INTRODUCTION
Human acute myelogenous leukemia (AML) mainly occurring in adults is a genetically and epigenetically heterogeneous disorder, which presents a high morbidity and mortality all over the world including in China [1–5]. Despite genetic heterogeneity of the disease, patients excluding FAB-M3 AML have been treated with similar combinations of cytarabine and anthracyclines for over 4 decades with little improvement in clinical outcome [3, 4, 6, 7].
DNA (cytosine-5-)-methyltransferase 3 alpha (DNMT3A) gene encodes a de nove methyltransferase, which was known as one of the most frequently mutated genes in AML [8–12]. DNMT3A mutation, probably a driver mutation, shows high stability from AML initiation up to relapse [8, 13–17]. DNMT3A R882 mutation, the main form of DNMT3A mutations [10, 18–21], acts commonly as an unfavorable prognostic marker in AML patients [11, 21–27].
Previous literatures elicited that DNMT3A polymorphisms were mainly associated with susceptibility of solid tumors, such as gastric carcinoma [28–31], colorectal cancer [32], hepatocellular carcinoma [33], ovarian cancer [34, 35] and breast cancer [36, 37] Besides, DNMT3A polymorphisms can lead to changes in methylation of downstream genes, such as LINE-1 and the imprinted gene PEG3 [38–40]. So far, there is no report regarding association between DNMT3A polymorphisms and the outcome of AML patients, although polymorphisms in some genes are shown to play vital roles in initiation and development of hematologic malignancies [41–44].
In consideration that prognostic stratification was critical for determination of therapeutic regimens and improving survival for AML patients [8, 45–48], we determined the associations of DNMT3A single nucleotide polymorphisms (SNPs) with the prognosis of AML patients from southern China. And the associations of DNMT3A SNPs with the patients’ chemosensitivity were also assessed.
RESULTS
The baseline characteristics of patients
The entire study population consisted of 352 diagnostic non-FAB-M3 AML patients who received induction chemotherapy with cytarabine and anthracyclines and were evaluable for response. Genotypes were unavailable from 8 patients for all eleven SNPs. The rest of 344 subjects were eligible for this study. Baseline characteristics in overall subjects were shown in Table 1. Undergoing a median follow-up of 442 days (range, 15-1500 days), 60% patients achieved a complete remission (CR). All the subjects were self-reported Chinese ancestry. The median age was 42 years (range, 14-79 years), and 45% were women. Forty-four patients (12.8%) harbored FLT3-ITD mutation, 42 patients (12.2%) harbored NPM1 mutation, and 27 patients (7.8%) harbored DNMT3A R882 mutations (23 R882H mutation and 4 R882C mutation). Interestingly, patients positive for NPM1 mutations were usually accompanied by positive mutations for FLT3-ITD and DNMT3A R882 mutations (Supplementary Table S2). Thirty-nine patients (11.3%) received hematopoietic cell transplantation (HCT). For patients received HCT, survival data was censored at the time of undergoing transplantation.
Table 1: The baseline characteristics in entire AML patients
Characteristics |
Available patients (n,%) |
Median (Range) |
Missing (n,%) |
|---|---|---|---|
Sex |
|
|
|
Male |
188(54.7) |
— |
|
Female |
156(45.3) |
— |
|
Age, years |
344(100.0) |
42(14-79) |
|
BSA |
302(87.8) |
1.6(1.2-2.1) |
42(12.2) |
BMI |
302(87.8) |
22.4(15.6-35.1) |
42(12.2) |
FAB classification |
|
|
|
M0 |
1(0.3) |
— |
|
M1 |
20(5.8) |
— |
|
M2 |
165(48.0) |
— |
|
M4 |
50(14.5) |
— |
|
M5 |
63(18.3) |
— |
|
M6 |
5(1.5) |
— |
|
M7 |
1(0.3) |
— |
|
Other AML |
37(10.8) |
— |
|
Mix(AML+ALL) |
2(0.6) |
— |
|
FLT3-ITD mutation |
|
|
2(0.6) |
Positive |
44(12.8) |
— |
|
Negative |
298(86.6) |
— |
|
NPM1 mutation |
|
|
2(0.6) |
Positive |
42(12.2) |
— |
|
Negative |
300(87.2) |
— |
|
Risk stratificationsa |
|
|
|
Low risk |
62(18.0) |
— |
|
Intermediate risk |
168(48.8) |
— |
|
High risk |
114(33.2) |
— |
|
DNMT3A R882 mutation |
|
|
|
Positive |
27(7.8) |
— |
|
Negative |
317(92.2) |
— |
|
Chemotherapy regimens |
|
|
|
Mitoxantrone + AraC |
88(25.6) |
— |
|
AraC + ACLA + G-CSF |
107(31.1) |
— |
|
Daunorubicin + AraC |
42(12.2) |
— |
|
Idarubicin + AraC |
47(13.7) |
— |
|
THP + AraC |
29(8.4) |
— |
|
Others |
31(9.0) |
— |
|
BM blasts, % |
329(95.6) |
70.0(2.0-99.0) |
15(4.4) |
PB blasts, % |
277(80.5) |
60.0(6.0-97.0) |
67(19.5) |
WBC, ×109/L |
344(100.0) |
14.5(0.5-436.2) |
|
RBC, ×1012/L |
343(99.8) |
2.2(0.6-4.9) |
1(0.2) |
Hemoglobin, g/L |
344(100.0) |
71.0(27.0-155.0) |
|
Platelets, ×109/L |
344(100.0) |
30.0(3.0-546.0) |
|
Neutrophils, ×109/L |
342(99.4) |
2.5(0.0-250.0) |
2(0.6) |
LDH, U/L |
336(97.7) |
355.6(44.7-7286.0) |
8(2.3) |
Treatment response |
|
|
|
Early death |
33(9.6) |
— |
|
CR after 1st course |
107(31.1) |
— |
|
CR after 2nd course |
98(28.5) |
— |
|
Partial remission |
37(10.8) |
— |
|
No remission |
69(20.1) |
— |
|
CR after 2 courses |
|
|
|
Yes |
205(59.6) |
|
|
No |
139(40.4) |
|
|
HCT |
|
|
|
Yes |
39(11.3) |
— |
|
No |
305(88.7) |
— |
|
aThe risk stratification based on validated cytogenetics and molecular data was classified according to NCCN Guidelines Version 1.2015 Acute Myeloid Leukemia.
Abbreviations: BSA, body surface area; BMI, Body Mass Index; FAB, French-American-British (classification); R882, Arginine 882; FLT3-ITD, fms-like tyrosine kinase3-intenal tandem duplication; BM, bone marrow; PB, peripheral blood; WBC, white blood cell; RBC, red blood cell; LDH, lactate dehydrogenase; AraC, cytarabine; ACLA, aclarubicin; G-CSF, granulocyte-colony stimulating factor; THP, pirarubicin; CR, complete remission; HCT, hematopoietic cell transplantation.
Genotype Information and linkage disequilibrium analysis
Eleven SNPs (rs11695471, rs2289195, rs734693, rs2276598, rs1465825, rs7590760, rs13401241, rs7581217, rs749131, rs41284843 and rs7560488) at DNMT3A gene locus were chosen from a Deep Catalog of Human Genetic Variation database--1000 Genomes Project (available at http://www.1000genomes.org/home). Table 2 showed detailed information regarding these 11 SNPs. Rs2276598 (Leu 422 Leu) and rs41284843 (Pro 9 Pro) were synonymous variations, while the other nine SNPs were intron variations. The minor allele frequencies (MAF) of DNMT3A polymorphisms in this study were comparable to those of Han population in Southern China (CHS) from 1000 Genomes Project. Hardy–Weinberg equilibrium of the genotype distribution was discovered for 11 SNPs in this study (Table 2). The linkage disequilibrium (LD) of DNMT3A polymorphisms at 11 loci were shown in Figure 1. There was no obvious LD between each pair of the 11 SNPs (r2 < 0.8).
Table 2: Characteristics of 11 selected tag SNPs from DNMT3A
SNP |
Position |
MAF (refa) |
Alleles |
Variation |
Pb |
|---|---|---|---|---|---|
rs11695471 |
2:25234839 |
0.083(0.10) |
T:A |
Intron22 |
1.0000 |
rs2289195 |
2:25240614 |
0.268(0.29) |
G:A |
Intron18 |
0.5917 |
rs734693 |
2:25241002 |
0.345(0.37) |
C:T |
Intron17 |
0.3794 |
rs2276598 |
2:25246633 |
0.330(0.30) |
C:T |
Exon10 (Leu422Leu) |
0.4515 |
rs1465825 |
2:25255584 |
0.373(0.38) |
T:C |
Intron6 |
0.6282 |
rs7590760 |
2:25266314 |
0.301(0.34) |
G:C |
Intron6 |
0.0287 |
rs13401241 |
2:25295601 |
0.310(0.24) |
C:A |
Intron3 |
1.0000 |
rs7581217 |
2:25302075 |
0.423(0.45) |
C:T |
Intron2 |
0.3109 |
rs749131 |
2:25306755 |
0.359(0.30) |
A:C |
Intron2 |
0.3950 |
rs41284843 |
2:25313958 |
0.086(0.09) |
G:A |
Exon2(Pro9Pro) |
0.5456 |
rs7560488 |
2:25345952 |
0.178(0.12) |
T:C |
5-Flanking |
0.4571 |
aref indicates the MAF value of CHS from 1000 Genomes.
bP value for Hardy–Weinberg equilibrium analysis.
Abbreviations: SNP, single nucleotide polymorphism; MAF, minor allele frequency.
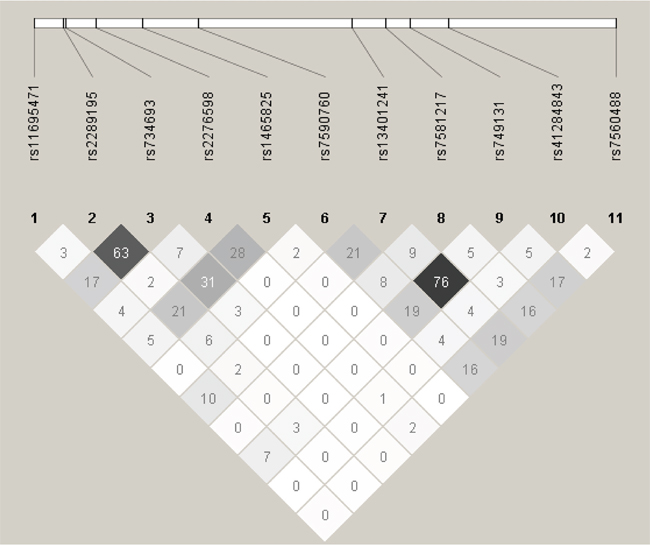
Figure 1: Pairwise linkage disequilibrium of the 11 DNMT3A SNPs in the studied AML patients. The linkage disequilibrium (LD) of DNMT3A polymorphisms at 11 loci were analyzed using Haploview software and indicated by r2 value (%).
Associations between individual SNPs and chemosensitivity status in AML patients
Univariate analysis
Associations of DNMT3A SNPs with chemosensitivity after first or second induction chemotherapy were shown in Table 3. Chemosensitivity was defined as good response (GR) or poor response (PR) in this study. Significant associations between three SNPs (rs11695471, rs2276598, rs734693) and good response (GR) ratio were observed. Rs734693 CT/TT genotypes showed a trend of lower GR ratio (P=0.093, dominant model) with an OR of 1.50 (95% CI: 0.93-2.41). A higher GR ratio was observed for rs2276598 CT/TT genotypes (P=0.046, dominant model) with an OR of 0.62 (95% CI: 0.39-0.99). Rs11695471 AT/AA genotypes also showed a lower GR ratio (P=0.041, dominant model) with an OR of 1.84 (95% CI: 1.02-3.33). No significant associations of other 8 SNPs with GR ratio were found (Table 3).
Table 3: Univariate and Logistic regression analysis of associations between DNMT3A SNPs and chemosensitivity in entire AML patients
SNP |
Genotype |
N |
P |
OR (95% CI)a |
P a |
|---|---|---|---|---|---|
rs11695471 |
TA+AA v TT |
344 |
0.041 |
2.012 (1.060-3.819) |
0.033 |
rs2289195 |
GA+AA v GG |
343 |
0.584 |
0.755 (0.265-2.153) |
0.599 |
rs734693 |
CT+TT v CC |
344 |
0.093 |
1.691 (1.004-2.849) |
0.048 |
rs2276598 |
CT+TT v CC |
337 |
0.046 |
0.587 (0.347-0.995) |
0.048 |
rs1465825 |
TC+CC v TT |
342 |
0.654 |
0.960 (0.570-1.618) |
0.878 |
rs7590760 |
GC+CC v GG |
342 |
0.553 |
0.910 (0.410-2.020) |
0.816 |
rs13401241 |
CA+AA v CC |
342 |
0.927 |
1.013 (0.606-1.694) |
0.959 |
rs7581217 |
CT+TT v CC |
342 |
0.534 |
0.901 (0.527-1.540) |
0.702 |
rs749131 |
AC+CC v AA |
341 |
0.876 |
1.035 (0.615-1.741) |
0.898 |
rs41284843 |
GA+AA v GG |
340 |
0.382 |
1.362 (0.721-2.574) |
0.342 |
rs7560488 |
TC+CC v TT |
337 |
0.916 |
1.017 (0.589-1.758) |
0.951 |
NOTE: Bold font indicates statistical significance.
aAdjusted for gender, age and FAB classification.
Abbreviations: SNP, single nucleotide polymorphism; OR, odds ratio; CI, confidence interval.
The characteristics of AML patients between or among genotypes of rs11695471, rs2276598, rs734693 and other eight SNPs were described (data not shown). These characteristics, such as gender, age or FAB classification at diagnosis showed no significant difference between or among genotypes of these SNPs.
Logistic regression analysis
In order to ascertain independent predictive factors for chemosensitivity, except for DNMT3A SNP rs11695471, rs2276598 and rs734693, covariates including gender, age and FAB classification were included in a logistic regression model. The rationale for including these variables was that they showed significant association with chemosensitivity in univariate analysis (P<0.05, data not shown). Results of logistic regression analysis showed that rs2276598 CT/TT carriers showed a significantly higher GR ratio (OR=0.587; 95% CI: 0.347-0.995; P=0.048] as compared to the CC homozygotes, whereas rs734693 CT/TT carriers (OR=1.691; 95% CI: 1.004-2.849; P=0.048) and rs11695471 AT/AA carriers (OR=2.012; 95% CI: 1.060-3.819; P=0.033) showed markedly lower GR ratio as compared with rs734693 CC genotype and rs11695471 TT genotype, respectively (Table 3).
A combined genotype analysis of rs11695471, rs2276598 and rs734693 for AML chemosensitivity
DNMT3A polymorphisms (rs11695471, rs2276598 and rs734693) were shown to exert individual effect on chemosensitivity in AML, but their combined effect had not been elucidated yet. Hence, a DNMT3A combined genotype score model was generated and results of combined genotype analysis were presented in Table 4. Chi-square test indicated that a defined favorable response group (composite score 1, 2 or 3) had a higher GR ratio compared with a defined unfavorable response group (composite score 0) (favorable group: 73.1% vs unfavorable group: 47.1%, P=0.002). Adjusted for covariates including gender, age and FAB classification, binary logistic regression analysis indicated that the defined favorable response group from a DNMT3A combined genotype score model had a higher GR ratio (OR=0.242; 95% CI: 0.109-0.536; P=0.001).
Table 4: Combined analysis of the association between rs11695471, rs2276598 and rs734693 genotypes and AML chemosensitivity
Composite Scorea |
GR ratio |
N |
P |
OR (95% CI)b |
Pb |
|---|---|---|---|---|---|
Composite score 0 |
47.1% |
34 |
1.00 (reference) |
|
|
composite score 1 |
70.6% |
85 |
0.017 |
0.315 (0.130-0.763) |
0.011 |
composite score 2 |
73.6% |
121 |
0.004 |
0.248 (0.105-0.587) |
0.001 |
composite score 3 |
76.3% |
97 |
0.002 |
0.244 (0.101-0.588) |
0.002 |
composite score1/2/3 |
73.1% |
305 |
0.002 |
0.242 (0.109-0.536) |
0.001 |
NOTE: Bold font indicates statistical significance.
Abbreviations: SNP, single nucleotide polymorphism; GR, good response; OR, odd ratio; CI, confidence interval.
aA DNMT3A combined genotype score model was created by figuring up the genotyped data of SNPs rs11695471, rs734693 and rs2276598. Score 1 indicated favorable alleles (i.e. rs11695471 TT genotype, or rs734693 CC genotype, or rs2276598 CT/TT genotype for chemosensitivity) and score 0 indicated unfavorable alleles (i.e. rs11695471 TA/AA genotype, or rs734693 CT/TT genotype, or rs2276598 CC genotype for chemosensitivity). After adding up these scores, four composite score groups were generated: composite score 0, composite score 1, composite score 2 and composite score 3. Two chemotherapy response groups were also defined: unfavorable response (composite score 0) and favorable response (composite score 1/2/3).
b Adjusted for gender, age and FAB classification.
DNMT3A genetic polymorphisms predicted disease prognosis in AML patients
Survival analysis verified that the DNMT3A R882 mutation AML patients showed markedly shorter median OS and DFS compared with R882 wildtype patients (R882 mutation: 212 days vs R882 wildtype: 560 days, P<0.0001 for OS; R882 mutation: 228 days vs R882 wildtype: 469 days, P=0.004 for DFS, Figure 2G, 2H).
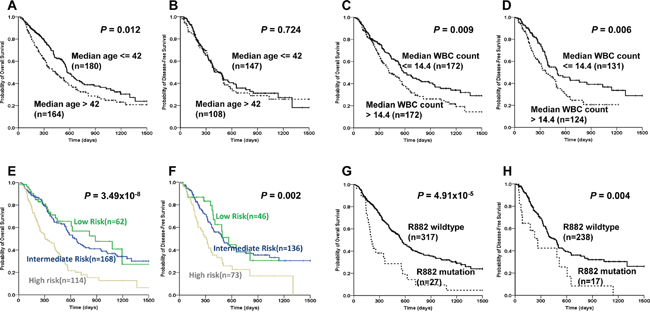
Figure 2: Associations of age, WBC count, risk stratification and DNMT3A R882 mutation with disease survivals in AML patients. Kaplan-Meier evaluation of OS A. and DFS B. based on the median age in the entire AML cohort. Kaplan-Meier evaluation of OS C. and DFS D. based on median WBC count in the entire AML cohort. Kaplan-Meier evaluation of OS E. and DFS F. based on risk stratification in the entire AML cohort. Kaplan-Meier evaluation of OS G. and DFS H. based on the status of DNMT3A R882 mutation in the entire AML cohort. Patients failed to achieve CR were omitted in the DFS analysis.
Univariate analysis for disease prognosis
Survival analysis showed that the six SNPs were significantly or in trend associated with prognosis of AML patients (Figure 3). Rs2289195 AA genotype showed significantly longer median OS and DFS versus GG/GA genotype group (P=0.005 for OS; P=0.013 for DFS, Figure 3A, 3B); rs734693 TT genotype had significantly longer OS and marginally longer median DFS compared with CC/CT genotype (TT: 1119 days vs CT/CC: 529 days, P=0.033 for OS; TT: 1035 days vs CT/CC: 393 days, P=0.071 for DFS, Figure 3C, 3D); and rs2276598 CT/TT genotype had a marginally longer DFS compared with CC genotype (CT/TT: 494 days vs CC: 360 days, P=0.058, Figure 3J). While rs1465825 TT genotype showed a markedly longer median OS and a in trend longer median DFS versus TC/CC genotype (TT: 622 days vs TC/CC: 456 days, P=0.044 for OS; TT: 491 days vs TC/CC: 370 days, P=0.104 for DFS, Figure 3E, 3F); rs11695471 TT genotype showed in trend longer median OS and DFS versus TA/AA genotype (TT: 548 days vs TA/AA: 379 days, P=0.088 for OS, TT: 484 days vs TA/AA: 330 days, P=0.169 for DFS, Figure 3G, 3H); and rs7590760 GG/GC genotype had a in trend longer DFS compared with CC genotype (GG/GC: 469 days vs CC: 320 days, P=0.136, Figure 3L). Other SNPs (Supplementary Figure S2) showed no associated with median OS and DFS.
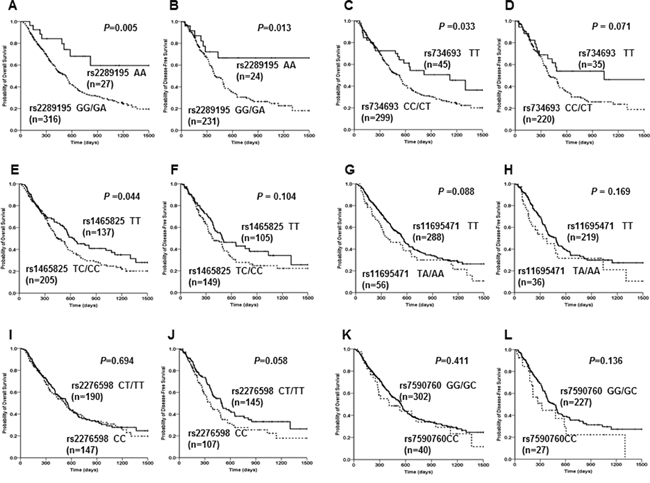
Figure 3: Associations of DNMT3A rs2289195, rs7346693, rs1465825, rs11695471, rs2276598 and rs7590760 polymorphisms with disease survivals in AML patients. Kaplan-Meier evaluation of OS A. and DFS B. based on the rs2289195 genotypes in the AML cohort. Kaplan-Meier evaluation of OS C. and DFS D. based on the rs7346693 genotypes in the AML cohort; Kaplan-Meier evaluation of OS E. and DFS F. based on the rs1465825 genotypes in the AML cohort; Kaplan-Meier evaluation of OS G. and DFS H. based on the rs11695471 genotypes in the AML cohort. Kaplan-Meier evaluation of OS I. and DFS J. based on the rs2276598 genotypes in the AML cohort; Kaplan-Meier evaluation of OS K. and DFS L. based on the rs7590760 genotypes in the AML cohort. Patients failed to achieve CR were omitted in the DFS analysis.
The characteristics of AML patients between genotype groups of DNMT3A rs11695471, rs2289195, rs734693, rs2276598, rs1465825 rs7590760 and other five SNPs were described (data not shown). The characteristics, such as gender, age, WBC count, BM blasts percentage, chemotherapy regimen, risk stratification or FAB classification at diagnosis showed no significant difference among the genotypes of all SNPs associated with OS or DFS.
Multivariate analysis for disease prognosis
For the multivariate survival analysis, four well-established prognostic factors [27, 49–53]: age, WBC count, risk stratification and DNMT3A R882 mutation status were also considered for OS as these factors affected prognosis of AML significantly in univariate analysis (Figure 2). While WBC count, the risk stratification and DNMT3A R882 mutation status were also considered for DFS, because age was not significantly associated with DFS in univariate analysis (Figure 2B). It is noteworthy that the risk stratification was based on validated cytogenetics and molecular data, which included FLT3-ITD and NPM1 mutation status in our study. Our results indicated that AML patients with FLT3-ITD mutation had a significantly shorter median survival than those without this mutation (P=0.002 for OS; P=0.023 for DFS, Figure S3A, S3B). NPM1 mutated patients also had a significantly shorter median OS and DFS (P<0.05, Figure S3C, S3D). Results of multivariate survival analysis with individual DNMT3A SNPs were shown in Table 5. Rs2289195 G>A polymorphism showed as a favorable prognostic predictor independently for both OS and DFS (HR=0.434, 95% CI: 0.213-0.887, P=0.022 for OS; HR=0.468, 95% CI: 0.217-1.008, P=0.052 for DFS). Rs2276598 C>T polymorphism was an independent favorable predictor for DFS (HR=0.680, 95% CI: 0.480-0.963, P=0.030), while rs11695471 T>A polymorphism was an independent poor predictor for OS (HR=1.453, 95% CI: 1.010-2.093, P=0.044).
Table 5: Multivariate analysis of association between each individual DNMT3A SNPs and disease prognosis of AML patients
Endpoint |
SNP |
Model |
N |
HR (95% CI) |
P a |
|---|---|---|---|---|---|
OS |
rs11695471 |
Dominant |
|
|
|
|
|
TT |
288 |
1.00 (reference) |
|
|
|
TA/AA |
56 |
1.453 (1.010-2.093) |
0.044 |
|
rs2289195 |
Recessive |
|
|
|
|
|
GG/GA |
316 |
1.00 (reference) |
|
|
|
AA |
27 |
0.434 (0.213-0.887) |
0.022 |
|
rs734693 |
Recessive |
|
|
|
|
|
CC/CT |
299 |
1.00 (reference) |
|
|
|
TT |
45 |
0.692 (0.436-1.098) |
0.116 |
|
rs2276598 |
Dominant |
|
|
|
|
|
CC |
147 |
1.00 (reference) |
|
|
|
CT/TT |
190 |
0.891 (0.664-1.194) |
0.439 |
|
rs1465825 |
Dominant |
|
|
|
|
|
TT |
137 |
1.00 (reference) |
|
|
|
TC/CC |
205 |
1.280 (0.951-1.724) |
0.103 |
|
rs7590760 |
Recessive |
|
|
|
|
|
GG/GC |
302 |
1.00 (reference) |
|
|
|
CC |
40 |
0.997 (0.645-1.540) |
0.988 |
DFS |
rs11695471 |
Dominant |
|
|
|
|
|
TT |
219 |
1.00 (reference) |
|
|
|
TA/AA |
36 |
1.484 (0.911-2.419) |
0.111 |
|
rs2289195 |
Recessive |
|
|
|
|
|
GG/GA |
231 |
1.00 (reference) |
|
|
|
AA |
24 |
0.468 (0.217-1.008) |
0.052 |
|
rs734693 |
Recessive |
|
|
|
|
|
CC/CT |
220 |
1.00 (reference) |
|
|
|
TT |
35 |
0.707 (0.410-1.221) |
0.212 |
|
rs2276598 |
Dominant |
|
|
|
|
|
CC |
107 |
1.00 (reference) |
|
|
|
CT/TT |
145 |
0.680 (0.480-0.963) |
0.030 |
|
rs1465825 |
Dominant |
|
|
|
|
|
TT |
105 |
1.00 (reference) |
|
|
|
TC/CC |
149 |
1.256 (0.881-1.792) |
0.207 |
|
rs7590760 |
Recessive |
|
|
|
|
|
GG/GC |
227 |
1.00 (reference) |
|
|
|
CC |
27 |
1.186 (0.693-2.033) |
0.533 |
aAdjusted for age, WBC count, risk stratification and R882 mutation (OS); and for WBC count, risk stratification and R882 mutation (DFS).
Bold font indicates statistical significance.
Abbreviations: SNP, single nucleotide polymorphism; HR, hazard ratio; CI, confidence interval; OS, Overall survival; DFS, disease-free survival.
Furthermore, the prognostic significance of all 6 SNPs was analyzed simultaneously in a multivariate analysis model. After adjusting for age, WBC count, risk stratification and DNMT3A R882 mutational status, the results showed that only the rs2289195 G>A SNP had an independently favorable prognosis effect on OS; after adjusting by WBC count, risk stratification and DNMT3A R882 mutational status, both rs2289195 G>A and rs2276598 C>T polymorphisms appeared to be favorable prognostic predictors for DFS independently. However, other DNMT3A polymorphisms did not show association with prognosis indicated by either OS or DFS (Table 6).
Table 6: Multivariate analysis of association between DNMT3A polymorphisms and disease prognosis of AML patients
Endpoint |
Variables in the model |
HR |
95% CI |
P a |
|---|---|---|---|---|
OS |
Age: >42 vs ≤42 |
1.457 |
1.087-1.953 |
0.012 |
|
WBC: >14.43 vs ≤14.43 |
1.328 |
0.987-1.788 |
0.061 |
|
Risk stratification |
|
|
|
|
Low-risk vs Intermediate-risk |
0.807 |
0.494-1.320 |
0.393 |
|
High-risk vs Intermediate-risk |
2.067 |
1.509-2.831 |
6.13x10-6 |
|
R882 mutation: Yes vs No |
1.942 |
1.239-3.040 |
0.004 |
|
Rs2289195: AA vs GG/GA |
0.439 |
0.215-0.898 |
0.024 |
DFS |
WBC: >14.43 vs ≤14.43 |
1.492 |
1.040-2.141 |
0.030 |
|
Risk stratification |
|
|
|
|
Low-risk vs Intermediate-risk |
0.811 |
0.466-1.410 |
0.458 |
|
High-risk vs Intermediate-risk |
1.602 |
1.086-2.363 |
0.017 |
|
R882 mutation: Yes vs No |
2.066 |
1.174-3.636 |
0.012 |
|
Rs2276598: CT/TT vs CC |
0.657 |
0.463-0.933 |
0.019 |
|
Rs2289195: AA vs GG/GA |
0.450 |
0.208-0.970 |
0.042 |
Abbreviations: HR, hazard ratio; CI, confidence interval; OS, Overall survival; DFS, disease-free survival.
Bold font indicates statistical significance.
A combined genotype analysis of rs11695471, rs2289195 and rs2276598 for AML survival
As rs11695471, rs2289195 and rs2276598 showed associations with AML prognosis individually, DNMT3A combined genotype score model was created by figuring up the genotype data of rs11695471, rs2289195 and rs2276598. Survival analysis of combined genotype showed that a defined favorable prognosis group (including composite score 2 plus composite score 3) showed a in trend longer median OS and a significantly longer median DFS than a defined unfavorable prognosis group (including composite score 0 plus composite score 1) (favorable prognosis group: 569 days vs unfavorable prognosis group: 488 days, P=0.126 for OS; favorable prognosis group: 508 days vs unfavorable prognosis group: 330 days, P=0.004 for DFS, Figure 4). After adjusting for age, WBC count, risk stratification and DNMT3A R882 mutational status (OS), and for WBC count, risk stratification and DNMT3A R882 mutational status (DFS), the results of multivariate survival analysis indicated that the unfavorable prognosis group (composite score 0 plus composite score 1) appeared to be an independent predicting factor for shorter OS and DFS (HR=1.346, 95% CI: 1.003-1.805, P=0.047 for OS; HR =1.717, 95% CI: 1.212-2.432, P=0.002 for DFS, Table 7).
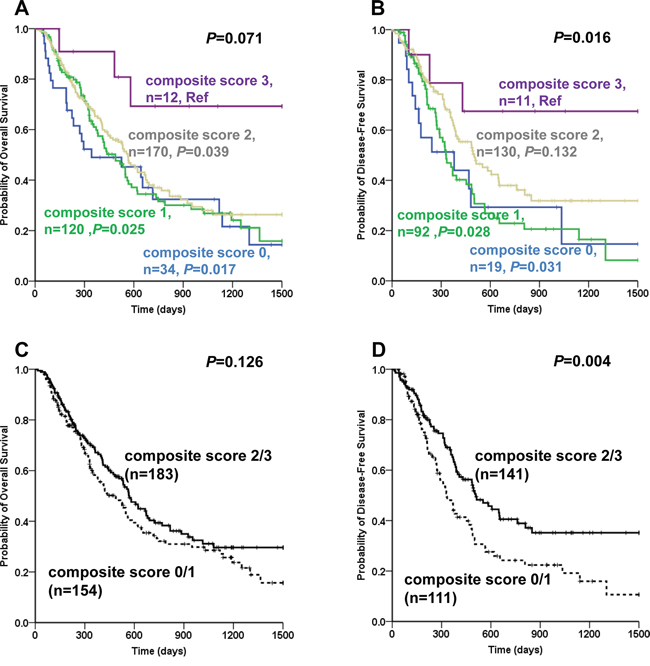
Figure 4: Combined genotype analysis of association between DNMT3A SNPs and disease prognosis in AML patients. Kaplan-Meier evaluation of OS A, C. and DFS B, D. based on the composite score group of rs11695471, rs2289195 and rs2276598 in the entire AML cohort; Patients failed to achieve CR were omitted in the DFS analysis.
Note: A DNMT3A polymorphism combined genotype score model was created by figuring up the genotyped data of SNPs rs11695471, rs2289195 and rs2276598. Score 1 indicated favorable alleles (i.e. rs11695471 TT genotype, or rs2289195 AA genotype, or rs2276598 CT/TT genotype for prognosis) and score 0 indicated unfavorable alleles (i.e. rs11695471 TA/AA genotype, or rs2289195 GG/GA genotype, or rs2276598 CC genotype for prognosis). After adding up these scores, four score groups were generated: composite score 0, composite score 1, composite score 2 and composite score 3. Two prognosis groups were also defined: unfavorable prognosis (composite score 0/1) and favorable prognosis (composite score 2/3).
Table 7: The potential association between DNMT3A genetic polymorphisms and AML prognosis by multivariate combined genotype analysis
Endpoint |
Composite Scorea |
N |
HR (95% CI) |
P b |
|---|---|---|---|---|
OS |
composite score 3 |
12 |
1.00 (reference) |
|
|
composite score 2 |
170 |
2.639 (0.828-8.418) |
0.101 |
|
composite score 1 |
120 |
3.218 (1.004-10.311) |
0.049 |
|
composite score 0 |
34 |
3.841 (1.152-12.806) |
0.028 |
|
composite score 2/3 |
183 |
1.00 (reference) |
|
|
composite score 0/1 |
154 |
1.346 (1.003-1.805) |
0.047 |
DFS |
composite score 3 |
11 |
1.00 (reference) |
|
|
composite score 2 |
130 |
1.984 (0.618-6.372) |
0.250 |
|
composite score 1 |
92 |
3.139 (0.974-10.112) |
0.055 |
|
composite score 0 |
19 |
3.854 (1.092-13.598) |
0.036 |
|
composite score 2/3 |
141 |
1.00 (reference) |
|
|
composite score 0/1 |
111 |
1.717 (1.212-2.432) |
0.002 |
NOTE: Bold font indicates statistical significance.
Abbreviations: SNP, single nucleotide polymorphism; HR, hazard ratio; CI, confidence interval; OS, Overall survival; DFS, disease-free survival.
a A DNMT3A combined genotype score model was created by figuring up the genotyped data of SNPs rs11695471, rs2276598 and rs2289195. Score 1 indicated favorable alleles (i.e. rs11695471 TT genotype, or rs2276598 CT/TT genotype, or rs2289195 AA genotype for prognosis) and score 0 indicated unfavorable alleles (i.e. rs11695471 TA/AA genotype, or rs2276598 CC genotype, or rs2289195 GG/GA genotype for prognosis). After adding up these scores, four score groups were generated: composite score 0, composite score 1, composite score 2 and composite score 3. Two prognosis groups were also defined: unfavorable prognosis (composite score 0/1) and favorable prognosis (composite score 2/3).
b Adjusted for age, WBC count, risk stratification and DNMT3A R882 mutation (OS); and for WBC count, risk stratification and DNMT3A R882 mutation (DFS).
DNMT3A mutation/SNP may reduce DNMT3A mRNA expression in AML patients
DNMT3A mRNA levels were detected using semi-quantitative reverse-transcriptase PCR method in PBMCs from 29 healthy volunteers and 74 unselected de novo AML patient samples with known CR status, DNMT3A mutation (R882 mutation, n=9) and DNMT3A SNP status. As shown in Figure 5A, PBMCs DNMT3A mRNA levels were markedly higher in AML patients than those in healthy volunteers (P<0.001). In order to assess potential influence of DNMT3A mRNA levels on complete remission (CR), 61 AML patients were divided into good response (GR) group and poor response (PR) group. As shown in Figure 5B, no difference in PBMCs DNMT3A mRNA levels was observed between GR and PR patients (P=0.414). Mean DNMT3A mRNA levels in DNMT3A R882 wildtype AML patients (n=64) was 2.06 times that of patients with R882 mutation (n=9, P=0.043, Figure 5C). Mean DNMT3A mRNA levels in patients with rs7590760 GG/GC genotype (n=66) was 2.38 times that of rs7590760 CC genotype (n=5, P=0.098, Figure 5D). No difference in DNMT3A mRNA expression among genotypes of other DNMT3A polymorphisms was observed in AML patients (data not shown).
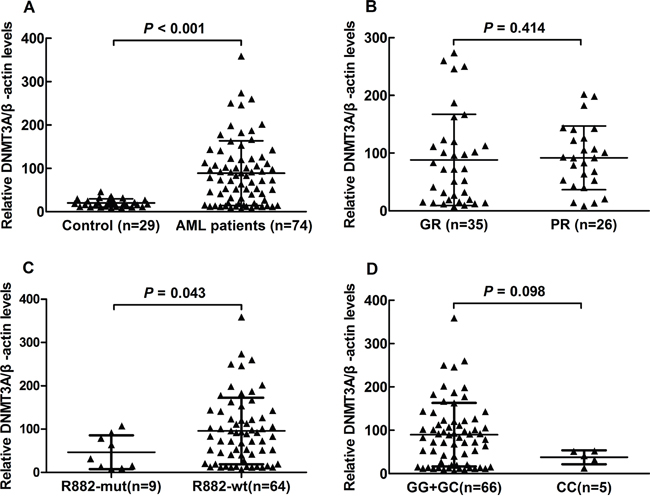
Figure 5: DNMT3A mRNA levels in AML patients with different CR status, R882 mutation status and rs7590760 genotypes. A. Comparison of DNMT3A mRNA levels in AML patients and healthy volunteers. B. Comparison of DNMT3A mRNA levels in the AML patients with good response (GR) or poor response (PR). C. Comparison of DNMT3A mRNA levels in AML patients with or without R882 mutation. D. Comparison of DNMT3A mRNA levels in AML patients with rs7590760 GG/GC genotype and rs7590760 CC genotype. Each triangle represents one patient and the number of patients in each group is shown. Horizontal lines indicate the mean DNMT3A/β-actin ratio. Relative expression levels in each patient were determined in duplicates and normalized to β-actin. Relative expression was calculated by 2−ΔCt.
Lower DNMT3A levels predicted an inferior prognosis in AML patients from a TCGA dataset
Survival analysis from a TCGA data set (seen in PATIENTS AND METHODS) found that the AML patients with lower DNMT3A expression showed significantly shorter median OS than those with higher DNMT3A expression (lower DNMT3A: 335 days vs higher DNMT3A: 822 days, P<0.001, Figure 6).
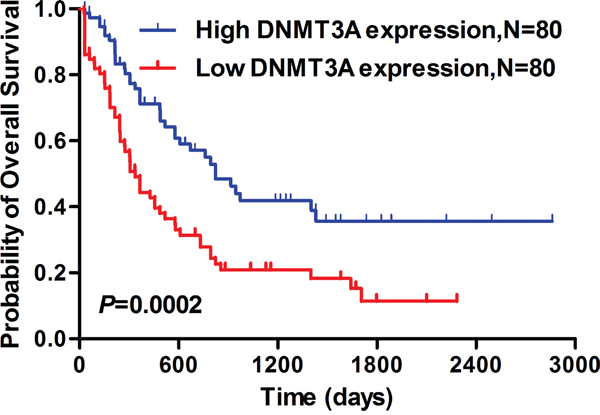
Figure 6: Lower DNMT3A mRNA expression predicted significantly shorter median overall survival in AML patients. Kaplan-Meier comparison of overall survival in AML patients with low DNMT3A expression and high DNMT3A expression.
To sum up, our results indicated that rs11695471, rs734693 and rs2276598 were significantly associated with chemosensitivity in AML patients, while rs11695471, rs2289195, and rs2276598 can independently predict AML patients’ disease prognosis.
DISCUSSION
To our knowledge, this study firstly identified a strong association between DNMT3A genetic polymorphisms and chemosensitivity and prognosis in AML patients. We observed that rs11695471, rs734693 and rs2276598 were significantly associated with chemosensitivity, while rs11695471, rs2289195 and rs2276598 had significant associations with the prognosis (OS or DFS) in AML patients. Further, DNMT3A combined genotype score model (created by figuring up data from rs11695471, rs734693 and rs2276598 genotypes for chemosensitivity; rs11695471, rs2289195, rs2276598 genotypes for prognosis) also showed similar results in our study population. In agreement with previous investigations [9, 11, 21, 23, 27], we confirmed that DNMT3A R882 mutations predicted a significantly poor prognosis in our AML patients.
For the multivariate survival analysis, the prognostic effect of DNMT3A SNPs was adjusted by four well-established prognostic factors including age, WBC count, risk stratification of disease and DNMT3A R882 mutation status. In our study, FLT3-ITD and NPM1 mutations were considered in the risk stratification according to NCCN Guidelines (Version 1.2015 Acute Myeloid Leukemia), we included risk stratification but not mutational status of FLT3-ITD and NPM1 as individual variables in our multivariate analysis. In agreement with previous reports [54–56], FLT3-ITD predicted a significantly poor prognosis in our AML patients. However, patients positive for NPM1 mutation had a significantly shorter median survival, though some researchers reported that NPM1 mutation conferred a favorable prognosis in AML patients with younger age or normal karyotype [57, 58]. This disagreement may be explained by the significant co-occurrence of NPM1 and FLT3-ITD mutations, and prognosis of patients carrying both mutations was similar with that of patients with FLT3-ITD mutation alone [57–59]. In consistent with previous reports [23, 60, 61], we observed that NPM1 mutations occurred concurrently with FLT3-ITD and DNMT3A R882 mutation in our samples. Besides, Boissel N et al [62] showed no significant difference in AML prognosis between NPM-mutated and -nonmutated patients with normal karyotype.
Hayette S et al [63] reported that DNMT3B overexpression may serve as a poor prognostic marker in AML, and DNMT3B expression was inversely correlated with DNMT3A level. Therefore, wondering whether the outcome effect of DNMT3A mutation/SNP on AML could act through affecting DNMT3A mRNA expression, we determined DNMT3A transcript levels in PBMCs from de novo AML patients. Consistent with previous reports [64, 65], we observed that AML patients showed significantly increased DNMT3A mRNA level than healthy volunteers, which indicated that high DNMT3A expression might contribute to the pathogenesis of leukemia [64]. Though evidence has shown that DNMT3A mutation can reduce the enzyme activity [9, 11, 66, 67], influence of R882 mutation on DNMT3A mRNA expression has not been described yet. Here we found that AML patients with R882 mutation showed significantly lower DNMT3A mRNA level. Fortunately, homozygote mutation of rs7590760 G>C polymorphism which could predicted a trend of shorter OS and DFS in AML, also showed a trend of decreased DNMT3A mRNA levels in our study. Furthermore, survival analysis indicated that lower DNMT3A expression displayed significantly shorter median OS in AML patients with data from a TCGA data set.
Although there is no report regarding association between DNMT3A polymorphisms and the outcome of AML patients, we speculate the possible mechanisms underline association between DNMT3A SNPs and chemosensitivity or disease prognosis may be explained by change in DNMT3A mRNA expression. Herein we found that AML patients positive for R882 mutation showed significantly lower DNMT3A mRNA levels. And also, rs7590760 G>C rare homozygotes presented a trend of decreased DNMT3A mRNA levels as compared to carriers of the rs7590760 G wildtype allele. In our study, the significant SNPs rs11695471, rs2289195, rs734693, rs2276598, rs1465825 and rs7590760 in univariate analysis were located in introns or synonymous, pair-wise LD analysis showed that r2 ranges from 0 to 0.63 for these SNPs. For the three significant SNPs (rs11695471, rs2289195 and rs2276598) associated with AML prognosis in multivariate analysis, the r2 ranges from 0.02 to 0.04. Therefore, the positive association may not be explained by LD between these SNPs (Figure 1). Of note, the 6 significant SNPs appeared to cluster at 3’ region of the gene. Evidence has shown that variants in 3’ UTR may change miRNAs targeting and subsequent change in mRNA expression. For instance, Cheng et al [68]reported that a polymorphic nucleotide T deletion (rs34351976) in NPM1 3’UTR could lead to the formation of an illegitimate miR-337-5p binding site and thus predicted an adverse prognosis in AML. Similarly, there may exist SNP(s) located in DNMT3A 3’-UTR which were in high LD with the positively associated SNPs. Of course, additional studies are required to determine the functional SNP(s) at the locus.
Certainly, several limitations of our study are unneglectable. First, though risk factors such as gender, age, blood routine indexes (clinical factors), karyotype, risk stratification (cytogenetics factor) and somatic mutations (including FLT3-ITD, NPM1 and DNMT3A) were adjusted during data analysis, other important risk factors, such as CEBPA, KIT and IDH1/2 mutations, which may also contribute to the prognosis of AML [22, 69–72], were ignored in this study. Second, the number of samples for mutant homozygote of some DNMT3A SNPs was relatively small because of the low MAF values, which may have led to limited statistical power. Third, only the genetic model (dominant or recessive model) which had statistical significance was chosen to evaluate the association between each SNP and AML outcomes in our study. Finally, this study required confirmation in independent studies with more sufficient power to detect single and combined genotype effects of DNMT3A genetic polymorphisms.
In summary, the current study firstly indicated that (i) there was strong association between DNMT3A genetic polymorphisms and the treatment outcomes in AML patients, and (ii) an available score model evaluating multiple polymorphisms in DNMT3A could predict chemosensitivity or prognosis in AML patients, and (iii) especially, a strategy estimating multiple noncoding polymorphisms in epigenetic modification related genes, such as DNMT3A, could predict the treatment outcomes in AML patients, which may improve rational design for more precise therapeutic regimens and prognostic stratification.
PATIENTS AND METHODS
Patients and treatment
A prospective analysis of 352 Chinese Han AML patients with available whole peripheral blood or bone marrow samples who hospitalized in Xiangya Hospital of Central South University between June 2011 and July 2015 was evaluated. FAB classification was applied to diagnose AML patients in our study [73]. Patients with the FAB-M3 subtype were excluded because of the occurrence of the specific fusion gene PML-RARa caused by translocation t(15;17) which makes the treatment effect better than other types of AML. And patients with serious diseases or other cancers, therapy-related AML or secondary AML arising from a prior myelodysplastic syndrome were also excluded. Clinical study admission (registration number: ChiCTR-PPC-14005297) was approved. Informed consent has been obtained for this study, which was approved by the Ethics Committee of Institute of Clinical Pharmacology of Central South University (registerNo.CTXY-120025-2) and was implemented in strict accordance with the Declaration of Helsinki. All patients received standard induction chemotherapy consisted of recommended dose anthracyclines for three consecutive days and 100-200 mg/m2 cytarabine for seven consecutive days. Genomic DNA for polymorphism analyses was available for 344 cases from this cohort. Once CR was achieved, the patients were treated with hematopoietic cell transplantation (HCT) or cycles of high-dose cytarabine, followed by periods of consolidation chemotherapy consisted of anthracyclines and cytarabine.
Definition and end points
Major end points were defined as described previously [50, 74, 75]. Complete remission (CR) following one or two cycles of induction chemotherapy was used to evaluate chemosensitivity. CR was characterized by < 5% bone marrow (BM) blast cells and the absolute counts of neutrophils >1 x 109/L and platelets >100 x 109/L in addition to no evidence of extramedullary leukemia. Partial remission was characterized by: achieving all criteria of CR in hematology; BM blasts declining to 5% to 25% or to 5% if Auer rod positive. In this study, chemosensitivity groups were defined: the relatively good response (GR) group (including receiving CR or partial remission) and the relatively poor response (PR) group (except for CR or partial remission). Disease-free survival (DFS) was equal to the time interval from the first CR to relapse or death for any cause. Overall survival (OS) was equal to the time length from diagnosis to death for any cause. For patients without relapse or death event by the end of the study follow-up, survival end points were censored by the last follow-up. Patients failed to follow-up were excluded in the study. The risk stratification including FLT3-ITD status and NPM1 mutation status based on validated cytogenetics and molecular data was classified according to NCCN Guidelines Version 1.2015 Acute Myeloid Leukemia [76].
FLT3-ITD and PCR-gel-electrophoresis
The internal tandem duplications of FLT3 (FLT3-ITD) were tested as described [55, 77]. Briefly, the fragment from exon 14 to exon 15 of FLT3 were amplified using previously described primers (Supplementary Table S1) [78] from genomic DNA by the following steps: denaturing at 94°C for 45 seconds, annealing at 57°C for 45 seconds, and extension at, 72°C for 30 seconds, for 35 cycles on a Master cycler® pro (Eppendorf, Germany) including a pre-denaturation at 94°C and a post-extension at 72°C for 5 minutes, respectively. PCR products were electrophoresed on 2% agarose gels and visualized under UV light with Gel Red (red) staining. A length of 329 base pair (bp) was produced from wild-type alleles. PCR bands larger than 329 bp were seen as FLT3-ITD. Repeat analyses were done on samples with additional bands.
Detection of DNMT3A R882 and NPM1 mutations by PCR-pyrosequencing
Detection of DNMT3A R882 (exon 23) and NPM1 (exon 12) mutations was carried out by PCR and analyzed with Pyro Mark Q24 Advance software (Qiagen, Germany) similarly as previous described [79–81]. DNA separated from whole peripheral blood or bone marrow of patients was amplified with biotinylated primer to harvest single-stranded templates for pyrosequencing. PCR was performed on a Master cycler® pro (Eppendorf, Germany). Next, the single-stranded templates were analyzed using Pyro Mark Q24 station (Qiagen, Germany) according to instrument instructions. Pyrosequencing assay involves 3 primers: forward, reverse and sequencing (Supplementary Table S1). Necessarily, Sanger sequencing was also applied to verify the reliability of the pyrosequencing result. The results displayed that two R882 variants (R882H and R882C) could be detected simultaneously, and the results of pyrosequencing and Sanger sequencing were consistent completely.
SNPs selection
The full sequence of the human DNMT3A gene investigated in this study included all exons and introns (109.63 kb), which were pinpointed to chromosome 2, position 25232961-25342590 (www.ncbi.nlm.nih.gov/genbank/). Genetic variation data for the entire DNMT3A gene were obtained from the 1000 Genomes Project (http://www.1000genomes.org/) for 105 Han Chinese individuals from Southern China (CHS). From this database, a total of 300 SNPs in DNMT3A have been identified in the CHS population, and 121 were common SNPs with minor allele frequencies (MAF) ≥0.05. We identify 44 SNPs (tagging SNPs) that efficiently tags 112 of the SNPs using Haploview, version 4.2 (Broad Institute of MIT and Harvard, Cambridge, M A, USA), a software package that provides computation of linkage disequilibrium (LD) statistics [82]. r2>0.8 indicates a strong LD between two pairwise SNPs. Based on consulting literature material, we selected 15 tagging SNPs that best represented potential functional SNPs (priority: preferring missense to synonymous codon to intron variant). Finally, we successfully genotyped 10 tagging SNPs using A Mass ARRAY system from Sequenom (Supplementary Figure S1). DNMT3A SNP (rs7560488) was also selected according to the publication by Wu H et al [31], which showed an increased risk of gastric cancer for the C allele (variant) by modulating promoter activity in Chinese population.
Genotyping of DNMT3A polymorphisms
For DNMT3A genotyping, whole peripheral blood genomic DNA was extracted from AML patients using a E.Z.N.A.® SQ Blood DNA Kit II (Omega Bio-Tek, USA). A Mass ARRAY system from Sequenom was used to genotype DNMT3A 10 SNPs. The assays were validated by Polymerase chain reaction–restriction fragment length polymorphism (PCR-RFLP) or by Sanger sequencing (ABI 3730XL DNA Analyzer, Applied Biosystems, USA) or by pyrosequencing methods for 5% of the samples, and the results were 100% concordant. DNMT3A rs7560488 was genotyped by Sanger sequencing. The forward and reverse primer pairs of PCR, and the following extending primer or sequencing primer were shown in Supplementary Table S1.
DNMT3A combined genotype score model
A DNMT3A combined genotype score model was created by figuring up the genotyped data of multi-SNPs. Score 1 indicated favorable genotype(s) (i.e. rs11695471 TT genotype, or rs734693 CC genotype, or rs2276598 CT/TT genotype for chemosensitivity; rs2289195 AA genotype, or rs11695471 TT genotype, or rs2276598 CT/TT genotype for prognosis) and score 0 indicated unfavorable genotype(s) (i.e. rs11695471 TA/AA genotype, or rs734693 CT/TT genotype, or rs2276598 CC genotype for chemosensitivity; rs2289195 GG/GA genotype, or rs11695471 TA/AA genotype, or rs2276598 CC genotype for prognosis). After adding up the score of every SNP, four score groups were generated: composite score 0, composite score 1, composite score 2 and composite score 3. And then another two chemosensitivity groups were defined: the relatively unfavorable response group (composite score 0) and the relatively favorable response group (including composite score 1, composite score 2 and composite score 3). Two prognosis groups were also defined: the relatively unfavorable prognosis group (including composite score 0 and composite score 1) and the relatively favorable prognosis group (including composite score 2 and composite score 3).
Analysis of DNMT3A mRNA expression
We unbiasedly selected 74 cases’ bone marrow (BM) or peripheral blood mononuclear cells (PBMCs) from this cohort, whose baseline characteristics were shown in Supplementary Table S3. DNMT3A mRNA expression was also assessed by RT-qPCR and compared with 29 normal peripheral blood samples obtained from healthy donors who had no prior history of malignancy. Mononuclear cells were separated by centrifuging (400 × g) and layering for 30 minutes at 18°C~25°C mixing equal volume of Histopaque-1077 (Sigma–Aldrich, St. Louis, MO). Total RNA was purified with RNAiso Plus reagents (Takara-Bio, Dalian, China) and stored at −80 °C until analysis. The cDNA was reverse transcribed using Prime Script™ RT reagent kit (Takara-Bio, Dalian, China). QRT-PCR was performed on a LC480® real-time PCR system (Roche, USA) using the real-time PCR quant kit (Takara, Biotechnology, Dalian, China). The forward primer of DNMT3A was 5’-AGTAC GACGACGA CGGCTA-3’, the reverse primer was 5’-CACACTCCACGCAAAAGCAC-3’. The β-actin mRNA expression was used as an endogenous control.
DNMT3A expression Data extract from TCGA database
A dataset including the information on DNMT3A mRNA expression and overall survival (OS) of AML (n=173 for LAML gene expression, pancan normalized) was obtained from the UCSC Cancer Genomics Browser of TCGA (https://genome-cancer.soe.ucsc.edu/). DNMT3A expression was divided into lower DNMT3A expression group and higher DNMT3A expression group according to a cut-off value of median DNMT3A expression. Then, we evaluated the association of DNMT3A expression with overall survival (OS) in this AML cohort. Thirteen samples were not included in survival analysis due to lacking of survival information.
Statistical analysis
Chi-square (χ2) test was used for determining whether DNMT3A 11 SNPs agreed with Hardy–Weinberg equilibrium. The linkage disequilibrium (LD) of genotypes was analyzed by Haploview software (available at http://www.broad.mit.edu/mpg/haploview). The frequencies of the genotypes were compared using χ2 test or Fisher’s test in baseline’ characteristics of patients or status of achieving CR after one or two cycles of induction chemotherapy. A logistic regression model was used to ascertain independent predictive factors for chemosensitivity, including gender, age, FAB classification, rs11695471, rs2276598 and rs734693. The OR and 95% CI indicated the relative risk level. Comparisons of continuous variables between or among genotype groups were performed using nonparametric tests. The significance of observed differences in proportions was tested by the χ2 test and Fisher’s exact test when data were spare. The primary analysis was performed on OS. Sensitivity analyses were conducted on CR rate and DFS. The median OS and DFS evaluations were performed by Kaplan-Meier curve. Cox’s proportional hazard model was used to identify independent prognostic predictors for OS and DFS using a backward conditional procedure. The following variables were included for the analyses: rs11695471, rs2289195, rs7346693, rs2276598, rs1465825 and rs7590760, age, WBC count, risk stratification and DNMT3A R882 mutational status. The HR and 95% CI indicated the relative risk level for survivals. Two-sided P <0.05 represented a statistical significance of difference. SPSS 18.0 was used for statistical analyses.
ACKNOWLEDGMENTS AND FUNDINGS
This project was supported by Chinese National Science Foundation (No. 81422052), Special topic of the major subject of national science and technology (2012ZX09509-107), and Hunan Provincial Natural Science Foundation of China (13JJ1010).
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
REFERENCES
1. Dohner H, Weisdorf DJ, Bloomfield CD. Acute Myeloid Leukemia. N Engl J Med. 2015; 373:1136-1152.
2. Buchi F, Masala E, Rossi A, Valencia A, Spinelli E, Sanna A, Gozzini A, Santini V. Redistribution of H3K27me3 and acetylated histone H4 upon exposure to azacitidine and decitabine results in de-repression of the AML1/ETO target gene IL3. Epigenetics. 2014; 9:387-395.
3. Gamazon ER, Lamba JK, Pounds S, Stark AL, Wheeler HE, Cao X, Im HK, Mitra AK, Rubnitz JE, Ribeiro RC, Raimondi S, Campana D, Crews KR, et al. Comprehensive genetic analysis of cytarabine sensitivity in a cell-based model identifies polymorphisms associated with outcome in AML patients. Blood. 2013; 121:4366-4376.
4. Emadi A, Karp JE. The clinically relevant pharmacogenomic changes in acute myelogenous leukemia. Pharmacogenomics. 2012; 13:1257-1269.
5. Zheng R, Zeng H, Zhang S, Chen T, Chen W. National estimates of cancer prevalence in China, 2011. Cancer Lett. 2015.
6. Mehdipour P, Santoro F, Minucci S. Epigenetic alterations in acute myeloid leukemias. FEBS J. 2015; 282:1786-1800.
7. Becker H, Bloomfield CD. Acute myeloid leukaemia in 2012: En route to improved treatment options. Nat Rev Clin Oncol. 2013; 10:76-79.
8. Bolli N, Manes N, McKerrell T, Chi J, Park N, Gundem G, Quail MA, Sathiaseelan V, Herman B, Crawley C, Craig JI, Conte N, Grove C, et al. Characterization of gene mutations and copy number changes in acute myeloid leukemia using a rapid target enrichment protocol. Haematologica. 2015; 100:214-222.
9. Ley TJ, Ding L, Walter MJ, McLellan MD, Lamprecht T, Larson DE, Kandoth C, Payton JE, Baty J, Welch J, Harris CC, Lichti CF, Townsend RR, et al. DNMT3A mutations in acute myeloid leukemia. N Engl J Med. 2010; 363:2424-2433.
10. Marcucci G, Metzeler KH, Schwind S, Becker H, Maharry K, Mrozek K, Radmacher MD, Kohlschmidt J, Nicolet D, Whitman SP, Wu YZ, Powell BL, Carter TH, et al. Age-related prognostic impact of different types of DNMT3A mutations in adults with primary cytogenetically normal acute myeloid leukemia. J Clin Oncol. 2012; 30:742-750.
11. Yan XJ, Xu J, Gu ZH, Pan CM, Lu G, Shen Y, Shi JY, Zhu YM, Tang L, Zhang XW, Liang WX, Mi JQ, Song HD. Exome sequencing identifies somatic mutations of DNA methyltransferase gene DNMT3A in acute monocytic leukemia. Nat Genet. 2011; 43:309-315.
12. Renneville A, Abdelali RB, Chevret S, Nibourel O, Cheok M, Pautas C, Dulery R, Boyer T, Cayuela JM, Hayette S, Raffoux E, Farhat H, Boissel N, et al. Clinical impact of gene mutations and lesions detected by SNP-array karyotyping in acute myeloid leukemia patients in the context of gemtuzumab ozogamicin treatment: results of the ALFA-0701 trial. Oncotarget. 2014; 5:916-932. doi: 10.18632/oncotarget.1536.
13. Hou HA, Kuo YY, Liu CY, Chou WC, Lee MC, Chen CY, Lin LI, Tseng MH, Huang CF, Chiang YC, Lee FY, Liu MC, Liu CW, et al. DNMT3A mutations in acute myeloid leukemia: stability during disease evolution and clinical implications. Blood. 2012; 119:559-568.
14. Kronke J, Bullinger L, Teleanu V, Tschurtz F, Gaidzik VI, Kuhn MW, Rucker FG, Holzmann K, Paschka P, Kapp-Schworer S, Spath D, Kindler T, Schittenhelm M, et al. Clonal evolution in relapsed NPM1-mutated acute myeloid leukemia. Blood. 2013; 122:100-108.
15. Tan Y, Liu H, Chen S. Mutant DNA methylation regulators endow hematopoietic stem cells with the preleukemic stem cell property, a requisite of leukemia initiation and relapse. Front Med. 2015.
16. Walter MJ, Ding L, Shen D, Shao J, Grillot M, McLellan M, Fulton R, Schmidt H, Kalicki-Veizer J, O'Laughlin M, Kandoth C, Baty J, Westervelt P, et al. Recurrent DNMT3A mutations in patients with myelodysplastic syndromes. Leukemia. 2011; 25:1153-1158.
17. Ploen GG, Nederby L, Guldberg P, Hansen M, Ebbesen LH, Jensen UB, Hokland P, Aggerholm A. Persistence of DNMT3A mutations at long-term remission in adult patients with AML. Br J Haematol. 2014; 167:478-486.
18. El Ghannam D, Taalab MM, Ghazy HF, Eneen AF. DNMT3A R882 mutations in patients with cytogenetically normal acute myeloid leukemia and myelodysplastic syndrome. Blood Cells Mol Dis. 2014; 53:61-66.
19. Lin J, Yao DM, Qian J, Chen Q, Qian W, Li Y, Yang J, Wang CZ, Chai HY, Qian Z, Xiao GF, Xu WR. Recurrent DNMT3A R882 mutations in Chinese patients with acute myeloid leukemia and myelodysplastic syndrome. PLoS One. 2011; 6:e26906.
20. Im AP, Sehgal AR, Carroll MP, Smith BD, Tefferi A, Johnson DE, Boyiadzis M. DNMT3A and IDH mutations in acute myeloid leukemia and other myeloid malignancies: associations with prognosis and potential treatment strategies. Leukemia. 2014; 28:1774-1783.
21. Kao HW, Liang DC, Kuo MC, Wu JH, Dunn P, Wang PN, Lin TL, Shih YS, Liang ST, Lin TH, Lai CY, Lin CH, Shih LY. High frequency of additional gene mutations in acute myeloid leukemia with MLL partial tandem duplication: DNMT3A mutation is associated with poor prognosis. Oncotarget. 2015; 6:33217-33225. doi: 10.18632/oncotarget.5202.
22. Patel JP, Gonen M, Figueroa ME, Fernandez H, Sun Z, Racevskis J, Van Vlierberghe P, Dolgalev I, Thomas S, Aminova O, Huberman K, Cheng J, Viale A, et al. Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. N Engl J Med. 2012; 366:1079-1089.
23. Thol F, Damm F, Ludeking A, Winschel C, Wagner K, Morgan M, Yun H, Gohring G, Schlegelberger B, Hoelzer D, Lubbert M, Kanz L, Fiedler W, et al. Incidence and prognostic influence of DNMT3A mutations in acute myeloid leukemia. J Clin Oncol. 2011; 29:2889-2896.
24. Ribeiro AF, Pratcorona M, Erpelinck-Verschueren C, Rockova V, Sanders M, Abbas S, Figueroa ME, Zeilemaker A, Melnick A, Lowenberg B, Valk PJ, Delwel R. Mutant DNMT3A: a marker of poor prognosis in acute myeloid leukemia. Blood. 2012; 119:5824-5831.
25. Ostronoff F, Othus M, Ho PA, Kutny M, Geraghty DE, Petersdorf SH, Godwin JE, Willman CL, Radich JP, Appelbaum FR, Stirewalt DL, Meshinchi S. Mutations in the DNMT3A exon 23 independently predict poor outcome in older patients with acute myeloid leukemia: a SWOG report. Leukemia. 2013; 27:238-241.
26. Hou HA, Lin CC, Chou WC, Liu CY, Chen CY, Tang JL, Lai YJ, Tseng MH, Huang CF, Chiang YC, Lee FY, Kuo YY, Lee MC, et al. Integration of cytogenetic and molecular alterations in risk stratification of 318 patients with de novo non-M3 acute myeloid leukemia. Leukemia. 2014; 28:50-58.
27. Yuan XQ, Peng L, Zeng WJ, Jiang BY, Li GC, Chen XP. DNMT3A R882 Mutations Predict a Poor Prognosis in AML: A Meta-Analysis From 4474 Patients. Medicine (Baltimore). 2016; 95:e3519.
28. Fan H, Liu D, Qiu X, Qiao F, Wu Q, Su X, Zhang F, Song Y, Zhao Z, and Xie W. A functional polymorphism in the DNA methyltransferase-3A promoter modifies the susceptibility in gastric cancer but not in esophageal carcinoma. BMC Med. 2010; 8:12.
29. Yang XX, He XQ, Li FX, Wu YS, Gao Y, Li M. Risk-association of DNA methyltransferases polymorphisms with gastric cancer in the southern chinese population. Int J Mol Sci. 2012; 13:8364-8378.
30. Wu Q, Lu S, Wang L, Hu J, Qiao F, Qiu X, Zhao C, Lao Y, Song Y, Fan H. DNMT3A rs36012910 A>G polymorphism and gastric cancer susceptibility in a Chinese population. Mol Biol Rep. 2012; 39:10949-10955.
31. Wu H, Zhang K, Gong P, Qiao F, Wang L, Cui H, Sui X, Gao J, Fan H. A novel functional TagSNP Rs7560488 in the DNMT3A1 promoter is associated with susceptibility to gastric cancer by modulating promoter activity. PLoS One. 2014; 9:e92911.
32. Zhao Z, Li C, Song Y, Wu Q, Qiao F, Fan H. Association of the DNMT3A -448A>G polymorphism with genetic susceptibility to colorectal cancer. Oncol Lett. 2012; 3:450-454.
33. Zhao C, Yan F, Wu H, Qiao F, Qiu X, Fan H. DNMT3A -448A>G polymorphism and the risk for hepatocellular carcinoma. Biomed Rep. 2013; 1:664-668.
34. Kelemen LE, Sellers TA, Schildkraut JM, Cunningham JM, Vierkant RA, Pankratz VS, Fredericksen ZS, Gadre MK, Rider DN, Liebow M, Goode EL. Genetic variation in the one-carbon transfer pathway and ovarian cancer risk. Cancer Res. 2008; 68:2498-2506.
35. Mostowska A, Sajdak S, Pawlik P, Lianeri M, Jagodzinski PP. DNMT1, DNMT3A and DNMT3B gene variants in relation to ovarian cancer risk in the Polish population. Mol Biol Rep. 2013; 40:4893-4899.
36. Kullmann K, Deryal M, Ong MF, Schmidt W, Mahlknecht U. DNMT1 genetic polymorphisms affect breast cancer risk in the central European Caucasian population. Clin Epigenetics. 2013; 5:7.
37. Sun MY, Yang XX, Xu WW, Yao GY, Pan HZ, Li M. Association of DNMT1 and DNMT3B polymorphisms with breast cancer risk in Han Chinese women from South China. Genet Mol Res. 2012; 11:4330-4341.
38. Inoue-Choi M, Nelson HH, Robien K, Arning E, Bottiglieri T, Koh WP, Yuan JM. Plasma S-adenosylmethionine, DNMT polymorphisms, and peripheral blood LINE-1 methylation among healthy Chinese adults in Singapore. BMC Cancer. 2013; 13:389.
39. Tajuddin SM, Amaral AF, Fernandez AF, Rodriguez-Rodero S, Rodriguez RM, Moore LE, Tardon A, Carrato A, Garcia-Closas M, Silverman DT, Jackson BP, Garcia-Closas R, Cook AL, et al. Genetic and non-genetic predictors of LINE-1 methylation in leukocyte DNA. Environ Health Perspect. 2013; 121:650-656.
40. Haggarty P, Hoad G, Horgan GW, Campbell DM. DNA methyltransferase candidate polymorphisms, imprinting methylation, and birth outcome. PLoS One. 2013; 8:e68896.
41. Agundez JA. Polymorphisms of human N-acetyltransferases and cancer risk. Curr Drug Metab. 2008; 9:520-531.
42. Dong LM, Potter JD, White E, Ulrich CM, Cardon LR, Peters U. Genetic susceptibility to cancer: the role of polymorphisms in candidate genes. JAMA. 2008; 299:2423-2436.
43. Bolufer P, Barragan E, Collado M, Cervera J, Lopez JA, Sanz MA. Influence of genetic polymorphisms on the risk of developing leukemia and on disease progression. Leuk Res. 2006; 30:1471-1491.
44. McLeod HL, Krynetski EY, Relling MV, Evans WE. Genetic polymorphism of thiopurine methyltransferase and its clinical relevance for childhood acute lymphoblastic leukemia. Leukemia. 2000; 14:567-572.
45. Saad ED, Abbruzzese JL. Prognostic stratification in UPC: a role for assessing the value of conventional-dose and high-dose chemotherapy for unknown primary carcinoma. Crit Rev Oncol Hematol. 2002; 41:205-211.
46. Marcucci G, Mrozek K, Bloomfield CD. Molecular heterogeneity and prognostic biomarkers in adults with acute myeloid leukemia and normal cytogenetics. Curr Opin Hematol. 2005; 12:68-75.
47. Meyer SC, Levine RL. Translational implications of somatic genomics in acute myeloid leukaemia. Lancet Oncol. 2014; 15:e382-394.
48. Domingues P, Gonzalez-Tablas M, Otero A, Pascual D, Ruiz L, Miranda D, Sousa P, Goncalves JM, Lopes MC, Orfao A, Tabernero MD. Genetic/molecular alterations of meningiomas and the signaling pathways targeted. Oncotarget. 2015; 6:10671-10688. doi: 10.18632/oncotarget.3870.
49. Kuhnl A, Valk PJ, Sanders MA, Ivey A, Hills RK, Mills KI, Gale RE, Kaiser MF, Dillon R, Joannides M, Gilkes A, Haferlach T, Schnittger S, et al. Downregulation of the Wnt inhibitor CXXC5 predicts a better prognosis in acute myeloid leukemia. Blood. 2015; 125:2985-2994.
50. Mortland L, Alonzo TA, Walter RB, Gerbing RB, Mitra AK, Pollard JA, Loken MR, Hirsch B, Raimondi S, Franklin J, Pounds S, Cao X, Rubnitz JE, et al. Clinical significance of CD33 nonsynonymous single-nucleotide polymorphisms in pediatric patients with acute myeloid leukemia treated with gemtuzumab-ozogamicin-containing chemotherapy. Clin Cancer Res. 2013; 19:1620-1627.
51. Parkin B, Ouillette P, Yildiz M, Saiya-Cork K, Shedden K, Malek SN. Integrated genomic profiling, therapy response, and survival in adult acute myelogenous leukemia. Clin Cancer Res. 2015; 21:2045-2056.
52. Groschel S, Schlenk RF, Engelmann J, Rockova V, Teleanu V, Kuhn MW, Eiwen K, Erpelinck C, Havermans M, Lubbert M, Germing U, Schmidt-Wolf IG, Beverloo HB, et al. Deregulated expression of EVI1 defines a poor prognostic subset of MLL-rearranged acute myeloid leukemias: a study of the German-Austrian Acute Myeloid Leukemia Study Group and the Dutch-Belgian-Swiss HOVON/SAKK Cooperative Group. J Clin Oncol. 2013; 31:95-103.
53. Niederwieser C, Kohlschmidt J, Volinia S, Whitman SP, Metzeler KH, Eisfeld AK, Maharry K, Yan P, Frankhouser D, Becker H, Schwind S, Carroll AJ, Nicolet D, et al. Prognostic and biologic significance of DNMT3B expression in older patients with cytogenetically normal primary acute myeloid leukemia. Leukemia. 2015; 29:567-575.
54. Frohling S, Schlenk RF, Breitruck J, Benner A, Kreitmeier S, Tobis K, Dohner H, Dohner K. Prognostic significance of activating FLT3 mutations in younger adults (16 to 60 years) with acute myeloid leukemia and normal cytogenetics: a study of the AML Study Group Ulm. Blood. 2002; 100:4372-4380.
55. Kottaridis PD, Gale RE, Frew ME, Harrison G, Langabeer SE, Belton AA, Walker H, Wheatley K, Bowen DT, Burnett AK, Goldstone AH, Linch DC. The presence of a FLT3 internal tandem duplication in patients with acute myeloid leukemia (AML) adds important prognostic information to cytogenetic risk group and response to the first cycle of chemotherapy: analysis of 854 patients from the United Kingdom Medical Research Council AML 10 and 12 trials. Blood. 2001; 98:1752-1759.
56. Whitman SP, Maharry K, Radmacher MD, Becker H, Mrozek K, Margeson D, Holland KB, Wu YZ, Schwind S, Metzeler KH, Wen J, Baer MR, Powell BL, et al. FLT3 internal tandem duplication associates with adverse outcome and gene- and microRNA-expression signatures in patients 60 years of age or older with primary cytogenetically normal acute myeloid leukemia: a Cancer and Leukemia Group B study. Blood. 2010; 116:3622-3626.
57. Dohner K, Schlenk RF, Habdank M, Scholl C, Rucker FG, Corbacioglu A, Bullinger L, Frohling S, Dohner H. Mutant nucleophosmin (NPM1) predicts favorable prognosis in younger adults with acute myeloid leukemia and normal cytogenetics: interaction with other gene mutations. Blood. 2005; 106:3740-3746.
58. Schnittger S, Schoch C, Kern W, Mecucci C, Tschulik C, Martelli MF, Haferlach T, Hiddemann W, Falini B. Nucleophosmin gene mutations are predictors of favorable prognosis in acute myelogenous leukemia with a normal karyotype. Blood. 2005; 106:3733-3739.
59. Thiede C, Koch S, Creutzig E, Steudel C, Illmer T, Schaich M, Ehninger G. Prevalence and prognostic impact of NPM1 mutations in 1485 adult patients with acute myeloid leukemia (AML). Blood. 2006; 107:4011-4020.
60. Renneville A, Boissel N, Nibourel O, Berthon C, Helevaut N, Gardin C, Cayuela JM, Hayette S, Reman O, Contentin N, Bordessoule D, Pautas C, Botton S, et al. Prognostic significance of DNA methyltransferase 3A mutations in cytogenetically normal acute myeloid leukemia: a study by the Acute Leukemia French Association. Leukemia. 2012; 26:1247-1254.
61. Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med. 2013; 368:2059-2074.
62. Boissel N, Renneville A, Biggio V, Philippe N, Thomas X, Cayuela JM, Terre C, Tigaud I, Castaigne S, Raffoux E, De Botton S, Fenaux P, Dombret H, et al. Prevalence, clinical profile, and prognosis of NPM mutations in AML with normal karyotype. Blood. 2005; 106:3618-3620.
63. Hayette S, Thomas X, Jallades L, Chabane K, Charlot C, Tigaud I, Gazzo S, Morisset S, Cornillet-Lefebvre P, Plesa A, Huet S, Renneville A, Salles G, et al. High DNA methyltransferase DNMT3B levels: a poor prognostic marker in acute myeloid leukemia. PLoS One. 2012; 7:e51527.
64. Mizuno S, Chijiwa T, Okamura T, Akashi K, Fukumaki Y, Niho Y, Sasaki H. Expression of DNA methyltransferases DNMT1, 3A, and 3B in normal hematopoiesis and in acute and chronic myelogenous leukemia. Blood. 2001; 97:1172-1179.
65. Haferlach T, Kohlmann A, Wieczorek L, Basso G, Kronnie GT, Bene MC, De Vos J, Hernandez JM, Hofmann WK, Mills KI, Gilkes A, Chiaretti S, Shurtleff SA, et al. Clinical utility of microarray-based gene expression profiling in the diagnosis and subclassification of leukemia: report from the International Microarray Innovations in Leukemia Study Group. J Clin Oncol. 2010; 28:2529-2537.
66. Russler-Germain DA, Spencer DH, Young MA, Lamprecht TL, Miller CA, Fulton R, Meyer MR, Erdmann-Gilmore P, Townsend RR, Wilson RK, Ley TJ. The R882H DNMT3A mutation associated with AML dominantly inhibits wild-type DNMT3A by blocking its ability to form active tetramers. Cancer Cell. 2014; 25:442-454.
67. Gowher H, Loutchanwoot P, Vorobjeva O, Handa V, Jurkowska RZ, Jurkowski TP, Jeltsch A. Mutational analysis of the catalytic domain of the murine Dnmt3a DNA-(cytosine C5)-methyltransferase. J Mol Biol. 2006; 357:928-941.
68. Cheng CK, Kwan TK, Cheung CY, Ng K, Liang P, Cheng SH, Chan NP, Ip RK, Wong RS, Lee V, Li CK, Yip SF, Ng MH. A polymorphism in the 3'-untranslated region of the NPM1 gene causes illegitimate regulation by microRNA-337-5p and correlates with adverse outcome in acute myeloid leukemia. Haematologica. 2013; 98:913-917.
69. Cairoli R, Beghini A, Grillo G, Nadali G, Elice F, Ripamonti CB, Colapietro P, Nichelatti M, Pezzetti L, Lunghi M, Cuneo A, Viola A, Ferrara F, et al. Prognostic impact of c-KIT mutations in core binding factor leukemias: an Italian retrospective study. Blood. 2006; 107:3463-3468.
70. Schnittger S, Kohl TM, Haferlach T, Kern W, Hiddemann W, Spiekermann K, Schoch C. KIT-D816 mutations in AML1-ETO-positive AML are associated with impaired event-free and overall survival. Blood. 2006; 107:1791-1799.
71. Bienz M, Ludwig M, Leibundgut EO, Mueller BU, Ratschiller D, Solenthaler M, Fey MF, Pabst T. Risk assessment in patients with acute myeloid leukemia and a normal karyotype. Clin Cancer Res. 2005; 11:1416-1424.
72. Paschka P, Marcucci G, Ruppert AS, Mrozek K, Chen H, Kittles RA, Vukosavljevic T, Perrotti D, Vardiman JW, Carroll AJ, Kolitz JE, Larson RA, Bloomfield CD. Adverse prognostic significance of KIT mutations in adult acute myeloid leukemia with inv(16) and t(8;21): a Cancer and Leukemia Group B Study. J Clin Oncol. 2006; 24:3904-3911.
73. Bennett JM, Catovsky D, Daniel MT, Flandrin G, Galton DA, Gralnick HR, Sultan C. Proposed revised criteria for the classification of acute myeloid leukemia. A report of the French-American-British Cooperative Group. Ann Intern Med. 1985; 103:620-625.
74. Cheson BD, Bennett JM, Kopecky KJ, Buchner T, Willman CL, Estey EH, Schiffer CA, Doehner H, Tallman MS, Lister TA, Lo-Coco F, Willemze R, Biondi A, et al. Revised recommendations of the International Working Group for Diagnosis, Standardization of Response Criteria, Treatment Outcomes, and Reporting Standards for Therapeutic Trials in Acute Myeloid Leukemia. J Clin Oncol. 2003; 21:4642-4649.
75. Ho PA, Kuhn J, Gerbing RB, Pollard JA, Zeng R, Miller KL, Heerema NA, Raimondi SC, Hirsch BA, Franklin JL, Lange B, Gamis AS, Alonzo TA, et al. WT1 synonymous single nucleotide polymorphism rs16754 correlates with higher mRNA expression and predicts significantly improved outcome in favorable-risk pediatric acute myeloid leukemia: a report from the children's oncology group. J Clin Oncol. 2011; 29:704-711.
76. National Comprehensive Cancer Network. NCCN Clinical Practice Guidelines in Oncology: Acute Myeloid Leukemia (Version 1.2015). 2014 Dec 3.
77. Yin J, Xie X, Zhang F, Chen Z, Hu C, Su G, Liu H, Zheng Y, He C, Shen H, Qiu Q, He J, Pan Z, et al. Low frequency of mutations in Chinese with acute myeloid leukemia: Different disease or different aetiology? Leuk Res. 2015; 39:646-648.
78. Kiyoi H, Naoe T, Nakano Y, Yokota S, Minami S, Miyawaki S, Asou N, Kuriyama K, Jinnai I, Shimazaki C, Akiyama H, Saito K, Oh H, et al. Prognostic implication of FLT3 and N-RAS gene mutations in acute myeloid leukemia. Blood. 1999; 93:3074-3080.
79. Alderborn A, Kristofferson A, Hammerling U. Determination of single-nucleotide polymorphisms by real-time pyrophosphate DNA sequencing. Genome Res. 2000; 10:1249-1258.
80. Wan Z, Wang G, Li T, Xu B, Pei Q, Peng Y, Sun H, Cheng L, Zeng Y, Yang G, Zhu YS. Marked Alteration of Rosuvastatin Pharmacokinetics in Healthy Chinese with ABCG2 34G>A and 421C>A Homozygote or Compound Heterozygote. J Pharmacol Exp Ther. 2015; 354:310-315.
81. Fusco G, Amoroso MG, Gesualdi Montesano N, Viscardi M. Development of a pyrosequencing assay for the typing of alphaherpesviruses. MethodsX. 2015; 2:47-52.
82. Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005; 21:263-265.

