (95% CI) |
p |
|---|---|---|---|
rs13130787 |
|||
C/C |
Ref |
||
T/C |
1.641 (0.975–2.760) |
0.062 |
|
T/T |
1.868 (0.856–4.073) |
0.116 |
|
rs1042026 |
|||
G/G |
Ref |
||
A/G |
0.668 (0.420–1.060) |
0.087 |
|
A/A |
1.898 (0.550–6.543) |
0.310 |
|
rs1229984 |
|||
A/A |
Ref |
||
G/A |
0.537 (0.340–0.848) |
0.008 |
|
G/G |
0.352 (0.084–1.477) |
0.154 |
|
rs1000589 |
|||
T/T |
Ref |
||
G/T |
0.773 (0.483–1.236) |
0.282 |
|
G/G |
1.734 (0.999–3.010) |
0.050 |
|
rs9573163 |
|||
G/G |
Ref |
||
C/G |
1.515 (0.958–2.394) |
0.075 |
|
C/C |
1.573 (0.811–3.050) |
0.180 |
|
rs9543325 |
|||
C/C |
Ref |
||
T/C |
0.791 (0.505–1.240) |
0.307 |
|
T/T |
0.673 (0.378–1.200) |
0.180 |
|
rs944289 |
|||
C/C |
Ref |
||
T/C |
0.831 (0.477–1.448) |
0.514 |
|
T/T |
2.391 (0.513–11.139) |
0.267 |
|
rs9929218 |
|||
G/G |
Ref |
||
A/G |
0.998 (0.637–1.565) |
0.994 |
|
A/A |
6.074 (1.426–25.870) |
0.015 |
|
rs17761864 |
|||
C/C |
Ref |
||
A/C |
0.577 (0.328–1.018) |
0.058 |
|
A/A |
1.82 (0.523–6.338) |
0.347 |
|
rs4924935 |
|||
T/T |
Ref |
||
C/T |
0.692 (0.380–1.261) |
0.229 |
|
C/C |
1.334 (0.501–3.549) |
0.564 |
|
rs2257205 |
|||
G/G |
Ref |
||
A/G |
0.783 (0.478–1.284) |
0.333 |
|
A/A |
0.863 (0.454–1.642) |
0.654 |
|
rs455804 |
|||
G/G |
Ref |
||
T/G |
1.199 (0.783–1.838) |
0.404 |
|
T/T |
1.872 (0.812–4.312) |
0.141 |
HR, hazard ratio; 95% CI, 95% confidence interval.
All p values were calculated using the Wald test.
A p < 0.05 indicates statistical significance.
DISCUSSION
We evaluated the effects of 37 SNPs in 26 genes on the prognosis of 170 Han Chinese male LC patients. We demonstrated that ADH1B rs1042026, ADH1B rs1229984, and CDH1 rs9929218 were significantly associated with overall survival. Our data shed new light on the association between genetic variations in the ADH1B and CDH1 genes and LC prognosis in the Han Chinese population.
The ADH1B gene is located on chromosome 4q21-q23. The rs1229984 variant in the ADH1B gene causes a missense mutation (R48H) which increases the activity of the ADH1B enzyme (i.e. faster acetaldehyde production generated by ethanol oxidation) [10, 11]. Following alcohol consumption, elevated ADH1B activity is thought to transiently increase the level of acetaldehyde, which leads to unpleasant effects that limit the desire to continue drinking. A meta-analysis of this variant in Asian, European, and African Americans populations (where the rs1229984 A allele is common) demonstrated a strong association with alcohol-related disorder risk [12–14].
The CDH1 gene encodes the E-cadherin protein, which is a 120 kDa glycoprotein that consists of an extracellular domain containing five tandem repeats, a cytoplasmic domain, and a single transmembrane domain [15, 16]. CDH1 hypermethylation is one of the mechanisms by which E-cadherin expression is silenced. Abnormal CDH1 expression has been linked to many human diseases including cancer, nephrolithiasis, pre-eclampsia, and ectopic pregnancy [17, 18]. Association between rs9929218 and both colorectal cancer risk and survival have also been observed [19–21]. Our results demonstrated that CDH1 rs9929218 AA genotype was associated with reduced overall survival in the Chinese Han population. However, additional studies with larger cohorts derived from other populations are necessary.
Differences in survival were most apparent in individuals with T stage. There are several possible explanations for this finding. First, because individuals with T stage have already acquired many somatic mutations that could drive tumor growth or therapeutic resistance, subtle variations that alter the DNA repair capacity will not have a significant impact. Second, the differences in survival may reflect radiation-related outcomes, given that most T stage individuals received radiation treatment for the primary tumor, whereas only a minority of T stage individuals received radiation for treatment of the primary tumor. However, the latter explanation does not account for the common occurrence of relapsed metastatic disease outside the field of radiation.
In summary, our data raise the possibility that three polymorphisms may be one of major driving forces of LC progression, and could be valuable prognostic markers for LC patients. Further studies will focus on the functional experiments based on the relevant genes on animal models, to investigate detailed mechanism involved.
MATERIALS AND METHODS
Study participants
A total of 170 male patients (median age 60 years, range 32–82) who were diagnosed with LC at the First Affiliated Hospital of the Medical College of Xi’an Jiaotong University before 2002 and who were followed-up from January 2002 to April 2013 were included in the study. All patients underwent resection for LC at the same hospital. Additionally, all patients were Han Chinese from Xi’an city and the surrounding regions. The research protocol was performed according to the Declaration of Helsinki and was approved by the Human Research Committee of the First Affiliated Hospital of the Medical College of Xi’an Jiaotong University for the Approval of Research Involving Human Subjects. Informed consent was obtained from all patients.
Demographic and clinical data
Patient demographic and clinical data including age, sex, ethnicity, residential region, smoking status, alcohol use, education status, body mass index, and family history of cancer were collected through in-person interviews using a standardized epidemiological questionnaire. Detailed clinical information including the time of diagnosis, time of surgery and/or treatment with chemotherapy, time of recurrence and/or death, tumor stage, degree of differentiation, location, whether lymph node dissection was performed, and the treatment protocol was collected through medical chart review or physician consultation. Standard follow-up was performed by a trained specialist through on-site interviews, direct calls, or written communication with either patients or family members. The most recent follow-up data in this analysis were obtained in April 2013. No patients were lost during follow-up.
SNP selection and genotyping
We selected 37 SNPs with a minor allele frequency (MAF) > 5% in the HapMap Han Chinese population in Beijing that were previously associated with head and neck cancer [22–24] for genotyping. Genomic DNA was extracted from peripheral blood leukocytes using GoldMag® nanoparticles (GoldMag Ltd. Xi’an, China) according to the manufacturer’s instructions. DNA concentrations were estimated using a NanoDrop 2000 (Thermo Scientific, Waltham, Massachusetts, USA). The Sequenom MassARRAY Assay Design 3.0 Software was used to design Multiplexed SNP MassEXTEND assays [25]. Genotyping was performed using a Sequenom MassARRAY RS1000 [25]. Data management and analysis were performed using the Sequenom Typer 4.0 software as previously described [25, 26].
Data analysis
All follow-up survey and experimental data were analyzed using SPSS 17.0 (SPSS, Chicago, IL, USA). Survival time was defined as the time between the date of diagnosis and either the date of death (deceased patients) or last contact date (living patients). The Kaplan-Meier method was used to estimate overall survival. The survival curves were compared using Log-rank tests. Univariate analysis included the following factors: age, degree of tumor differentiation, pathologic tumor stage (pT), pathologic nodal stage (pN), WHO grade, surgical method, whether cervical lymph node dissection was performed, and the 37 candidate SNPs. Univariate and multivariable Cox proportional hazard models were used to calculate hazard ratio (HRs) and 95% confidence intervals (CIs). Two-sided p values < 0.05 were considered statistically significant and were calculated using the Wald test.
CONFLICTS OF INTEREST
The authors declare that there are no conflicts of interest.
REFERENCES
1. Markou K, Christoforidou A, Karasmanis I, Tsiropoulos G, Triaridis S, Constantinidis I, Vital V, Nikolaou A. Laryngeal cancer: epidemiological data from Nuorthern Greece and review of the literature. Hippokratia. 2013; 17:313–318.
2. Burduk PK. Association between infection of virulence cagA gene Helicobacter pylori and laryngeal squamous cell carcinoma. Med Sci Monit. 2013; 19:584–591.
3. Jaworowska E, Trubicka J, Lener MR, Masojc B, Zlowocka-Perlowska E, McKay JD, Renard H, Oszutowska D, Wokolorczyk D, Lubinski J, Grodzki T, Serwatowski P, Nej-Wolosiak K, et al. Smoking related cancers and loci at chromosomes 15q25, 5p15, 6p22.1 and 6p21.33 in the Polish population. PloS one. 2011; 6:e25057.
4. Ohshima H, Tazawa H, Sylla BS, Sawa T. Prevention of human cancer by modulation of chronic inflammatory processes. Mutat Res. 2005; 591:110–122.
5. Hashibe M, Brennan P, Benhamou S, Castellsague X, Chen C, Curado MP, Dal Maso L, Daudt AW, Fabianova E, Fernandez L, Wunsch-Filho V, Franceschi S, Hayes RB, et al. Alcohol drinking in never users of tobacco, cigarette smoking in never drinkers, and the risk of head and neck cancer: pooled analysis in the International Head and Neck Cancer Epidemiology Consortium. J Natl Cancer Inst. 2007; 99:777–789.
6. Mutlu P, Mutlu M, Yalcin S, Yaylaci A, Unsoy G, Saylam G, Akin I, Gunduz U, Korkmaz H. Association between XRCC3 Thr241Met polymorphism and laryngeal cancer susceptibility in Turkish population. Eur Arch Oto rhino laryngol. 2015; 272:3779–3784.
7. Lin H, Lin D, Zheng C. Association of XPD Lys751Gln polymorphism with head and neck cancer susceptibility: evidence from 11,443 subjects. Diagn Pathol. 2014; 9:15.
8. Li X, Xu J, Yang X, Wu Y, Cheng B, Chen D, Bai B. Association of single nucleotide polymorphisms of nucleotide excision repair genes with laryngeal cancer risk and interaction with cigarette smoking and alcohol drinking. Tumour Biol. 2014; 35:4659–4665.
9. Lu B, Li J, Gao Q, Yu W, Yang Q, Li X. Laryngeal cancer risk and common single nucleotide polymorphisms in nucleotide excision repair pathway genes ERCC1, ERCC2, ERCC3, ERCC4, ERCC5 and XPA. Gene. 2014; 542:64–68.
10. Edenberg HJ, Foroud T. Genetics and alcoholism. Nat Rev Gastro Hepa. 2013; 10:487–494.
11. Hurley TD, Edenberg HJ. Genes encoding enzymes involved in ethanol metabolism. Alcohol Res. 2012; 34:339–344.
12. Li D, Zhao H, Gelernter J. Strong association of the alcohol dehydrogenase 1B gene (ADH1B) with alcohol dependence and alcohol-induced medical diseases. Biol Psychiat. 2011; 70:504–512.
13. Bierut LJ, Goate AM, Breslau N, Johnson EO, Bertelsen S, Fox L, Agrawal A, Bucholz KK, Grucza R, Hesselbrock V, Kramer J, Kuperman S, Nurnberger J, et al. ADH1B is associated with alcohol dependence and alcohol consumption in populations of European and African ancestry. Mol Psychiatr. 2012; 17:445–450.
14. Gelernter J, Kranzler HR, Sherva R, Almasy L, Koesterer R, Smith AH, Anton R, Preuss UW, Ridinger M, Rujescu D, Wodarz N, Zill P, Zhao H, et al. Genome-wide association study of alcohol dependence:significant findings in African- and European-Americans including novel risk loci. Mol Psychiatr. 2014; 19:41–49.
15. Nollet F, Kools P and van Roy F. Phylogenetic analysis of the cadherin superfamily allows identification of six major subfamilies besides several solitary members. J Mol Biol. 2000; 299:551–572.
16. Blaschuk OW, Sullivan R, David S, Pouliot Y. Identification of a cadherin cell adhesion recognition sequence. Dev Biol. 1990; 139:227–229.
17. Stemmler MP, Bedzhov I. A Cdh1HA knock-in allele rescues the Cdh1-/- phenotype but shows essential Cdh1 function during placentation. Dev Dyn. 2010; 239:2330–2344.
18. Zhang J, Jin T, Yunus Z, Li X, Geng T, Wang H, Cui Y, Chen C. Genetic polymorphisms of VIP variants in the Tajik ethnic group of northwest China. BMC Genet. 2014; 15:102.
19. Lubbe SJ, Di Bernardo MC, Broderick P, Chandler I, Houlston RS. Comprehensive evaluation of the impact of 14 genetic variants on colorectal cancer phenotype and risk. Am J Epidemiol. 2012; 175:1–10.
20. Burnett-Hartman AN, Newcomb PA, Hutter CM, Peters U, Passarelli MN, Schwartz MR, Upton MP, Zhu LC, Potter JD, Makar KW. Variation in the association between colorectal cancer susceptibility loci and colorectal polyps by polyp type. Am J Epidemiol. 2014; 180:223–232.
21. Abuli A, Lozano JJ, Rodriguez-Soler M, Jover R, Bessa X, Munoz J, Esteban-Jurado C, Fernandez-Rozadilla C, Carracedo A, Ruiz-Ponte C, Cubiella J, Balaguer F, Bujanda L, et al. Genetic susceptibility variants associated with colorectal cancer prognosis. Carcinogenesis. 2013; 34:2286–2291.
22. Bediaga NG, Marichalar-Mendia X, Rey-Barja N, Setien-Olarra A, Gonzalez-Garcia JA, de Pancorbo MM, Aguirre-Urizar JM, Acha-Sagredo A. Polymorphisms in alcohol and tobacco metabolism genes in head and neck cancer in the Basque Country. J Oral Pathol Med. 2015; 44:769–775.
23. Zhang Y, Gu N, Miao L, Yuan H, Wang R, Jiang H. Alcohol dehydrogenase-1B Arg47His polymorphism is associated with head and neck cancer risk in Asian: a meta-analysis. Tumour Biol. 2015; 36:1023–1027.
24. Yuan H, Ma H, Lu F, Yuan Z, Wang R, Jiang H, Hu Z, Shen H, Chen N. Genetic variants at 4q23 and 12q24 are associated with head and neck cancer risk in China. Mol Carcinogen. 2013; 52:E2–9.
25. Gabriel S, Ziaugra L, Tabbaa D. SNP genotyping using the Sequenom MassARRAY iPLEX platform. Current protocols in human genetics / editorial board, Jonathan L Haines [et al]. 2009; Chapter 2:Unit 2.12.
26. Thomas RK, Baker AC, Debiasi RM, Winckler W, Laframboise T, Lin WM, Wang M, Feng W, Zander T, MacConaill L, Lee JC, Nicoletti R, Hatton C, et al. High-throughput oncogene mutation profiling in human cancer. Nat Genet. 2007; 39:347–351.

