INTRODUCTION
Prostate cancer (PCa) is a highly prevalent disease that remains clinically silent in the majority of cases and is mostly curable when localized. Advanced stages of PCa initially respond well to anti-androgen therapy but will usually progress to castration resistance with poor prognosis. Obviously, new prognostic tools are needed to allow for an early recognition of aggressive versus indolent forms of PCa. Furthermore, the development of novel targeted therapies for advanced PCa, e.g. immunotherapies, necessitates the development of future predictive biomarkers as companion diagnostics.
The α-chemokine receptor C-X-C chemokine receptor type 4 (CXCR4) and its endogenous ligand CXCL12, also called stromal-derived-factor 1 (SDF1), are attractive therapeutic targets as they are widely expressed in numerous epithelial, mesenchymal, and hematopoietic tumors [1–8]. Recent data suggest a chief position of the CXCR4/CXCL12 axis initializing androgen dependent proliferation, tumor cell motility, and metastatic growth in PCa [9].
Originally, CXCL12 was found to control hematopoietic cell trafficking and guide the homing of stem cells within the bone marrow [10, 11]. Thereby, cells with CXCR4 expression migrate towards compartments with high CXCL12 production. In the bone marrow, CXCL12 signals lead to cell chemotaxis until they are terminated through CXCR4 receptor internalization [12]. In the long-run, steady CXCL12 binding causes CXCR4 desensitization resulting in a resting, non-deploying state of hematopoietic stem cells and tumor cells, likewise [13]. This implies that a microenvironment with a constant CXCL12 production induces a down-regulation of CXCR4 and thereby antagonizes the process of metastasis [10]. Gene methylation adds to the major regulatory alterations interfering with CXCL12 homeostasis in tumor epithelium, leading to a silenced CXCL12 signal in various human malignant tumors [14–17].
Moreover, the CXCR4/CXCL12 axis seems to be tightly linked to the immune checkpoint regulator programmed death 1 (PD-1) and programmed death-ligand 1 (PD-L1) that co-operate to suppress anti-cancer immunity [18]. In the clinical setting, anti-PD-1 and anti-PD-L1 antibodies have shown promising outcomes in cancer patients. However, a subset of patients fails to respond to checkpoint inhibition [19]. Feig et al. recently managed to overcome anti-PD-L1 drug resistance in a murine pancreatic adenocarcinoma model by blocking CXCR4. These findings motivated us to investigate the CXCR4/CXCL12 axis within the cross-relation to PD-L1 expression in PCa
RESULTS
Analytical assay performance
Methylation levels of CXCL12 were determined by quantitative methylation-specific real-time PCR detecting methylated and unmethylated CXCL12 promoter sequences. The assay was designed within a CpG island upstream of the CXCL12 gene (Figure 1A). The analytical performance of the assay was tested using mixtures of bisulfite converted artificially methylated and unmethylated DNA. Figure 1B shows that the assay allowed for the accurate quantification of CXCL12 methylation within the whole spectrum of 0 – 100% methylation (r2 = 0.98).
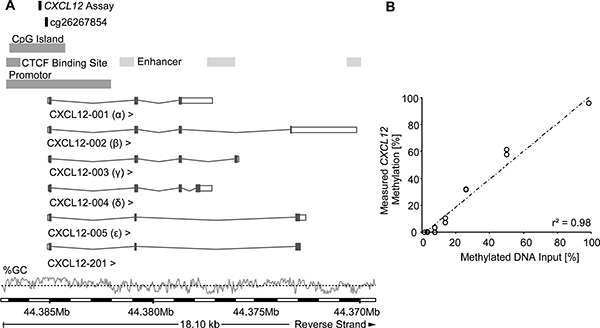
Figure 1: Genomic location and analytical performance verification of the quantitative CXCL12 methylation-specific real-time PCR and the bead cg26267854 from the Illumina Infinium HumanMethylation450 BeadChip. Location of the gene is on the reverse strand of chromosome 10, the information for this figure was taken from Ensembl Homo sapiens version 82.38 (GRCh38.p3). Six splice variants of CXCL12 have been identified in humans, with CXCL12-α and CXCL12-β as the main isoforms. The GC content [%] is shown with the black dotted line indicating 50% GC content (A). For analytical performance verification of the qPCR assay a dilution series of bisulfite-converted artificially methylated and unmethylated sperm DNA was analysed (mixtures: 0, 0.78, 1.56, 3.125, 6.25, 12.5, 25, 50, 100 % methylated DNA). Each DNA mixture was analysed in duplicate (B).
Frequency of CXCL12 gene methylation in human prostate tissue samples
In a test case control study, we evaluated a total of 66 human prostate tissue samples from 25 radical prostatectomy specimens regarding their CXCL12 methylation (mCXCL12) status in normal prostate parenchyma, hyperplastic adenomyomatous tissue, and carcinomatous tissue. These 25 PCa specimens were representative of a wide spectrum of the disease and comprised the following Gleason grading groups (GG): GG I 28%, GG II-III 28%, GG IV 24, and GG V 20%. Among non-carcinomatous samples, we analyzed 22 normal tissue samples remote from the tumor front and 19 benign hyperplastic cases. Compared to PCa samples, CXCL12 methylation was altogether significantly lower in benign and normal specimens compared to PCa samples (p < 0.05 and p < 0.01, Figure 2). Mean mCXCL12 values of normal and PCa tissue were narrowly confined to 39.5% [31.1%-48.1%] and 56% [45.4%-68.5%], respectively. Dispersion around the mean of mCXCL12 in PCa samples, however, was significantly more prominent (Χ2 = 14.1, p < 0.001, Bartlett’s test for equal variances). In detail, one group of patients showed values offsetting over approximately 70% methylation and another group was observed at a distance below approximately 30% methylation (Figure 2).
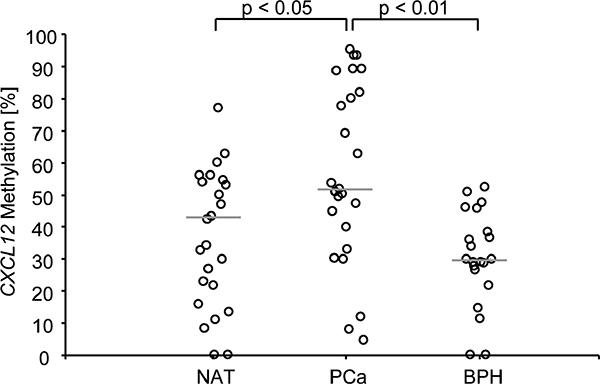
Figure 2: CXCL12 promoter methylation in tissue samples from PCa patients. Normal adjacent tissue samples and benign hyperplastic samples displayed a lower CXCL12 promoter methylation than prostate cancer samples. The median methylation is indicated by the grey line. The two cut-offs used for the survival analysis are indicted by the dotted lines. In Bonferroni’s multiple comparison test, PCa showed significantly higher CXCL12 methylation percentages than normal adjacent and benign hyperplastic tissue.
Differences in the spectrum of mCXCL12 prompted us to investigate the clinical performance of mCXCL12 in two independent cohorts of patients that had undergone radical prostatectomy.
Methylation and expression of CXCL12 in the training cohort
Full clinicopathological characteristics of the training cohort are given in Table 1. Among prognostic clinicopathological variables, a significant association was found between high mCXCL12 and high Gleason grading groups (p < 0.001) as well as nodal positive (pN1) patients (p = 0.038; Table 1).
Table 1: Clinico-pathological data and their association with CXCL12 methylation in the training (n = 247) and the testing (n = 498) cohort comprised of prostate cancer patients who underwent radical prostatectomy
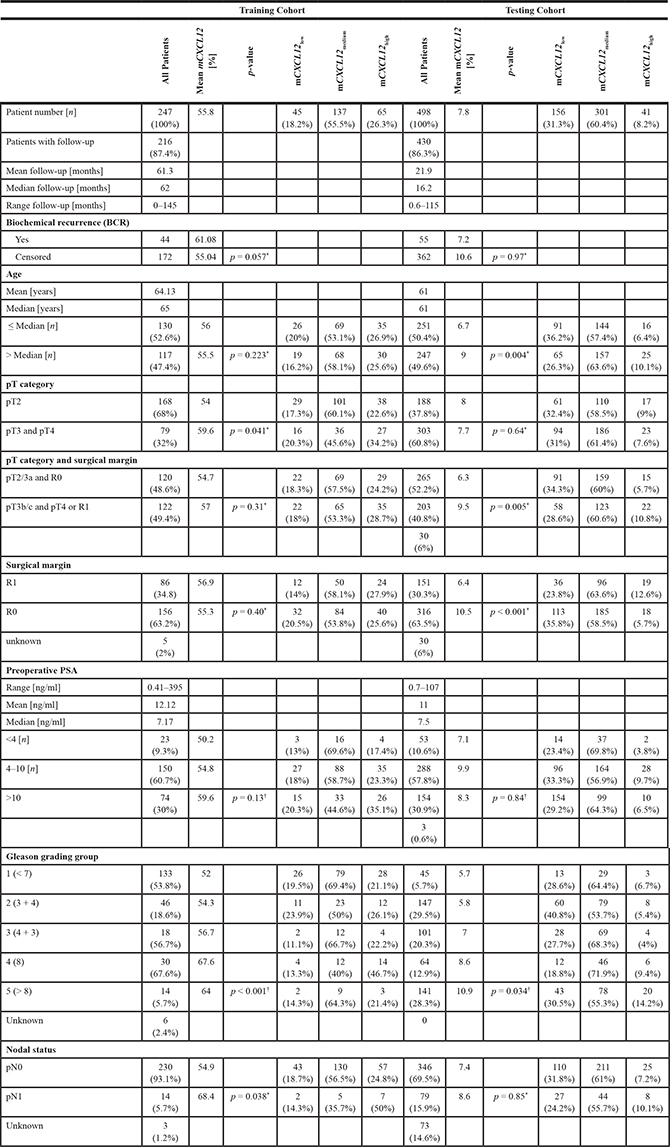
*Wilcoxon-Mann-Whitney test, †Kruskal-Wallis test.
In the training cohort, mCXCL12 revealed a symmetric distribution covering a broad spectrum of values (0.04% – 98.40%) with a major peak seen at 55% CXCL12 methylation, comparable with the results shown for the test case control series. Aiming at reproducing the characteristics of pathological mCXCL12 distribution in PCa, methylation values were trichotomized to obtain qualitative results. Two optimized cut-offs were introduced at 33.06% and 70.68%; mCXCL12low, mCXCL12medium, and mCXCL12high referring to the respective groups.
Immunohistochemical staining revealed a strong nuclear and cytoplasmic positivity of basal cells in normal prostate parenchyma. In carcinomatous glands, in contrast, CXCL12 expression was altogether fainter (Supplementary Figure 1). A significant association of CXCL12 staining intensity with categorical mCXCL12 was found (r = −0.21; p = 0.047; Χ2 = 3.97; p = 0.046).
CXCL12 methylation and biochemical recurrence-free survival analyses: Training cohort
Subsequently, we analyzed whether mCXCL12 allows for the stratification of patients at risk for biochemical recurrence (BCR, Table 2). Trichotomized mCXCL12 stratified patients according to BCR in the Kaplan-Meier analysis (χ2 = 6.13, p = 0.047, Figure 3A).
Table 2: Univariate and multivariate Cox proportional hazards analyses on BCR-free survival of the training and the testing cohort comprised of prostate cancer patients treated by radical prostatectomy
Training Cohort |
Testing Cohort |
|||||||
|---|---|---|---|---|---|---|---|---|
Univariate Cox |
Multivariate Cox |
Univariate Cox |
Multivariate Cox |
|||||
Clinico-pathologic parameter / biomarker |
Hazard ratio [95% CI] |
p-value |
Hazard ratio [95% CI] |
p-value |
Hazard ratio [95% CI] |
p-value |
Hazard ratio [95% CI] |
p-value |
Tumor stage (pT3 and pT4 vs. pT2) |
2.84 [1.57–5.14] |
0.001 |
5.37 [2.14–13.5] |
0.001 |
||||
Tumor stage (pT3b and pT4/R1 vs. pT3a and pT2/R0 ) |
3.47 [1.74–6.91] |
< 0.001 |
2.21 [1.04–4.72] |
0,041 |
1.99 [1.13–3.49] |
0.016 |
1.10 [0.57–2.12] |
0.77 |
Surgical margin (R1 vs. R0) |
2.46 [1.35–4.51] |
0.003 |
1.49 [0.87–2.56] |
0.15 |
||||
Gleason grading group |
1.90 [1.54–2.34] |
< 0.001 |
1.73 [1.38–2.18] |
< 0.001 |
1.69 [1.34–2.13] |
< 0.001 |
1.62 [1.22–2.14] |
0.001 |
Nodal status (pN1 vs. pN0) |
2.18 [0.86–5.55] |
0.102 |
1.84 [1.00–3.35] |
0.049 |
1.24 [0.65–2.36] |
0.54 |
||
Preoperative PSA level < 4 ng/ml vs. 4–10 ng/ml, vs. > 10 ng/ml |
1.70 [0.99–2.92] |
0.053 |
1.54 [0.98–2.43] |
0.060 |
||||
CXCL12 methylation (mCXCL12low/highvs. mCXCL12medium) |
2.11 [1.15–3.87] |
0.016 |
1.48 [0.79–2.77] |
0,22 |
1.92 [1.12–3.27] |
0.017 |
1.76 [1.00–3.09] |
0.049 |
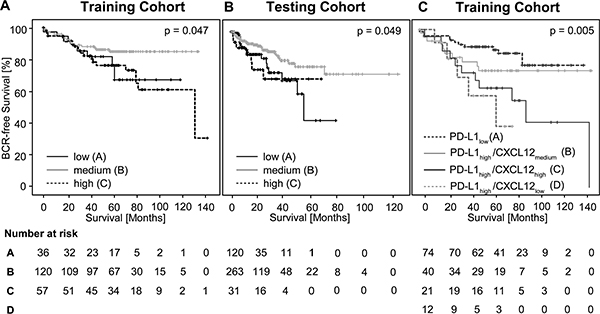
Figure 3: Kaplan-Meier survival of BCR-free survival in prostate cancer patients treated with radical prostatectomy and stratified by trichotomized CXCL12 DNA methylation levels (mCXCL12low, mCXCL12medium, and mCXCL12high) in the training cohort (A) and in the testing cohort (B). Kaplan-Meier survival analysis of BCR-free survival in prostate cancer patients stratified according to the CXCL12 methylation and PD-L1 expression levels in the training cohort (C), respectively.
For univariate Cox proportional hazard analysis, mCXCL12low and mCXCL12high were combined in one group. In the training cohort, 44.5% of patients fell within the mCXCL12low/high group and 55.5% fell within the mCXCL12medium group. Abnormal mCXCL12low/high was associated with significantly shorter BCR-free survival (hazard ratio (HR) = 2.11 [95%CI: 1.15–3.87],p = 0.016, Table 2).
Clinical performance of mCXCL12 in the testing cohort
Table 1 displays the full clinicopathological characteristics of the testing cohort. The results from the testing cohort are entirely based upon data generated by the TCGA Research Network: http://cancergenome.nih.gov/. mCXCL12 significantly correlated with age (p = 0.004), Gleason grading group (p = 0.034), and surgical margins/pT3b-pT4 categories (p = 0.005).
According to the approach chosen for the training cohort, mCXCL12 was trichotomized using optimized cut-offs (Table 1). Trichotomized mCXCL12 stratified patients according to BCR in the Kaplan-Meier analysis (χ2 = 6.01, p = 0.049, Figure 3B).
Analogous to the approach in the training cohort, mCXCL12low and mCXCL12 high groups were combined to mCXCL12low/high for univariate Cox proportional hazard analysis. In the testing cohort, 39.6% of patients fell within the mCXCL12low/high group and 60.4% fell within the mCXCL12medium group.
In univariate Cox proportional hazard analysis, mCXCL12low/high was associated with significantly shorter BCR-free survival (HR = 1.92 [95%CI: 1.12–3.27], p = 0.017, Table 2, Figure 4A). In multivariate analysis including pT/R, pN, and Gleason grading group, mCXCL12low/high qualified as an independent prognostic parameter (HR = 1.76 [95%CI: 1.00–3.09], p = 0.049, Table 2, Figure 4B).
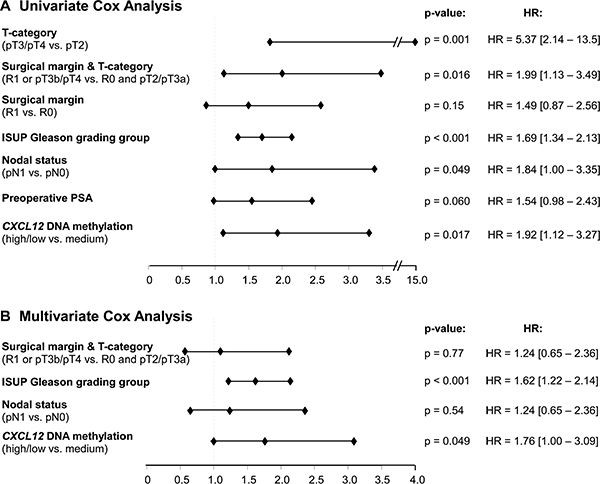
Figure 4: Forest Plot of the univariate (A) and multivariate (B) Cox proportional hazards analysis of BCR-free survival in the testing cohort.
mCXCL12 in relation to preoperative serum PSA
It has been shown that CXCL12 can help to distinguish between benign lesions and PCa among patients presenting with low serum PSA [20], which prompted us to evaluate mCXCL12 with respect to the initial serum PSA in the full cohort of patients. Patients were stratified according to initial PSA levels. Mean mCXCL12 did not differ between low-level PSA secreting (PSA < 4 ng/ml), medium-level PSA secreting (PSA ≥ 4 ng/ml, ≤ 10 ng/ml), and high-level PSA secreting (PSA > 10 ng/ml) tumors in both cohorts under investigation (Table 1). mCXCL12low/high was significantly associated with BCR in medium-level PSA secreting PCa in the univariate Cox proportional hazard model (Table 3).
Table 3: Univariate Cox proportional hazards analyses on BCR-free survival of the training and the testing cohort comprised of prostate cancer patients treated by radical prostatectomy
Training Cohort |
Testing Cohort |
||||||
|---|---|---|---|---|---|---|---|
Univariate Cox |
Univariate Cox |
||||||
Stratification according to initial PSA |
Hazard ratio [95% CI] |
n |
p-value |
Clinico-pathologic parameter / biomarker |
Hazard ratio [95% CI] |
n |
p-value |
PSA < 4 ng/ml |
ND |
23 |
PSA < 4 ng/ml |
3.46 [0.22–55.78] |
53 |
0.38 |
|
PSA 4–10 ng/ml |
2.02 [0.91 – 4.44] |
150 |
0.082 |
PSA 4–10 ng/ml |
2.09 [1.00–4.36] |
288 |
0.049 |
PSA > 10 ng/ml |
ND |
74 |
PSA > 10 ng/ml |
1.55 [0.67–3.59] |
154 |
0.3 |
|
Patients were stratified according to CXCL12 DNA methylation and BCR-free survival was analyzed in three subgroups of patients: PSA < 4 ng/ml, PSA 4–10 ng/ml, and PSA > 10 ng/ml.
ND: not determined due to low number of events.
Association with PD-L1 expression in the training cohort
Very recently, Gevensleben et al. showed that PD-L1 expression served as an independent prognostic marker in PCa in the same cohort of patients [21]. We correlated mCXCL12 with the previously published PD-L1 expression data. Mean mCXCL12 did not differ between PD-L1high and PD-L1low PCa. However, Spearman’s rank correlation revealed a trend towards an association of PD-L1with mCXCL12 (ρ = 0.132, p = 0.084). Therefore, subgroups were defined according to PD-L1 expression in PCa: PD-L1high PCa were flagged and allocated to mCXCL12low, mCXCL12medium, and mCXCL12high groups, respectively. A total of 85 patients presented with PD-L1low, while 87 PD-L1high patients were split into 15 mCXCL12low, 45 mCXCL12medium, and 27 mCXCL12high. In Kaplan-Meier analysis, PD-L1low PCa showed the longest BCR-free survival (mean estimated 112 months), while mCXCL12medium showed best BRC-free survival rates among PD-L1high PCa (mean estimated 107 months), and PD-L1high/mCXCL12low, and PD-L1high/mCXCL12high showed short BCR-free survival times (mean estimated 52 months and 83 months, respectively; n = 151, χ2 = 12.99; p = 0.005; Figure 3C).
Concordance, accuracy, and robustness of mCXCL12 detection in biopsies and ectomies
It has previously been shown that methylation assays precisely report methylation status even in formalin-fixed paraffin-embedded (FFPE) tissues [22, 23]. In order to test the utility of our assay for the analysis of biopsy specimens, we evaluated the robustness, accuracy, and concordance of mCXCL12 testing in small biopsy specimens and the matched ectomy in a biospy test case series including 10 patients. Therefore, we correlated mCXCL12 obtained from pooled biopsy cores of an individual patient with mCXCL12 detected in his total tumor volume after radical prostatectomy (Figure 5A). mCXCL12 levels detected in biopsies significantly correlated with those obtained from the total tumor volume (r = 0.76; p = 0.019) indicating that mCXCL12 in biopsies represents the methylation level found in the tumor. In a second step, we tested whether mCXCL12 can robustly and accurately be tested in small samples and over a wide range of DNA yield. Patients of the biopsy test case series were divided into two groups: five patients with mCXCL12 ranging from 15.5% – 41.2% were assigned to group A and 5 patients with mCXCL12 ranging from 42.7%–65.2% were allocated to group B. Tumor DNA samples of each group were pooled and measured tenfold with two different input amounts (30 ng and 6 ng, respectively) as displayed in Figure 5B. Mean mCXCL12 was independent from the amount of DNA applied (T = 0.23, p = 0.82 for pool A and T = 0.62, p = 0.55 for pool B). The coefficients of variance of paired results neither differed in pool A nor in pool B (Levene test of homoscedastics: F = 1.76, p = 0.20 for pool A and F = 0.25, p = 0.62 for pool B, respectively).
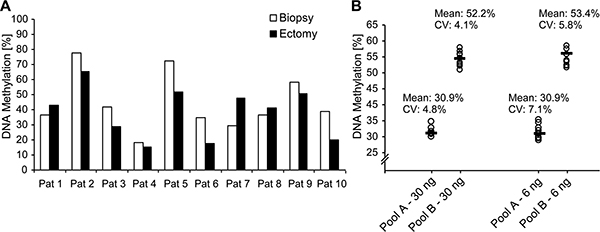
Figure 5: CXCL12 DNA methylation on matching biopsies and ectomy samples from ten PCa patients. (A) Performance evaluation of the CXCL12 DNA methylation assay with two DNA pools. (B) Pool A was comprised of the five low methylated ectomy samples and pool B of the five highly methylated ectomy samples. Ten replicates were measured for each pool with two different input amounts (6 and 30 ng per reaction).
DISCUSSION
Recent studies have attracted notice to the CXCR4/CXCL12 axis in metastasized PCa [24, 25]. In breast and colon cancer, lower intrinsic levels of the chemoattractant chemokine CXCL12 due to hypermethylation of its promoter disrupt cellular feedback mechanisms to internalize membranous CXCR4 [14, 15]. This, in consequence, results in higher tumor cell motility and an augmented capability for metastasis [14, 15]. Our data support the postulate, that CXCL12 methylation down-regulates tumor intrinsic CXCL12 protein expression in PCa as well, thereby fostering metastasis.
Due to its negative effect on CXCR4 expression, tumor intrinsic CXCL12 may be a valuable biomarker in clinical situations that require decision-making. Immunohistochemical CXCL12 staining, however, did not discriminate between PCa with favorable and adverse outcome in the present study. Abnormal methylation of the CXCL12 gene, in contrast, was associated with a shorter BCR-free survival of primary PCa. In two independent cohorts of patients after radical prostatectomy, CXCL12 hypo- and hypermethylation served as a prognostic biomarker identifying patients with early BCR and qualified as an independent prognostic factor in the testing cohort.
The fact that the cohort is based on radical prostatectomy specimens, however, is a major limitation of this study. Gene methylation can be robustly determined even in little formalin-fixed pathological material [22, 23]. If the prognostic value of CXCL12 methylation can be transferred to a biopsy-based cohort, its analysis may be a promising tool in order to identify patients with innocuous tumors that can safely be followed by active surveillance protocols. Conversely, aberrant methylation of CXCL12 may indicate a high metastatic potential. In consequence, affected patients possibly will benefit from radical treatment, including radical prostatectomy. In a paired analysis of CXCL12 methylation based on biopsy and matched ectomy specimens, we were able to show that the results of mCXCL12 assessments are concordant. In addition, our assay seems to be able to reliably determine CXCL12 methylation in biopsy material. The variation coefficients of our results were slightly higher the lower the amount of applied DNA was, however, in homoscedastic testing, no difference could be observed. In practice, growing variances in small samples may easily be counteracted by increasing the number of replicates in the analysis. If the assay will be established for future routine use, a broad spectrum of biopsy material will be needed to precisely set the measuring range and achieve tight tolerances.
Immunotherapy has been emerging as a promising treatment strategy in cancer patients [26–28]. Among major immune checkpoints, e.g. PD-1 and CTLA4, the pharmacologic blockage of CXCR4 and its endogenous ligand CXCL12 has appeared as a prime target for blocking strategies in the treatment of cancer patients [10]. Recent studies have focused on pharmacological CXCR4 inhibition within the cross relation of anti-PD-1/anti-PD-L1 drug resistance [29, 30]. In vitro and murine in vivo studies have provided data that ADM3100 induced blockage of CXCR4 restores sensitivity to PD-1 and CTLA-4 checkpoint inhibitors in anti-PD1 and anti-CTLA-4 drug-resistant tumors and thus might help to overcome resistance to immune therapy [10, 18, 30]. Consequently, a combined anti-CXCR4/CXCL12 and anti-PD1/PD-L1 (Ulocuplumab, BMS-936564/Nivolumab, BMS-936558) therapy is under clinical investigation in metastatic solid tumors (CXCessoR4 trial, ClinicalTrials.gov Identifier: NCT02472977). In our study, patients with high PD-L1 expression and aberrant CXCL12 methylation presented with significantly shorter BRC-free survival intervals than patients with either low PD-L1 expression or high PD-L1 expression plus normal CXCL12 methylation. Accordingly, a combined anti-PD-L1/PD and antagonistic CXCR4/CXCL12 treatment seems to be a promising approach in PCa as well. For the identification of eligible patients, methylation of CXCL12 together with immunohistochemical staining of PD-L1 might be a powerful tool. Whether carcinomatous CXCL12 methylation predicts responsiveness to therapeutic intervention targeting the CXCR4/CXCL12 axis remains the object of future research.
MATERIALS AND METHODS
Subjects
Patients and clinical endpoint
1. The test case control series included 66 FFPE tissue samples from prostate tissue of 25 PCa patients. The samples included 25 prostate cancers, 24 normal adjacent tissue (NAT) and 22 benign prostatic hyperplasia (BPH) specimens.
2. Patient training cohort: A patient cohort comprised of 247 patients with histologically confirmed adenocarcinoma of the prostate who underwent radical prostatectomy at the University Hospital Bonn between 1998 and 2008 was retrospectively enrolled. BCR-free survival was considered as the primary endpoint of the study.
3. Patient testing cohort: The TCGA cohort provided data of 498 patients with histologically confirmed adenocarcinoma of the prostate. Furthermore, transcription data were available from additional 50 specimens obtained from patients with simultaneous PCa. BCR-free survival was considered as the primary endpoint of the study.
4. The biospy test case series included 10 patients with primary PCa diagnosed at the University Hospital Bonn based on 4 to 12 transrectal core needle biopsies. All patients underwent radical prostatectomy during the course of their disease.
Choice of illumina human methylation450 bead
CXCL12 methylation was available from 498 PCa specimens and 50 normal tissues. Clinical follow-up was provided in 417 individuals (mean follow-up period 22 months, range 1–115 months). The TCGA methylation data has been created by TCGA Research Network: http://cancergenome.nih.gov/ using the Infinium HumanMethylation450 BeadChip (Illumina, Inc., San Diego, CA, USA). Methylation values for each bead pair comprised of a variant specific for the methylated and the unmethylated status, respectively, were calculated using the formula 100*Intensity_Bead_Methylated/(Intensity_Bead_Methylated + Intensity_Bead_Unmethylated). Bead pair cg26267854 located within the upstream CpG-island of the CXCL12 promoter was selected.
Ethics
The Institutional Review Board (IRB) at the University Hospital of Bonn approved the study (Lfd. Nr. 071/14). Informed consent has been obtained from all patients that were included in the TCGA cohort in accordance with the Helsinki Declaration of 1975.
Sample preparation
For methylation analysis, ectomy samples were processed according to the InnuConvert Bisulfite All-In-One Kit (Analytik Jena, Germany) as previously published [31]. Bisulfite DNA from biopsies were prepared as described earlier [23]. Length of tumor infiltrate in one tissue biopsy ranged from 0.1 mm to 12 mm (median 4 mm). Number of tumorous cores in one set of biopsies ranged from 1–8 (median 4). Tumorous tissue was marked, micro-dissected, and further processed as described above. For comparison with methylation results obtained from matched biopsies, total tumorous tissue from radical prostatectomy specimens was micro-dissected and bisulfite converted.
For the analytical performance verification of the assay, a dilution series of bisulfite-converted artificially methylated DNA (CpGenomeTM Universal Methylated DNA; Merck Millipore, Darmstadt, Germany) and unmethylated DNA (NW Andrology & Cryobank Inc., Spokane, WA, USA) was used.
CXCL12 quantitative methylation-specific real-time PCR
Gene methylation of CXCL12 was quantified by a quantitative-methylation real-time PCR assay, with the primers and probes shown in Table 4. The assay was performed using an AB 7500 Fast Real-Time PCR System (Life Technologies Corporation, Carlsbad, CA, USA). If not stated otherwise in the result section, DNA input was 25 ng of bisulfite-converted DNA from FFPE ectomy tissue for each PCR reaction. 5 μl DNA from biopsy samples was applied without prior quantification. A calibrator sample (50:50 mixture of bisulfite-converted artificially methylated and unmethylated DNA) was used. Each sample was measured in triplicate. Sample measurements were considered valid when the following quality criterion was met:
Table 4: Primer and probe sequences of the quantitative CXCL12 methylation-specific real-time PCR
Primer/Probe name |
Primer/Probe sequence |
CXCL12-F |
5′- attggagagattgaggatttta -3′ |
CXCL12-R |
5′- aaactaacttatttacttttcatta -3′ |
CXCL12-P-M |
5′- Atto647N-cgccatcgaaaaaccccgtccc-BHQ-2 -3′ |
CXCL12-P-U |
5′- HEX-caccatcaaaaaaccccatccc-BHQ-1 -3′ |
CTCXCL12-P-U or CTCXC12-P-M < 35
Other samples were excluded from the analysis due to insufficient DNA yield. The CXCL12 methylation was calculated using the ΔΔCT method:
ΔCT = ΔCTCXCL12-P-U – ΔCT CXCL12-P-M, ΔΔCT = ΔCTsample – ΔCTcalibrator, mCXCL12 = 100%/(1 + 2ΔΔCT).
Immunohistochemical analyses of PD-L1 expression and CXCL12 expression
PD-L1 expression data were collected from our previous study [21]. In brief, the anti-PD-L1 antibody clone EPR1161(2) (Abcam, Cambridge, UK) was used. Specific membranous and cytoplasmic staining of epithelial tumor cells was considered positive [21]. The intensity of PD-L1–staining was scored semi-quantitatively as negative (0), weak (1), moderate (2), or strong (3) by two independent and blinded pathologists. For each patient a PD-L1 expression value was computed. Up to five cores per patient were scored semiquantitatively and mean averaged. Patients’ stratification for Kaplan-Meier analysis was conducted using the median PD-L1 expression of the total cohort.
For immunohistochemical CXCL12 staining, TMA blocks were cut (3 μm) and mounted on Superfrost slides (Menzel Gläser, Braunschweig, Germany). A polyclonal anti-CXCL12 antibody (bios #bs-4938R, 1:600) was used. A Medac Autostainer 480 performed the staining (Medac, Wedel, Germany). Antigen-antibody-binding was visualized by horse-radish-peroxidase polymer method. To assess the specificity of staining, negative controls with PBS were run.
Ninety-one cases could be evaluated in immunohistochemical CXCL12 staining. CXCL12 staining was observed in the nuclei as well as in the cytoplasma of basal cells of PCa glands. Nuclear and cytoplasmic staining was assessed as negative (0), weak (1), or strong (2). Scores were added and averaged over 2–3 tissue scores. For correlation analysis and survival analysis, scores were dichotomized by the median of the total cohort.
Statistical analyses
Statistical analysis was performed using SPSS, version 22 (SPSS Inc., Chicago, IL). The relationship between input DNA methylation and measured DNA methylation was assessed by linear regression. In the training and testing cohort, comparative studies of dichotomized CXCL12 methylation values were tested using the Mann-Whitney U test for paired analysis and using the Kruskal-Wallis test in cases when more than two groups were compared. Statements regarding potential correlations were made using the Spearman’s correlation coefficient ranked between two assessed variables. Comparative analysis of methylation in the biopsy case test series were performed using the Pearson correlation coefficient. Diluted tumor DNA of the biopsy case test series were analysed using the t-test and the Levene test of homoscedastics. BCR-free survival estimates were calculated according to Kaplan-Meier and Cox proportional hazards regression analysis. Multivariate analysis was performed with all parameters tested significant in the univariate analysis. P-values (two-sided) lower than 0.05 were considered significant. The test case control series was analysed using GraphPadPrism (GraphPad, La Jolla, CA). For comparison of independent groups of the test case control study, one-way analysis of variance (one-way ANOVA including Barlett’ statistic) with Bonferroni-post-hoc testing was applied.
CONFLICTS OF INTEREST
Dimo Dietrich has been an employee and is a stockholder of Epigenomics AG, a company that aims to commercialize the DNA methylation biomarkers SEPT9 and SHOX2. Dimo Dietrich is inventor and co-inventor and owns patents on methylation biomarkers and related technologies. Some of these patents are commercially exploited by Epigenomics AG. Dimo Dietrich receives inventor’s compensation from Epigenomics AG. Dimo Dietrich is a consultant for AJ Innuscreen GmbH (Berlin, Germany), a 100% daughter company of Analytik Jena AG (Jena, Germany), and receives royalties from product sales. Dimo Dietrich is a consultant and receives or received compensation from Therawis GmbH (Munich, Germany), Oncgnostics GmbH (Jena, Germany), MDxHealth, Inc. (Irvine, CA, USA), Epigenomics AG (Berlin, Germany) and R-Biopharm AG (Darmstadt, Germany).
GRANT SUPPORT
The study has been funded by the University Hospital of Bonn.
REFERENCES
1. Furusato B, Mohamed A, Uhlen M, Rhim JS. CXCR4 and cancer. Pathol Int. 2010; 60:497–505.
2. Zlotnik A. New insights on the role of CXCR4 in cancer metastasis. J Pathol. 2008; 215:211–213.
3. Muller A, Homey B, Soto H, Ge N, Catron D, Buchanan ME, McClanahan T, Murphy E, Yuan W, Wagner SN, Barrera JL, Mohar A, Verastegui E, et al. Involvement of chemokine receptors in breast cancer metastasis. Nature. 2001; 410:50–56.
4. Vandercappellen J, Van Damme J, Struyf S. The role of the CXC chemokines platelet factor-4 (CXCL4/PF-4) and its variant (CXCL4L1/PF-4var) in inflammation, angiogenesis and cancer. Cytokine Growth Factor Rev. 2011; 22:1–18.
5. Balkwill F. The significance of cancer cell expression of the chemokine receptor CXCR4. Semin Cancer Biol. 2004; 14:171–179.
6. Darash-Yahana M, Pikarsky E, Abramovitch R, Zeira E, Pal B, Karplus R, Beider K, Avniel S, Kasem S, Galun E, Peled A. Role of high expression levels of CXCR4 in tumor growth, vascularization, and metastasis. FASEB J. 2004; 18:1240–1242.
7. Teicher BA, Fricker SP. CXCL12 (SDF-1)/CXCR4 pathway in cancer. Clin Cancer Res. 2010; 16:2927–2931.
8. Domanska UM, Kruizinga RC, Nagengast WB, Timmer-Bosscha H, Huls G, de Vries EG, Walenkamp AM. A review on CXCR4/CXCL12 axis in oncology: no place to hide. Eur J Cancer. 2013; 49:219–230.
9. Hsiao JJ, Ng BH, Smits MM, Wang J, Jasavala RJ, Martinez HD, Lee J, Alston JJ, Misonou H, Trimmer JS, Wright ME. Androgen receptor and chemokine receptors 4 and 7 form a signaling axis to regulate CXCL12-dependent cellular motility. BMC Cancer. 2015; 15:204.
10. Scala S. Molecular Pathways: Targeting the CXCR4-CXCL12 Axis-Untapped Potential in the Tumor Microenvironment. Clin Cancer Res. 2015; 21:4278–4285.
11. Bilgin YM, de Greef GE. Plerixafor for stem cell mobilization: the current status. Curr Opin Hematol. 2016; 23:67–71.
12. Roland J, Murphy BJ, Ahr B, Robert-Hebmann V, Delauzun V, Nye KE, Devaux C, Biard-Piechaczyk M. Role of the intracellular domains of CXCR4 in SDF-1-mediated signaling. Blood. 2003; 101:399–406.
13. Marchese A. Endocytic trafficking of chemokine receptors. Curr Opin Cell Biol. 2014; 27:72–77.
14. Wendt MK, Cooper AN, Dwinell MB. Epigenetic silencing of CXCL12 increases the metastatic potential of mammary carcinoma cells. Oncogene. 2008; 27:1461–1471.
15. Wendt MK, Johanesen PA, Kang-Decker N, Binion DG, Shah V, Dwinell MB. Silencing of epithelial CXCL12 expression by DNA hypermethylation promotes colonic carcinoma metastasis. Oncogene. 2006; 25:4986–4997.
16. Zhi Y, Chen J, Zhang S, Chang X, Ma J, Dai D. Down-regulation of CXCL12 by DNA hypermethylation and its involvement in gastric cancer metastatic progression. Dig Dis Sci. 2012; 57:650–659.
17. Zhou W, Jiang Z, Song X, Liu Y, Wen P, Guo Y, Xu F, Kong L, Zhang P, Han A, Yu J. Promoter hypermethylation-mediated down-regulation of CXCL12 in human astrocytoma. J Neurosci Res. 2008; 86:3002–3010.
18. Fearon DT. The carcinoma-associated fibroblast expressing fibroblast activation protein and escape from immune surveillance. Cancer Immunol Res. 2014; 2:187–193.
19. Topalian SL, Hodi FS, Brahmer JR, Gettinger SN, Smith DC, McDermott DF, Powderly JD, Carvajal RD, Sosman JA, Atkins MB, Leming PD, Spigel DR, Antonia SJ, et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N Engl J Med. 2012; 366:2443–2454.
20. Macoska JA, Begley LA, Dunn RL, Siddiqui J, Wei JT, Sarma AV. Pilot and feasibility study of serum chemokines as markers to distinguish prostatic disease in men with low total serum PSA. Prostate. 2008; 68:442–452.
21. Gevensleben H, Dietrich D, Golletz C, Steiner S, Jung M, Thiesler T, Majores M, Stein J, Uhl B, Mueller S, Ellinger J, Stephan C, Jung K, et al. The immune checkpoint regulator PD-L1 is highly expressed in aggressive primary prostate cancer. Clin Cancer Res. 2015.
22. Dietrich D, Hasinger O, Banez LL, Sun L, van Leenders GJ, Wheeler TM, Bangma CH, Wernert N, Perner S, Freedland SJ, Corman JM, Ittmann MM, Lark AL, et al. Development and clinical validation of a real-time PCR assay for PITX2 DNA methylation to predict prostate-specific antigen recurrence in prostate cancer patients following radical prostatectomy. J Mol Diagn. 2013; 15:270–279.
23. Jung M, Uhl B, Kristiansen G, Dietrich D. Bisulfite Conversion of DNA from Tissues, Cell Lines, Buffy Coat, FFPE Tissues, Microdissected Cells, Swabs, Sputum, Aspirates, Lavages, Effusions, Plasma, Serum, and Urine. Methods Mol Biol. 2015.
24. Chen Q, Zhong T. The association of CXCR4 expression with clinicopathological significance and potential drug target in prostate cancer: a meta-analysis and literature review. Drug Des Devel Ther. 2015; 9:5115–5122.
25. Delongchamps NB, Beuvon F, Mathieu JR, Delmas S, Metzger I, Prats H, Cabon F. CXCR4 is highly expressed at the tumor front but not in the center of prostate cancers. World J Urol. 2015; 33:281–287.
26. Pardoll DM. The blockade of immune checkpoints in cancer immunotherapy. Nat Rev Cancer. 2012; 12:252–264.
27. Sharma P, Allison JP. The future of immune checkpoint therapy. Science. 2015; 348:56–61.
28. Postow MA, Callahan MK, Wolchok JD. Immune Checkpoint Blockade in Cancer Therapy. J Clin Oncol. 2015; 33:1974–1982.
29. Joyce JA, Fearon DT. T cell exclusion, immune privilege, and the tumor microenvironment. Science. 2015; 348:74–80.
30. Feig C, Jones JO, Kraman M, Wells RJ, Deonarine A, Chan DS, Connell CM, Roberts EW, Zhao Q, Caballero OL, Teichmann SA, Janowitz T, Jodrell DI, et al. Targeting CXCL12 from FAP-expressing carcinoma-associated fibroblasts synergizes with anti-PD-L1 immunotherapy in pancreatic cancer. Proc Natl Acad Sci USA. 2013; 110:20212–20217.
31. Holmes EE, Jung M, Meller S, Leisse A, Sailer V, Zech J, Mengdehl M, Garbe LA, Uhl B, Kristiansen G, Dietrich D. Performance evaluation of kits for bisulfite-conversion of DNA from tissues, cell lines, FFPE tissues, aspirates, lavages, effusions, plasma, serum, and urine. PLoS One. 2014; 9:e93933.

