Introduction
Hepatocellular carcinoma (HCC) is a common cancer, with high morbidity and mortality, which places a heavy financial burden on sufferers, especially those in developing countries [1]. Despite developments in imaging, chemotherapy, interventional radiology, surgical techniques, and liver transplantation for HCC within the past two decades, the prognosis is still poor, and HCC remains the third most frequent cause of cancer-related deaths [2, 3]. At present, the best hope of improving the long-term survival of HCC patients is to detect the cancer at an early stage [4]. However, current noninvasive screening tools, such as serum α-fetoprotein and imaging (ultrasonography), are not highly sensitive, especially in the early diagnosis of HCC [5-7]. As with many other cancers, drug resistance, recurrence, and metastasis are the most important and difficult problems in the treatment of HCC [3].
Recently, tumor-derived exosomes (TEXs) have shown promise in the field of cancers [8]. The contents of TEXs, such as microRNA (miRNA) and oncoproteins, reflect the specific characteristics of their endosomal origin [9, 10]. TEXs are protected against degradation from enzymes (e.g., RNAse), as they are enclosed in a lipid bilayer [11]. Importantly, these enriched miRNAs and proteins in TEXs are selected, suggesting that the exosomal content may provide novel serological biomarkers for various types of cancer [12-14]. For example, research has suggested that glypican-1 could be used to detect early-stage pancreatic cancer [12] and that miR-21 and miR-93 could be used to detect early-stage HCC [13, 15]. In addition, TEXs can influence cancer progression and metastasis by the transfer of genetic materials between cells in the tumor microenvironment [16]. Thus, the evidence indicates that exosomes have various key roles in the development of cancer. An improved understanding of these roles may lead to the development of a powerful new strategy for preventing and treating HCC.
In this review, we summarize the roles and probable mechanisms of exosomes in HCC. In particular, considering the key role of miRNAs in cancer, we focus on exosomal miRNAs and their potential utility in HCC. The aim of the review is to illustrate the potential clinical applications of exosomes in the detection and treatment of HCC.
Generation of Exosomes in the Liver
As shown in Figure 1, exosomes are formed by endocytosis, exocytosis, protein transport, and protein sorting. Virus and transmembrane proteins are endocytosed and transported into early endosomes. Following, early endosomes adevelop into late endosomes. In late endosomes, intraluminal vesicles (ILVs) form through inward budding of endosomal membranes and finally result in a large multivesicular body (MVB). ILVs in a MVB released into the extracellular space are referred to as “exosomes,” which typically have a size of 30-120 nm [17, 18]. Various types of cells, particularly tumor cells, can secrete exosomes into the extracellular matrix after multiple intracellular vesicles fuse with the cell membrane [17, 18]. Exosomes are found not only in serum, plasma, and urine but also in other body fluids, such as amniotic fluid, ascites, nasal lavage fluid, breast milk, saliva, and cell culture media [9, 15, 19-21].
With regard to the liver, three main cell types release exosomes (Figure 1): hepatocytes, nonparenchymal immune cells (e.g., Kupffer cells, natural killer cells, T cells, and B cells), and nonparenchymal liver cells (e.g., liver stellate cells). Exosomes derived from different cells in the liver have different functions, for example, exosomes derived from T cells and B cells are important mediators of inflammation [22], and exosomes derived from hepatic stellate cells may be involved in fibrosis processes [23].
Functions of Exosomes in HCC
Exosomal miRNAs and HCC
Many studies have suggested that miRNAs participate in the development of HCC and that they may serve as potential prognostic or diagnostic markers for HCC [24-31]. The expression of the miRNAs miR-21, miR-221, and miR-222 is upregulated in HCC tissues [27-29], whereas the expression of miR-122a, miR-145, miR-199a, and miR-223 is downregulated, as compared to that of normal controls [27, 30, 32]. Recent studies suggest that exosomes may function as biological delivery vehicles for miRNAs [9, 33-35]. The expression of miRNAs in exosomes in HCC is shown in Table 1.
The miRNA of exosomes is different from that of their cells of origin. In miRNA expression profiling of Hep3B cells containing miRNA of exosomes derived from those cells, the expression levels of 25 miRNAs were significantly enriched in the exosomes, whereas those of 30 miRNAs were downregulated. Notably, 11 miRNAs were detected exclusively in the exosomes [9]. In another study of SMMC-7721 hepatocarcinoma cells, some miRNAs, such as miR-486-5p and miR-10b-5p, were highly abundant in exosomes but less abundant in cellular RNAs, and let-7d-5p, let-7b-5p, and let-7c-5p, which were highly abundant in cellular miRNAs, occurred only in very low numbers in exosomal RNAs [36]. These results point to selective enrichment of exosomes with specific miRNAs in HCC cells.
Exosomal miRNAs are transferable and functional in recipient cells. miR-122, a liver-specific miRNA that plays various roles in liver physiology and promotes the replication of the hepatitis C virus (HCV) [37], can be transferred via exosomes between two human hepatoma cell lines Huh7 and HepG2. When exosomal miR-122 released by Huh7 cells is transferred to HepG2 cells, it reduces the growth and proliferation of recipient cells [38]. Interestingly, exosomal miRNAs can also be transferred between different types of cells. Exosomes from adipose-derived mesenchymal stem cells (ADMSCs) can transfer miR-122 to HepG2 [35]. In addition, human and mouse liver cells, as well as primary human B cells, can exchange cellular endogenous miRNA, mediated by exosome shuffling [39]. miR-21, miR-192, and miR-221 in exosomes derived from colorectal cancer cell lines can be transferred to HepG2 cells and A549 lung cancer cells [33]. These findings indicate that miRNAs can shuttle between cells via exosomes, potentially promoting the invasion and metastasis of HCC.
Following the transfer of exosomal miRNAs to recipient cells, they regulate the target gene expression of recipient cells. miR-122-transfected ADMSCs can effectively package miR-122 into secreted exosomes and then deliver it to HCC cells, thereby altering the expression of miR-122 target genes, such as cyclin G1 and insulin-like growth factor receptor 1 in hepatoma cells [35]. In a previous study, the recurrence of HCC was higher in HCC liver transplantation patients with reduced expression of serum exosomal miR-718 [40]. The authors attributed the increased recurrence to miR-718 markedly suppressing the proliferation of Huh7 cells by upregulating the expression of the HOXB8 gene.
Table 1: Specific expression of miRNAs in exosomes derived from HCC
miRNAs |
Source of exosomes |
Source of compared |
Function |
Ref |
Upregulated miRNAs |
||||
miR-584 |
Hep3B cells |
Hep3B donor cells |
Target TAK1, enhance transformed cell |
[9] |
miR-517c |
||||
miR-378 |
||||
miR-520f |
||||
miR-142-5p |
||||
miR-451 |
||||
miR-518d |
||||
miR-215 |
||||
miR-376a* |
||||
miR-133b |
||||
miR-367 |
||||
miR-21 |
Serum |
CHB and healthy volunteers. |
Correlated with cirrhosis and tumor stage |
[15] |
miR-486-5p |
SMMC-7721 cells |
HCC cells |
Promote Proliferation and Metastasis |
[36] |
miR-10b-5p |
||||
miR-18a, |
Serum |
LC and CHB patients |
Novel serological biomarkers for HCC |
[13] |
miR-221 |
||||
miR-222 |
||||
miR-224 |
||||
Downregulated miRNAs |
||||
let-7d-5p |
SMMC-7721 cells |
HCC cells |
Promote Proliferation and Metastasis |
[36] |
let-7b-5p |
||||
let-7c-5p |
||||
miR-718 |
Serum |
HOXB8 |
Target HOXB8, suppresses cell proliferation |
[40] |
miR-101 |
Serum |
LC and CHB patients |
Novel serological biomarkers for HCC |
[13] |
miR-106b |
Serum |
CHB patients |
Novel serological biomarkers for HCC |
[13] |
miR-122 |
||||
miR-195 |
||||
CHB, Chronic Hepatitis B; LC, Live Cirrhosis;
Exosomal proteomes and HCC
The content of exosomal proteins of hepatic cancer cell lines differs greatly, depending on their metastatic potential (high vs. low). In a quantitative analysis, specific proteins enriched in hepatocyte-derived exosomes included caveolae (caveolin 1), early endosomes, endoplasmic reticulum proteins, peroxisomes and mitochondria [41, 42]. Using mass spectrometry analysis, 213 unique proteins were detected in HCC-derived exosomes [43]. In the same study, a proteomic analysis revealed that the MET protein, caveolins, and S100 family members were significantly enriched in exosomes secreted by highly malignant HCC cells. Another study confirmed that the abundance of exosomal S100 family members, S100-A8 and S100-A9, significantly increased in response to HBV transfection in Huh-7 cells [44]. However, as shown by exosomal proteome profiling, the expression of these proteins does not seem to be specific to HCC. For example, the S100 protein is found in exosomes secreted from breast cancer cells [45], caveolin 1 is found in glioma [46], and MET is found in colorectal cancer cells [10]. MET signaling, caveolins and S100 family members are known to have pivotal roles in tumor growth, angiogenesis, and metastasis [47-49]. Thus, these exosomal proteomes could be common targets for the treatment of various types of cancer.
RNA binding proteins (RBPs) act as miRNA carriers in cells [50]. Intriguingly, some RBPs, such as high-density lipoproteins and argonaute-2 (Ago2), are present in exosomes [51, 52].The involvement of lipid-mediated RNA loading in exosomes has been demonstrated [53]. ELAVL1 is a member of the ELAVL family of RBPs. It directly interacts with long noncoding RNA (lncRNA), a target of miRNA, in HCC cells [54]. Ahadi et al. identified ELAVL1 motifs on exosomal lncRNAs, suggesting that ELAVL1 may also stabilize the transcript [55]. In addition, Schilders et al. [56] demonstrated that M-phase phosphoprotein 6 is an exosome-associated RBP for the recruitment of the exosome to pre-rRNA. Thus, the literature suggests that miRNA loading of exosomes may be based on the interaction of the RBP-RNA complex. Research has also shown that replication of the HCV was associated with the Ago2-HSP90-miR-122 complex in exosomes isolated from HCV patients or HCV-infected Huh7.5 cells [51]. Thus, based on the current literature, RBPs not only mediate the loading of miRNAs into exosomes, they also play critical roles in stabilizing these miRNAs in exosomes.
Exosome-mediated invasion and metastasis
Tumor metastasis is a multistep process, which includes invasion, intravasation, and colonization of distal sites via the circulatory system [57]. The tumor microenvironment allows for tight integration of these processes [58]. Cancer-derived exosomes provide a means of cell-cell communication between cancer cells and may possibly recruit and reprogram the constituents of the tumor microenvironment [57]. The contents of exosomes, such as noncoding RNAs and proteins, may then influence cancer progression and metastasis [59].
Cellular TGF-beta-activated kinase 1 (TAK1) responds to environmental changes and intercellular modulation and is thought to contribute to the formation and growth of HCC [60]. In Hep3B cells, miRNAs are released into exosomes via a ceramide-dependent pathway, transferred to recipient cells, and then modulate the constitutive expression of TAK1 and downstream signaling associated with TAK1 in recipient cells [9]. Following the incubation of Hep3B and HepG2 with Hep3B-derived exosomes, the number of colonies of Hep3B and HepG2 cells in soft agar increasedFollowing ia, as well as the activity of caspase-3/7, indicating that exosomes have a positive effect on transformed cell growth and death in vitro [9]. CD90+ cancer stem cells are associated with metastasis and early recurrence in HCC [61, 62]. A previous study showed that CD90+ Huh7 cells derived from exosomes influenced the tumor microenvironment by promoting angiogenesis [63]. In that study, when endothelial cells were incubated with exosomes released by CD90+ Huh7 cells, the mRNA levels of the proangiogenic factor vascular endothelial growth factor (VEGF) and its receptor VEGF-R1 increased significantly in endothelial cells, in addition to adhesion between the CD90+ cells and endothelial cells. However, the aforementioned effects were not observed in exosomes released by Huh7 parental cells. Furthermore, H19, an lncRNA, was up-regulated (10-fold) in the exosomes derived from the CD90+ Huh7 cells compared with those derived from Huh7, suggesting that H19 lncRNA might stimulate angiogenesis and favor cell-cell interactions [63].
In common with noncoding RNAs, exosomal proteomes can promote the migration of cancer cells. Vasorin, a serum biomarker of hepatocarcinoma that is highly expressed in sera and tissues samples of hepatocarcinoma [64], is released by HepG2 cells in exosomes. Upon delivery of vasorin to human umbilical vein endothelial cells via exosomes, vasorin accelerates the migration of these cells [65]. In an immortalized hepatocyte line, the MET protein, caveolins, and S100 family members in HCC-derived exosomes triggered PI3K/AKT and MAPK signaling pathways and increased the secretion of matrix metalloproteinase-2 (MMP-2) and MMP-9, thereby facilitating the invasion activity of cancer cells [43].
Exosome-mediated immune evasion
Viral miRNA or proteins can also be packaged into exosomes [66]. Epstein-Barr virus miRNAs can be delivered to uninfected cells via exosomes [67]. The HIV accessory protein negative factor stimulates its own export by releasing exosomes [68]. Hepatitis virus infection is a major cause of HCC. Upon release from host cells, the hepatitis A virus cloaks itself in exosomes, thereby protecting the virion from antibody-mediated neutralization [69]. Similarly, the release of HCV from infected cells is linked to the exosomal pathway [70], and exosomes from HCV-infected cells are capable of transmitting infection to naive human hepatoma Huh7.5.1 cells [71]. Interestingly, levels of extracellular infectious HCV are significantly lower in exosomes released from autophagy knockdown cells [72]. The aforementioned sheds light on the potential role of the exosomal pathway as an immune evasion mechanism, enabling the hepatitis virus to replicate efficiently in the liver. Studies of HBV virion morphogenesis and secretion in HBV-infected hepatocytes demonstrated that multivesicular bodies and related cellular proteins were significantly involved in this process. Although HBV infection is known to alter the contents of exosomes [73], exosome-mediated immune mechanisms in HBV infection and HBV-related HCC remain unclear. Further research is needed on exosomal HBV miRNA and miRNA transport in HBV infection.
Applications of Exosomes in HCC
Exosomes for diagnosis
Exosome-associated miRNAs are more stable than nonexosome-associated miRNAs [74]. In addition, the expression of exosomal miRNAs is different to that of serum circulating miRNAs [75] and protein, as well as that of protein miRNAs [43]. Given the differences in the expression of these miRNAs, the exosomal content of cells can be used to aid the diagnosis of lung cancer [76], pancreatic cancer [12], and malignant melanoma [14]. As the miRNA and protein content of exosomes produced by HCC cells are completely different to those of normal cells [9, 36], with some miRNAs found only in HCC-derived exosomes [9], HCC-derived exosomes may serve as a novel diagnostic marker for HCC. Several exosomal miRNAs, such as miR-21, miR-18a, miR-221, miR-222, and miR-224, are candidate biomarkers [13, 15]. However, the diagnostic efficiency of exosomal miRNAs for HCC has not been determined. Furthermore, whether exosomal miRNA is a better biomarker than serum α-fetoprotein remains an intriguing topic for future investigations.
Exosomes for drug resistance
Drug resistance is a major obstacle in the treatment of HCC, and novel strategies need to be developed to combat drug toxicity and sensitivity in chemotherapy. In this regard, exosomes have attracted much interest. In one study, cisplatin enhanced cytotoxic T lymphocyte-specific cytotoxicity elicited by exosomes when compared that of controls, and significantly prolonged the survival time of mice with HCC [77]. Another study proposed that miR-122 could modulate the sensitivity of HCC cells to chemotherapeutic drugs by downregulating multidrug resistance-related genes, antiapoptotic genes, and the cell cycle-related gene cyclin B1 [24]. More recently, ADMSC-derived exosomes were shown to deliver miR-122 to HCC cells [35]. In vitro, the percentage of the G0/G1 population of HepG2 and Huh7 cells increased significantly when incubated with chemotherapeutic agents and/or ADMSC-derived exosomes (122-Exo) [35]. In the same study, Huh7 cells treated with 122-Exo showed similar G0/G1 arrest trends, and these effects were confirmed in vivo. These findings indicate that 122-Exo can enhance cell apoptosis and cell cycle arrest and increase the chemosensitivity of HCC [35]. Based on the literature, exosomes seems to show major promise in combating drug resistance and sensitivity in chemotherapy for HCC.
Exosomes as therapeutic tools
The function of exosomes as vehicles for the delivery of proteins and antigen chaperones illustrate their potential in immunotherapy of HCC [78]. For example, human macrophages can transfer miR-142 and miR-223 to HCC cells, decrease the expression of reporter proteins, and endogenously express stathmin-1 and the insulin-like growth factor-1 receptor, further inhibiting HCC cell proliferation or tumor growth [79]. Bone marrow stromal cells pulsed with homologous tumor-derived exosomes can increase the inhibition of homologous HCC H(22) cells, resulting in the arrest of H(22) cells in the G(0)/G(1) phase [80]. Exosomal proteins are actively released after stimulation. NY-ESO-1 is a tumor-specific antigen, with strong antigen immunogenicity [81]. The expression of the NY-ESO-1 protein in exosomes derived from HCC cell lines, as well as the expression of human leukocyte antigen, a tumor-specific immune-stimulating molecule, increases significantly after 5-Aza-2’-deoxycytidine treatment, indicating that exosomes secreted from hepatoma cells are able to stimulate an antitumor-specific immune response [82]. Notably, anticancer drugs also cause the release of exosomes from human HCC cells [83]. In vitro, after exposure to HCC cells resistant or sensitive to anticancer drugs, exosomes with superior immunogenicity were actively released by resistant anticancer drug-treated HepG2 cells, eliciting effective heat shock protein-specific natural killer cell antitumor responses [83]. In a rat model, exosomal ADMSCs facilitated HCC suppression by promoting NK T-cell anti-tumor responses [84]. These findings suggest that a vaccine that targeted exosomes could be efficient orin HCC immunotherapy.
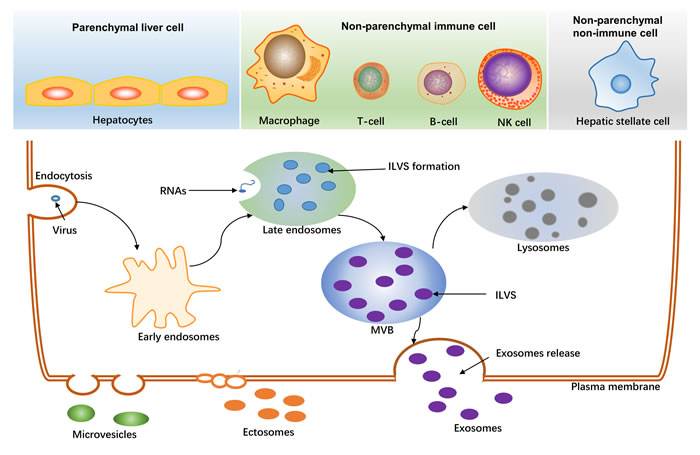
Figure 1: Generation of extracellular vesicles in liver. The main three cell-types released extracellular vesicles in liver are shown at the top. The biogenesis of exosomes is shown at the bottom. Virus, proteins, RNAs are sorted to late endosomes, and further formatted intraluminal vesicles(ILVs). ILVs in the multivesicular body (MVB) released to the extracellular space by an exocytic step is regarded as exosomes (30-120nm). MVB can also be degraded as lysosomes. Ectosomes (100-350nm) and other microvesicles (100-1000nm) are budded from plasma membrane.
Future Perspectives
Considerable advances have been made in understanding the functions of exosomes in HCC (Figure 2). These advances facilitate the development of a powerful new strategy to prevent and treat HCC. However, to exploit the potential of HCC-derived exosomes, more studies are needed of some specific areas, as described below:
First, there are no current technical standards established at present for the purification and isolation of exosomes. For example, ectosomes, which are very similar to exosomes, are oproduced by all cells (Figure 1). Thus, distinguishing purified and isolated exosomes and ectosomes is problematic [18]. In addition, the heterogeneity of exosomes (i.e., different density of exosomes and different expression of exosomal proteomic markers) needs to be considered as this can affect the purification of exosomespurified [85]. Therefore, more discriminant techniques are needed for the purification and isolation of exosomes to obtain more homogeneous exosomal preparations.
Second, cancer-associated fibroblasts (CAFs) promote tumorigenesis, progression, invasion, and chemoresistance in the tumor microenvironment of cancers [86]. In our previous work, we showed that CAFs, as well as energy metabolism reprogramming and oxygen stress, played important roles in abnormal proliferation [87]. The progress of HCC is always associated with CAFs [88, 89], as most cases of HCC are the result of liver cirrhosis [90]. As HCC-derived exosomes are known to play important roles in the tumor microenvironment, the next challenge is to better understand the role of HCC-derived exosomes in CAFs, energy metabolism reprogramming, and oxygen stress.
Third, the epithelial-mesenchymal transition (EMT) mediates HCC invasion and metastasis [91], and miRNA inhibits the expression of proteins involved in the EMT and promotes the EMT in cancer cells [92, 93]. The role of exosomes in the promotion of the EMT was delineated recently [94]. Hence, new studies could focus on the possible role of enriched and specific miRNAs in HCC-derived exosomes in the EMT.
Fourth, in addition to miRNAs, lncRNAs mediate the progress of HCC [95]. Recent work showed that circular RNAs functioned as miRNA sponges [96]. No studies have investigated the functions of lncRNAs and circular RNAs in HCC-derived exosomes. Considering the important functions of miRNAs in exosomes from HCC cells, further studies should investigate the potential roles of lncRNAs and circRNAs in exosomes in HCC.
Finally, treatment based on exosomes may be a double-edged sword, given the pro- and anticancer-inducing activity of these extracellular vehicles. Therefore, numerous large studies should evaluate the clinical efficacy and safety of exosomes for use in the development of a vaccine for HCC immunotherapy.
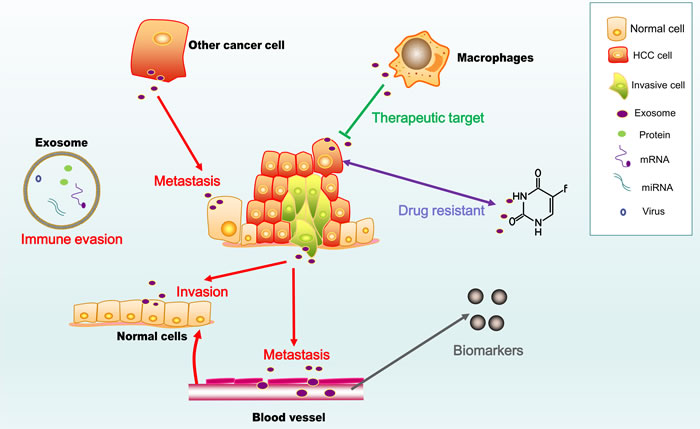
Figure 2: Two faces of exosomes in HCC. The figure shows the pro-cancer activity of exosomes (marked in red), including their roles in invasion, metastasis and immune evasion, as well as their potential use as diagnostic and prognostic biomarkers and immunotherapy for HCC.
Acknowledgments
This study was supported by Liuzhou scientific research and technological development programs (Grant No. 2015J030506, Sun Y); Natural Science Foundation of Guangxi University of Chinese Medicine (Grant No.YB14030, Sun Y) and The Self-funded research project of Guangxi Zhuang Autonomous Region health and Family Planning Commission(Grant No.Z2015206, Sun Y); Youth Science Foundation of Guangxi Medical University (GXMUYSF201545, Wu Z); National Natural Science Foundation of China (Grant No. 81301688, Cao K), Natural Science Foundation of Hunan Province (Grant No. 2016JJ6154, Cao K), 125 Talent Project/New xiangya project of the Third Xiangya Hospital of Central South University(Cao K).
Conflicts of interest
The authors declare that they have no conflict of interest.
References
1. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J and Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015; 65(2):87-108.
2. Ferlay J, Shin H-R, Bray F, Forman D, Mathers C and Parkin DM. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. International Journal of Cancer. 2010; 127(12):2893-2917.
3. Yang JD and Roberts LR. Hepatocellular carcinoma: a global view. Nature Reviews Gastroenterology & Hepatology. 2010; 7(8):448-458.
4. Santi V, Trevisani F, Gramenzi A, Grignaschi A, Mirici-Cappa F, Del Poggio P, Di Nolfo MA, Benvegnù L, Farinati F and Zoli M. Semiannual surveillance is superior to annual surveillance for the detection of early hepatocellular carcinoma and patient survival. J Hepatol. 2010; 53(2):291-297.
5. Sun WH, Liu Y, Shou DW, Sun QM, Shi JG, Chen L, Liang TB and Gong WH. AFP (alpha fetoprotein): Who are you in gastrology? Cancer Lett. 2015; 357(1):43-46.
6. Ertle JM, Heider D, Wichert M, Keller B, Kueper R, Hilgard P, Gerken G and Schlaak JF. A combination of alpha-fetoprotein and des-gamma-carboxy prothrombin is superior in detection of hepatocellular carcinoma. Digestion. 2013; 87(2):121-131.
7. Witjes CDM, van Aalten SM, Steyerberg EW, Borsboom G, de Man RA, Verhoef C and Ijzermans JNM. Recently introduced biomarkers for screening of hepatocellular carcinoma: a systematic review and meta-analysis. Hepatology International. 2013; 7(1):59-64.
8. Azmi AS, Bao B and Sarkar FH. Exosomes in cancer development, metastasis, and drug resistance: a comprehensive review. Cancer and Metastasis Reviews. 2013; 32(3-4):623-642.
9. Kogure T, Lin W-L, Yan IK, Braconi C and Patel T. Intercellular nanovesicle-mediated microRNA transfer: a mechanism of environmental modulation of hepatocellular cancer cell growth. Hepatology (Baltimore, Md). 2011; 54(4):1237-1248.
10. Ji H, Greening DW, Barnes TW, Lim JW, Tauro BJ, Rai A, Xu R, Adda C, Mathivanan S, Zhao W, Xue Y, Xu T, Zhu HJ and Simpson RJ. Proteome profiling of exosomes derived from human primary and metastatic colorectal cancer cells reveal differential expression of key metastatic factors and signal transduction components. Proteomics. 2013; 13(10-11):1672-1686.
11. Yuyama K, Sun H, Mitsutake S and Igarashi Y. Sphingolipid-modulated exosome secretion promotes clearance of amyloid-β by microglia. Journal of Biological Chemistry. 2012; 287(14):10977-10989.
12. Melo SA, Luecke LB, Kahlert C, Fernandez AF, Gammon ST, Kaye J, LeBleu VS, Mittendorf EA, Weitz J, Rahbari N, Reissfelder C, Pilarsky C, Fraga MF, Piwnica-Worms D and Kalluri R. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature. 2015; 523(7559):177-182.
13. Sohn W, Kim J, Kang SH, Yang SR, Cho J-Y, Cho HC, Shim SG and Paik Y-H. Serum exosomal microRNAs as novel biomarkers for hepatocellular carcinoma. Experimental & molecular medicine. 2015; 47:e184.
14. Alegre E, Zubiri L, Perez-Gracia JL, Gonzalez-Cao M, Soria L, Martin-Algarra S and Gonzalez A. Circulating melanoma exosomes as diagnostic and prognosis biomarkers. Clinica chimica acta. 2016; 454:28-32.
15. Wang H, Hou L, Li A, Duan Y, Gao H and Song X. Expression of serum exosomal microRNA-21 in human hepatocellular carcinoma. BioMed research international. 2014; 2014:864894.
16. Suchorska WM and Lach MS. The role of exosomes in tumor progression and metastasis (Review). Oncol Rep. 2016; 35(3):1237-1244.
17. Pan BT and Johnstone RM. Fate of the transferrin receptor during maturation of sheep reticulocytes in vitro: selective externalization of the receptor. Cell. 1983; 33(3):967-978.
18. Cocucci E and Meldolesi J. Ectosomes and exosomes: shedding the confusion between extracellular vesicles. Trends in cell biology. 2015; 25(6):364-372.
19. Oosthuyzen W, Sime NE, Ivy JR, Turtle EJ, Street JM, Pound J, Bath LE, Webb DJ, Gregory CD, Bailey MA and Dear JW. Quantification of human urinary exosomes by nanoparticle tracking analysis. The Journal of physiology. 2013; 591(23):5833-5842.
20. Street JM, Barran PE, Mackay CL, Weidt S, Balmforth C, Walsh TS, Chalmers RT, Webb DJ and Dear JW. Identification and proteomic profiling of exosomes in human cerebrospinal fluid. Journal of translational medicine. 2012; 10:5.
21. Lasser C, Alikhani VS, Ekstrom K, Eldh M, Paredes PT, Bossios A, Sjostrand M, Gabrielsson S, Lotvall J and Valadi H. Human saliva, plasma and breast milk exosomes contain RNA: uptake by macrophages. Journal of translational medicine. 2011; 9:9.
22. Wakim LM and Bevan MJ. Cross-dressed dendritic cells drive memory CD8+ T-cell activation after viral infection. Nature. 2011; 471(7340):629-632.
23. Greuter T and Shah VH. Hepatic sinusoids in liver injury, inflammation, and fibrosis: new pathophysiological insights. Journal of gastroenterology. 2016; 51(6):511-519.
24. Xu Y, Xia F, Ma L, Shan J, Shen J, Yang Z, Liu J, Cui Y, Bian X, Bie P and Qian C. MicroRNA-122 sensitizes HCC cancer cells to adriamycin and vincristine through modulating expression of MDR and inducing cell cycle arrest. Cancer letters. 2011; 310(2):160-169.
25. Wang B, Majumder S, Nuovo G, Kutay H, Volinia S, Patel T, Schmittgen TD, Croce C, Ghoshal K and Jacob ST. Role of microRNA-155 at early stages of hepatocarcinogenesis induced by choline-deficient and amino acid-defined diet in C57BL/6 mice. Hepatology (Baltimore, Md). 2009; 50(4):1152-1161.
26. Li S, Fu H, Wang Y, Tie Y, Xing R, Zhu J, Sun Z, Wei L and Zheng X. MicroRNA-101 regulates expression of the v-fos FBJ murine osteosarcoma viral oncogene homolog (FOS) oncogene in human hepatocellular carcinoma. Hepatology (Baltimore, Md). 2009; 49(4):1194-1202.
27. Wong QW, Lung RW, Law PT, Lai PB, Chan KY, To KF and Wong N. MicroRNA-223 is commonly repressed in hepatocellular carcinoma and potentiates expression of Stathmin1. Gastroenterology. 2008; 135(1):257-269.
28. Varnholt H, Drebber U, Schulze F, Wedemeyer I, Schirmacher P, Dienes HP and Odenthal M. MicroRNA gene expression profile of hepatitis C virus-associated hepatocellular carcinoma. Hepatology (Baltimore, Md). 2008; 47(4):1223-1232.
29. Meng F, Henson R, Wehbe-Janek H, Ghoshal K, Jacob ST and Patel T. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology. 2007; 133(2):647-658.
30. Gramantieri L, Ferracin M, Fornari F, Veronese A, Sabbioni S, Liu CG, Calin GA, Giovannini C, Ferrazzi E, Grazi GL, Croce CM, Bolondi L and Negrini M. Cyclin G1 is a target of miR-122a, a microRNA frequently down-regulated in human hepatocellular carcinoma. Cancer research. 2007; 67(13):6092-6099.
31. Huang XH, Wang Q, Chen JS, Fu XH, Chen XL, Chen LZ, Li W, Bi J, Zhang LJ, Fu Q, Zeng WT, Cao LQ, Tan HX and Su Q. Bead-based microarray analysis of microRNA expression in hepatocellular carcinoma: miR-338 is downregulated. Hepatology research. 2009; 39(8):786-794.
32. Murakami Y, Yasuda T, Saigo K, Urashima T, Toyoda H, Okanoue T and Shimotohno K. Comprehensive analysis of microRNA expression patterns in hepatocellular carcinoma and non-tumorous tissues. Oncogene. 2006; 25(17):2537-2545.
33. Chiba M, Kimura M and Asari S. Exosomes secreted from human colorectal cancer cell lines contain mRNAs, microRNAs and natural antisense RNAs, that can transfer into the human hepatoma HepG2 and lung cancer A549 cell lines. Oncol Rep. 2012; 28(5):1551-1558.
34. Kogure T and Patel T. Isolation of extracellular nanovesicle microRNA from liver cancer cells in culture. Methods in molecular biology (Clifton, NJ). 2013; 1024:11-18.
35. Lou G, Song X, Yang F, Wu S, Wang J, Chen Z and Liu Y. Exosomes derived from miR-122-modified adipose tissue-derived MSCs increase chemosensitivity of hepatocellular carcinoma. Journal of hematology & oncology. 2015; 8(1):122.
36. Wei J-x, Lv L-h, Wan Y-l, Cao Y, Li G-l, Lin H-m, Zhou R, Shang C-z, Cao J, He H, Han Q-f, Liu P-q, Zhou G and Min J. Vps4A functions as a tumor suppressor by regulating the secretion and uptake of exosomal microRNAs in human hepatoma cells. Hepatology (Baltimore, Md). 2015; 61(4):1284-1294.
37. Jopling C. Liver-specific microRNA-122: Biogenesis and function. RNA biology. 2012; 9(2):137-142.
38. Basu S and Bhattacharyya SN. Insulin-like growth factor-1 prevents miR-122 production in neighbouring cells to curtail its intercellular transfer to ensure proliferation of human hepatoma cells. Nucleic Acids Res. 2014; 42(11):7170-7185.
39. Pan Q, Ramakrishnaiah V, Henry S, Fouraschen S, de Ruiter PE, Kwekkeboom J, Tilanus HW, Janssen HLA and van der Laan LJW. Hepatic cell-to-cell transmission of small silencing RNA can extend the therapeutic reach of RNA interference (RNAi). Gut. 2012; 61(9):1330-1339.
40. Sugimachi K, Matsumura T, Hirata H, Uchi R, Ueda M, Ueo H, Shinden Y, Iguchi T, Eguchi H, Shirabe K, Ochiya T, Maehara Y and Mimori K. Identification of a bona fide microRNA biomarker in serum exosomes that predicts hepatocellular carcinoma recurrence after liver transplantation. British journal of cancer. 2015; 112(3):532-538.
41. Conde-Vancells J, Rodriguez-Suarez E, Embade N, Gil D, Matthiesen R, Valle M, Elortza F, Lu SC, Mato JM and Falcon-Perez JM. Characterization and comprehensive proteome profiling of exosomes secreted by hepatocytes. J Proteome Res. 2008; 7(12):5157-5166.
42. Wang K, Ye S, Song L, Cui J, Weng Y, Liang C, Sun R and Tang Z. [A preliminary study of the killing function in vitro by T lymphocytes activated by dendritic cells loaded with exosomes secreted by hepatic cancer cell lines with high or low metastatic potentials [Article in Chinese]. Zhonghua gan zang bing za zhi. 2007; 15(9):658-662.
43. He M, Qin H, Poon TCW, Sze S-C, Ding X, Co NN, Ngai S-M, Chan T-F and Wong N. Hepatocellular carcinoma-derived exosomes promote motility of immortalized hepatocyte through transfer of oncogenic proteins and RNAs. Carcinogenesis. 2015; 36(9):1008-1018.
44. Zhao X, Wu Y, Duan J, Ma Y, Shen Z, Wei L, Cui X, Zhang J, Xie Y and Liu J. Quantitative proteomic analysis of exosome protein content changes induced by hepatitis B virus in Huh-7 cells using SILAC labeling and LC-MS/MS. J Proteome Res. 2014; 13(12):5391-5402.
45. Fleming JM, Ginsburg E, Oliver SD, Goldsmith P and Vonderhaar BK. Hornerin, an S100 family protein, is functional in breast cells and aberrantly expressed in breast cancer. BMC cancer. 2012; 12(1):1.
46. Kucharzewska P, Christianson HC, Welch JE, Svensson KJ, Fredlund E, Ringner M, Morgelin M, Bourseau-Guilmain E, Bengzon J and Belting M. Exosomes reflect the hypoxic status of glioma cells and mediate hypoxia-dependent activation of vascular cells during tumor development. Proceedings of the National Academy of Sciences of the United States of America. 2013; 110(18):7312-7317.
47. Birchmeier C, Birchmeier W, Gherardi E and Vande Woude GF. Met, metastasis, motility and more. Nature reviews Molecular cell biology. 2003; 4(12):915-925.
48. Tse EY, Ko FC, Tung EK, Chan LK, Lee TK, Ngan ES, Man K, Wong AS, Ng IO and Yam JW. Caveolin-1 overexpression is associated with hepatocellular carcinoma tumourigenesis and metastasis. The Journal of pathology. 2012; 226(4):645-653.
49. Mishra SK, Siddique HR and Saleem M. S100A4 calcium-binding protein is key player in tumor progression and metastasis: preclinical and clinical evidence. Cancer metastasis reviews. 2012; 31(1-2):163-172.
50. Xu L, Yang BF and Ai J. MicroRNA transport: a new way in cell communication. Journal of cellular physiology. 2013; 228(8):1713-1719.
51. Bukong TN, Momen-Heravi F, Kodys K, Bala S and Szabo G. Exosomes from hepatitis C infected patients transmit HCV infection and contain replication competent viral RNA in complex with Ago2-miR122-HSP90. PLoS pathogens. 2014; 10(10):e1004424.
52. Angeloni NL, McMahon KM, Swaminathan S, Plebanek MP, Osman I, Volpert OV and Thaxton CS. Pathways for Modulating Exosome Lipids Identified By High-Density Lipoprotein-Like Nanoparticle Binding to Scavenger Receptor Type B-1. Scientific reports. 2016; 6:22915.
53. Janas T, Janas MM, Sapon K and Janas T. Mechanisms of RNA loading into exosomes. FEBS letters. 2015; 589(13):1391-1398.
54. Cao C, Sun J, Zhang D, Guo X, Xie L, Li X, Wu D and Liu L. The long intergenic noncoding RNA UFC1, a target of MicroRNA 34a, interacts with the mRNA stabilizing protein HuR to increase levels of beta-catenin in HCC cells. Gastroenterology. 2015; 148(2):415-426.e418.
55. Ahadi A, Brennan S, Kennedy PJ, Hutvagner G and Tran N. Long non-coding RNAs harboring miRNA seed regions are enriched in prostate cancer exosomes. Scientific reports. 2016; 6:24922.
56. Schilders G, Raijmakers R, Raats JM and Pruijn GJ. MPP6 is an exosome-associated RNA-binding protein involved in 5.8S rRNA maturation. Nucleic acids research. 2005; 33(21):6795-6804.
57. van Zijl F, Krupitza G and Mikulits W. Initial steps of metastasis: Cell invasion and endothelial transmigration. Mutation Research-Reviews in Mutation Research. 2011; 728(1-2):23-34.
58. Hui L and Chen Y. Tumor microenvironment: Sanctuary of the devil. Cancer letters. 2015; 368(1):7-13.
59. Kahlert C and Kalluri R. Exosomes in tumor microenvironment influence cancer progression and metastasis. Journal of molecular medicine (Berlin, Germany). 2013; 91(4):431-437.
60. Malato Y and Willenbring H. The MAP3K TAK1: a chock block to liver cancer formation. Hepatology (Baltimore, Md). 2010; 52(4):1506-1509.
61. Yamashita T, Honda M, Nakamoto Y, Baba M, Nio K, Hara Y, Zeng SS, Hayashi T, Kondo M, Takatori H, Yamashita T, Mizukoshi E, Ikeda H, Zen Y, Takamura H, Wang XW, et al. Discrete nature of EpCAM+ and CD90+ cancer stem cells in human hepatocellular carcinoma. Hepatology (Baltimore, Md). 2013; 57(4):1484-1497.
62. Guo Z, Li LQ, Jiang JH, Ou C, Zeng LX and Xiang BD. Cancer stem cell markers correlate with early recurrence and survival in hepatocellular carcinoma. World journal of gastroenterology. 2014; 20(8):2098-2106.
63. Conigliaro A, Costa V, Lo Dico A, Saieva L, Buccheri S, Dieli F, Manno M, Raccosta S, Mancone C, Tripodi M, De Leo G and Alessandro R. CD90+ liver cancer cells modulate endothelial cell phenotype through the release of exosomes containing H19 lncRNA. Molecular cancer. 2015; 14:155.
64. Li S, Li H, Yang X, Wang W, Huang A, Li J, Qin X, Li F, Lu G, Ding H, Su X, Hou L, Xia W, Shi M, Zhang H, Zhao Q, et al. Vasorin is a potential serum biomarker and drug target of hepatocarcinoma screened by subtractive-EMSA-SELEX to clinic patient serum. Oncotarget. 2015; 6(12):10045-10059. doi: 10.18632/oncotarget.3541.
65. Huang A, Dong J, Li S, Wang C, Ding H, Li H, Su X, Ge X, Sun L, Bai C, Shen X, Fang T, Li J and Shao N. Exosomal transfer of vasorin expressed in hepatocellular carcinoma cells promotes migration of human umbilical vein endothelial cells. Int J Biol Sci. 2015; 11(8):961-969.
66. Alenquer M and Amorim MJ. Exosome Biogenesis, Regulation, and Function in Viral Infection. Viruses. 2015; 7(9):5066-5083.
67. Pegtel DM, Cosmopoulos K, Thorley-Lawson DA, van Eijndhoven MA, Hopmans ES, Lindenberg JL, de Gruijl TD, Wurdinger T and Middeldorp JM. Functional delivery of viral miRNAs via exosomes. Proceedings of the National Academy of Sciences of the United States of America. 2010; 107(14):6328-6333.
68. Lenassi M, Cagney G, Liao M, Vaupotic T, Bartholomeeusen K, Cheng Y, Krogan NJ, Plemenitas A and Peterlin BM. HIV Nef is secreted in exosomes and triggers apoptosis in bystander CD4+ T cells. Traffic (Copenhagen, Denmark). 2010; 11(1):110-122.
69. Feng Z, Hensley L, McKnight KL, Hu F, Madden V, Ping L, Jeong SH, Walker C, Lanford RE and Lemon SM. A pathogenic picornavirus acquires an envelope by hijacking cellular membranes. Nature. 2013; 496(7445):367-371.
70. Tamai K, Shiina M, Tanaka N, Nakano T, Yamamoto A, Kondo Y, Kakazu E, Inoue J, Fukushima K, Sano K, Ueno Y, Shimosegawa T and Sugamura K. Regulation of hepatitis C virus secretion by the Hrs-dependent exosomal pathway. Virology. 2012; 422(2):377-385.
71. Ramakrishnaiah V, Thumann C, Fofana I, Habersetzer F, Pan Q, de Ruiter PE, Willemsen R, Demmers JAA, Stalin Raj V, Jenster G, Kwekkeboom J, Tilanus HW, Haagmans BL, Baumert TF and van der Laan LJW. Exosome-mediated transmission of hepatitis C virus between human hepatoma Huh7.5 cells. Proc Natl Acad Sci U S A. 2013; 110(32):13109-13113.
72. Shrivastava S, Devhare P, Sujijantarat N, Steele R, Kwon Y-C, Ray R and Ray RB. Knockdown of Autophagy Inhibits Infectious Hepatitis C Virus Release by the Exosomal Pathway. J Virol. 2015; 90(3):1387-1396.
73. Zhao X, Wu Y, Duan J, Ma Y, Shen Z, Wei L, Cui X, Zhang J, Xie Y and Liu J. Quantitative proteomic analysis of exosome protein content changes induced by hepatitis B virus in Huh-7 cells using SILAC labeling and LC-MS/MS. Journal of proteome research. 2014; 13(12):5391-5402.
74. Cheng L, Sharples RA, Scicluna BJ and Hill AF. Exosomes provide a protective and enriched source of miRNA for biomarker profiling compared to intracellular and cell-free blood. Journal of extracellular vesicles. 2014; 3.
75. Fornari F, Ferracin M, Trere D, Milazzo M, Marinelli S, Galassi M, Venerandi L, Pollutri D, Patrizi C, Borghi A, Foschi FG, Stefanini GF, Negrini M, Bolondi L and Gramantieri L. Circulating microRNAs, miR-939, miR-595, miR-519d and miR-494, Identify Cirrhotic Patients with HCC. PloS one. 2015; 10(10):e0141448.
76. Cazzoli R, Buttitta F, Di Nicola M, Malatesta S, Marchetti A, Rom WN and Pass HI. microRNAs derived from circulating exosomes as noninvasive biomarkers for screening and diagnosing lung cancer. Journal of thoracic oncology. 2013; 8(9):1156-1162.
77. Wang S-H, Shen Y, Li J, Xiang Z-W, Fan W-K and Chen L. [Experimental studies on anti-mouse hepatocellular carcinoma effects of cisplatin combined with exosomes]. Xi bao yu fen zi mian yi xue za zhi = Chinese journal of cellular and molecular immunology. 2009; 25(1):49-52.
78. Mignot G, Roux S, Thery C, Segura E and Zitvogel L. Prospects for exosomes in immunotherapy of cancer. Journal of cellular and molecular medicine. 2006; 10(2):376-388.
79. Aucher A, Rudnicka D and Davis DM. MicroRNAs transfer from human macrophages to hepato-carcinoma cells and inhibit proliferation. Journal of immunology (Baltimore, Md : 1950). 2013; 191(12):6250-6260.
80. Ma B, Jiang H, Jia J, Di L, Song G, Yu J, Zhu Y, Lu Z, Wang X, Zhou X and Ren J. Murine bone marrow stromal cells pulsed with homologous tumor-derived exosomes inhibit proliferation of liver cancer cells. Clinical & translational oncology. 2012; 14(10):764-773.
81. Nakamura S, Nouso K, Noguchi Y, Higashi T, Ono T, Jungbluth A, Chen YT, Old LJ, Nakayama E and Shiratori Y. Expression and immunogenicity of NY-ESO-1 in hepatocellular carcinoma. Journal of gastroenterology and hepatology. 2006; 21(8):1281-1285.
82. Xiao W-H, Sanren G-W, Zhu J-H, Li Q-W, Kang H-R, Wang R-L, Song L-P and Ye M. Effect of 5-aza-2’-deoxycytidine on immune-associated proteins in exosomes from hepatoma. World journal of gastroenterology. 2010; 16(19):2371-2377.
83. Lv L-H, Wan Y-L, Lin Y, Zhang W, Yang M, Li G-L, Lin H-M, Shang C-Z, Chen Y-J and Min J. Anticancer drugs cause release of exosomes with heat shock proteins from human hepatocellular carcinoma cells that elicit effective natural killer cell antitumor responses in vitro. The Journal of biological chemistry. 2012; 287(19):15874-15885.
84. Ko S-F, Yip H-K, Zhen Y-Y, Lee C-C, Lee C-C, Huang C-C, Ng S-H and Lin J-W. Adipose-Derived Mesenchymal Stem Cell Exosomes Suppress Hepatocellular Carcinoma Growth in a Rat Model: Apparent Diffusion Coefficient, Natural Killer T-Cell Responses, and Histopathological Features. Stem cells international. 2015; 2015:853506.
85. Bobrie A and Thery C. Exosomes and communication between tumours and the immune system: are all exosomes equal? Biochem Soc Trans. 2013; 41:263-267.
86. Franco OE, Shaw AK, Strand DW and Hayward SW. Cancer associated fibroblasts in cancer pathogenesis. Seminars in Cell & Developmental Biology. 2010; 21(1):33-39.
87. Tang S, Hou Y, Zhang H, Tu G, Yang L, Sun Y, Lang L, Tang X, Du YE, Zhou M, Yu T, Xu L, Wen S, Liu C and Liu M. Oxidized ATM promotes abnormal proliferation of breast CAFs through maintaining intracellular redox homeostasis and activating the PI3K-AKT, MEK-ERK, and Wnt-beta-catenin signaling pathways. Cell cycle (Georgetown, Tex). 2015; 14(12):1908-1924.
88. Li T, Yang Y, Hua X, Wang G, Liu W, Jia C, Tai Y, Zhang Q and Chen G. Hepatocellular carcinoma-associated fibroblasts trigger NK cell dysfunction via PGE2 and IDO. Cancer Lett. 2012; 318(2):154-161.
89. Sukowati CHC, Anfuso B, Croce LS and Tiribelli C. The role of multipotent cancer associated fibroblasts in hepatocarcinogenesis. BMC cancer. 2015; 15.
90. Yuen MF, Tanaka Y, Fong DY, Fung J, Wong DK, Yuen JC, But DY, Chan AO, Wong BC, Mizokami M and Lai CL. Independent risk factors and predictive score for the development of hepatocellular carcinoma in chronic hepatitis B. Journal of hepatology. 2009; 50(1):80-88.
91. Zhao XL, Sun T, Che N, Sun D, Zhao N, Dong XY, Gu QA, Yao Z and Sun BC. Promotion of hepatocellular carcinoma metastasis through matrix metalloproteinase activation by epithelial-mesenchymal transition regulator Twist1. J Cell Mol Med. 2011; 15(3):691-700.
92. Burk U, Schubert J, Wellner U, Schmalhofer O, Vincan E, Spaderna S and Brabletz T. A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. Embo Reports. 2008; 9(6):582-589.
93. Renjie W and Haiqian L. MiR-132, miR-15a and miR-16 synergistically inhibit pituitary tumor cell proliferation, invasion and migration by targeting Sox5. Cancer letters. 2015; 356(2 Pt B):568-578.
94. Garnier D, Magnus N, Lee TH, Bentley V, Meehan B, Milsom C, Montermini L, Kislinger T and Rak J. Cancer cells induced to express mesenchymal phenotype release exosome-like extracellular vesicles carrying tissue factor. The Journal of biological chemistry. 2012; 287(52):43565-43572.
95. Chang L, Wang G, Jia T, Zhang L, Li Y, Han Y, Zhang K, Lin G, Zhang R, Li J and Wang L. Armored long non-coding RNA MEG3 targeting EGFR based on recombinant MS2 bacteriophage virus-like particles against hepatocellular carcinoma. Oncotarget. 2016. 7:23988-4004. doi: 10.18632/oncotarget.8115.
96. Chen LL. The biogenesis and emerging roles of circular RNAs. Nature reviews Molecular cell biology. 2016; 17(4):205-211.

