INTRODUCTION
Aristolochic acid nephropathy (AAN) is a rapidly progressive acute or chronic tubulointerstitial nephritis (TIN) that occurs during end-stage renal failure and urothelial malignancy [1]. Patients taking aristolochic acid have a high risk (~50%) of upper urothelial tract carcinoma as well as bladder urothelial carcinoma within a few years; such cases have occurred in Europe and Asia [2, 3]. The predominant lesion of AAN is characterised by tubulointerstitial fibrosis, with tubular atrophy atrophy/loss and global glomerular sclerosis [2, 4, 5]. Apoptosis, epithelial-mesenchymal transition (EMT), and fibrosis are the main pathologic manifestations of the development of AAN. DNA damage, cell cycle arrest, and cell apoptosis have been studied in an AA-induced proximal tubular epithelial cell line (LLC-PK1) as well as proximal tubular epithelial cells (PTCs). Inflammation is also one of the mechanisms involved in AAN. TGF-/Smad3 may play a critical role in AAN. A previous study by Zhou et al [6]., found that AA could activate Smad signalling to mediate EMT and renal fibrosis through TGF-dependent and JNK/MAP kinase-dependent mechanisms. p53 is involved in the apoptotic injury seen in AAN. Previous studies also showed that AA stimulated the p53 pathway, mediating TEC apoptosis in acute AAN [7, 8]. Moreover, AA-DNA adducts are an AA exposure marker and risk factors of AA nephropathy-associated cancer [9]. AA commonly induces tumours in the renal cortex, renal pelvis and urinary bladder. Furthermore, some tumours have also been discovered in the forestomach, lungs, uterus and lymphoid organs [10]. However, the underlying pathogenesis of AAN remains largely unclear, which has impeded the development of effective therapies for AAN patients. Currently, a synthesize assay method is urgently needed that can identify the targets of AAN, and What 's more, explore molecular mechanisms of AAN, is necessary.
In disease network, system biology concentrates on complex molecular interaction, instead of changing single molecular component. Network biology analysis can discovery individual molecular component has little affect disease network. Network pharmacology, which is based on system biology and polypharmacology, provides a novel dominant paradigm for drug design that encompasses the network mode of “multiple targets, multiple effects, complex diseases” [11, 12]. Interestingly, research ideas of network pharmacology is correspond with the holistic and systemic views of Traditional Chinese medicine (TCM) [13, 14]. TCM network pharmacology provides a comprehensive analysis for TCM Formula, and TCM-based drug discovery [15]. For example, Li shao et al [16]., employed a network pharmacology method to identify bioactive ingredients and action mechanisms of Ge-Gen-Qin-Lian decoction for Type 2 Diabetes therapy. Moreover, network pharmacology can speculate and appraise toxicity and side effects of drugs by adopting the network analysis strategy. Thus, network pharmacology has become a powerful tool for identifying drug target, and then uncovering the therapeutic and toxicological mechanisms of TCM formulae. This shall promote modern study and development of TCM.
MicroRNAs (miRNAs) are a class of small (~22 nucleotides) non-coding RNAs that regulate gene expression by causing translational repression or degradation of target mRNA at the post-transcriptional level [17, 18]. miRNAs play key roles in a variety of biological process, such as cell apoptosis, proliferation, differentiation, development and tumorigenesis, and are involved in the development of many diseases, including cancer, nephropathy and renovascular diseasesuch as cell apoptosis, proliferation, differentiation, development and tumorigenesis, and involved in development of many diseases including cancer, nephropathy and renovascular disease [19–21]. Many studies have found that miRNAs play a role in the progression of renal disease. For example, miR-34a suppresses renal cancer cell growth, tube formation and metastasis [22], and miR-24 promotes renal ischemic injury by inducing apoptosis in endothelial and tubular epithelial cells [23].However, few previous studies have emphasised the value of circulating miRNAs as potential biomarkers for AAN therapy.
In the study, network pharmacology method was employed to unwind miRNA complexity in the progression of AAN. First, we analyzed the miRNA expression profiles from HK-2 normal samples and HK-2 samples treated with AA to identify DE-miRNAs using microarray analysis, and further the target genes of DE-miRNAs were predicted. Next, we performed a network analysis and functional analysis of these DE-miRNAs to uncover their roles in the progression of AAN. Finally, 6 DE-miRNAs (has-miR-192, has-miR-194, has-miR-542-3p, has-miR-450a, has-miR-584, has-miR-33a) were screened as potential biomarkers for AAN therapy. Our results may provide novel insight into the regulatory mechanisms of miRNAs in the progression of AAN and be useful for the prevention and treatment of AAN.
RESULTS
Screening of DE-miRNA
After preprocessing, we obtained miRNA expression data that composed of 6 samples and1222 miRNAs. The miRNA expression data was analyzed using SAM. According to the threshold of FDR<0.05, we identified 11 DE-miRNAs. The information for 11 DE-miRNAs were listed in Table 1.
Table 1: Eleven differentially expressed miRNAs
No. |
miRNA |
Fold change |
1 |
hsa-miR-33a |
6.59 |
2 |
hsa-miR-450a |
5.99 |
3 |
hsa-miR-194 |
6.30 |
4 |
hsa-miR-192 |
8.35 |
5 |
hsa-miR-4730 |
5.26 |
6 |
hsa-miR-542-3p |
5.19 |
7 |
hsa-miR-4747-3p |
4.21 |
8 |
hsa-miR-1234 |
4.51 |
9 |
hsa-miR-584 |
6.27 |
10 |
hsa-miR-3663-3p |
5.11 |
11 |
hsa-miR-3135b |
4.20 |
Note: FDR<0.05.
Construction of differentially expressed miRNA-gene regulation networks and miRNA-pathway regulation networks
To increase the reliability of the predicted DE-miRNA targets, we selected target genes that were stored in at least five databases. We predicted the target genes of each DE-miRNA and constructed a network using Cytoscape (Figure 1). The miRNA -gene regulation network consisted of 988 nodes for target genes, 7 nodes for DE-miRNAs, and 1074 regulatory relationships between 7 DE-miRNAs and 988 target genes. The yellow squares represent miRNA. Based on the numbers of DE-miRNAs regulating the target genes, the target genes are represented as bright green, light blue, orange and red circles. In the network, we observed certain target genes that were regulated by more than one miRNA. For example, GPR158 were regulated by hsa-miR-450a, hsa-miR-194, hsa-miR-584, and hsa-miR-33a.
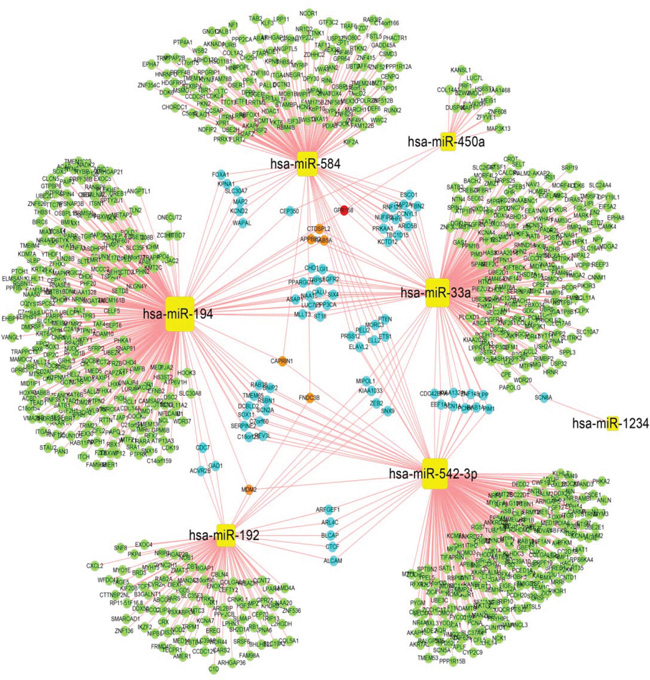
Figure 1: Regulatory network of differentially expressed miRNA-genes. In the miRNA-gene regulation network, yellow squares denote miRNAs. Target genes are represented in bright green, light blue, orange and red circles based on the number of regulatory relationships between the miRNAs and target genes. Bright green indicates one regulatory relationship, light blue denotes two regulatory relationships, orange denotes three regulatory relationships, and red indicates four regulatory relationships.
We performed pathway enrichment analysis of DE-miRNA target genes using the hypergeometric distribution test. Biological pathways with significant values (p<0.05) were identified as significant pathways. We identified the significant pathways for each DE-miRNA and built a miRNA-pathway regulation network using Cytoscape (Figure 2). In the network, 92 nodes were significant pathways, 6 nodes were DE-miRNAs, and 138 edges represented the regulatory relationships between significant pathways and DE-miRNAs. The DE-miRNAs are shown as yellow squares and the coloured circles denote target genes in light blue, pink, green and red based on the number of regulatory relationships between the miRNAs and target genes. Figure 2 shows that the pathways associated with inflammatory and immune functions were regulated by miRNAs, such as cytokine-cytokine receptor interactions, Wnt signalling pathways, MAPK signalling pathways, NOD-like receptor signalling pathways, B cell receptor signalling pathways and T cell receptor signalling pathways. Additionally, 12 cancer-related pathways were also represented, specifically prostate cancer, basal cell carcinoma, pathways in cancer, glioma, renal cell carcinoma, endometrial cancer, acute-myeloid leukaemia, colorectal cancer, pancreatic cancer, melanoma, small cell lung cancer, and non-small cell lung cancer. Furthermore, cell cycle, calcium signalling pathway, p53 signalling pathway, insulin signalling pathway, apoptosis, and others were also significantly regulated by miRNAs.
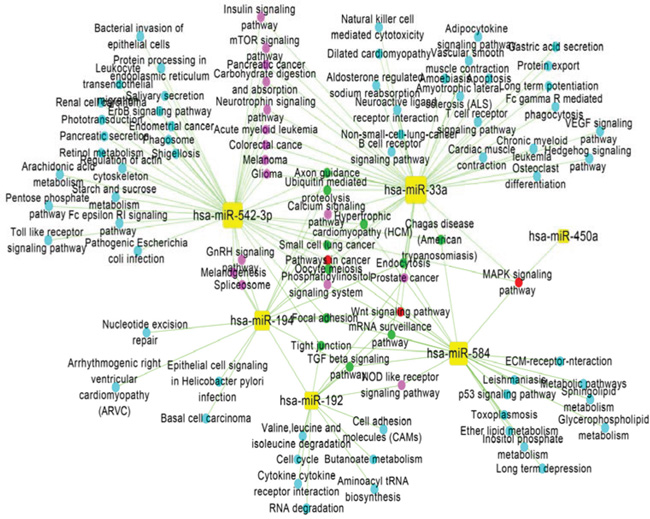
Figure 2: Regulatory-network analysis of miRNA-pathways. In the miRNA-pathway regulation network, yellow squares represent miRNAs, and the coloured circles denote pathways in light blue, pink, green and red based on the number of regulatory relationships between significant pathways and miRNAs. Light blue represents one regulatory relationship, pink represents two regulatory relationships, green represents three regulatory relationships, and red represents four regulatory relationships.
Given the miRNA-gene regulation network and the miRNA-pathway regulation network (Figures 1 and 2), information about the degrees of DE-miRNAs is shown in Figure 3; we found that has-miR-194 (305 genes and 15 pathways) had the greatest number of links to adjacent target genes, and has-miR-33a (230 genes and 43 pathways) had the greatest number of connections to adjacent pathways. Other miRNAs, including hsa-miR-542-3p (243 genes and 42 pathways), hsa-miR-584 (175 genes and 24 pathways), hsa-miR-192 (104 genes and 14 pathways), and hsa-miR-450a (17 genes and 1 pathways), also had connections with adjacent target genes and pathways.
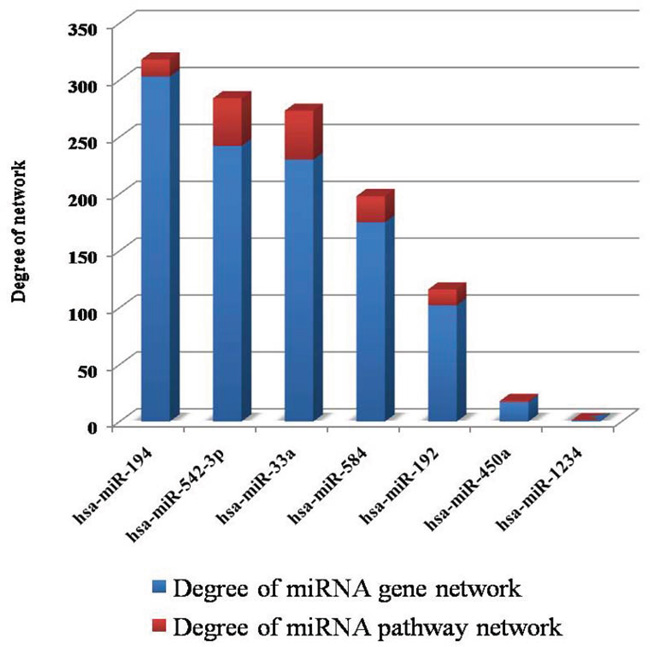
Figure 3: Information for the degrees of the DE-miRNAs. In the two networks, the degrees represent the numbers of connections between miRNAs and target genes/pathways.
Functional analysis of target genes of DE-miRNA
We performed GO biological process (BP) annotation analysis of DE-miRNA target genes using the hypergeometric distribution test. The GO terms with significant values (p<0.05) were identified as GO biological process. We identified the significant GO terms for each DE-miRNA and plotted the distribution of the GO biological process annotations (Figure 4). GO terms for hsa-miR-192, hsa-miR-542-3P, and hsa-miR-450a that were classified as biological processes by WEGO. The biological process terms suggested that hsa-miR-192 and hsa-miR-542-3p were mainly correlated with biological regulation, cell cycle, cellular metabolic process, cellular process, metabolic process, and others (Figure 4A. The biological process terms suggested that hsa-miR-33a, hsa-miR-194, and hsa-miR-584 are mainly involved in biological regulation, cellular metabolic process, cellular process, metabolic process, pigmentation (Figure 4B).
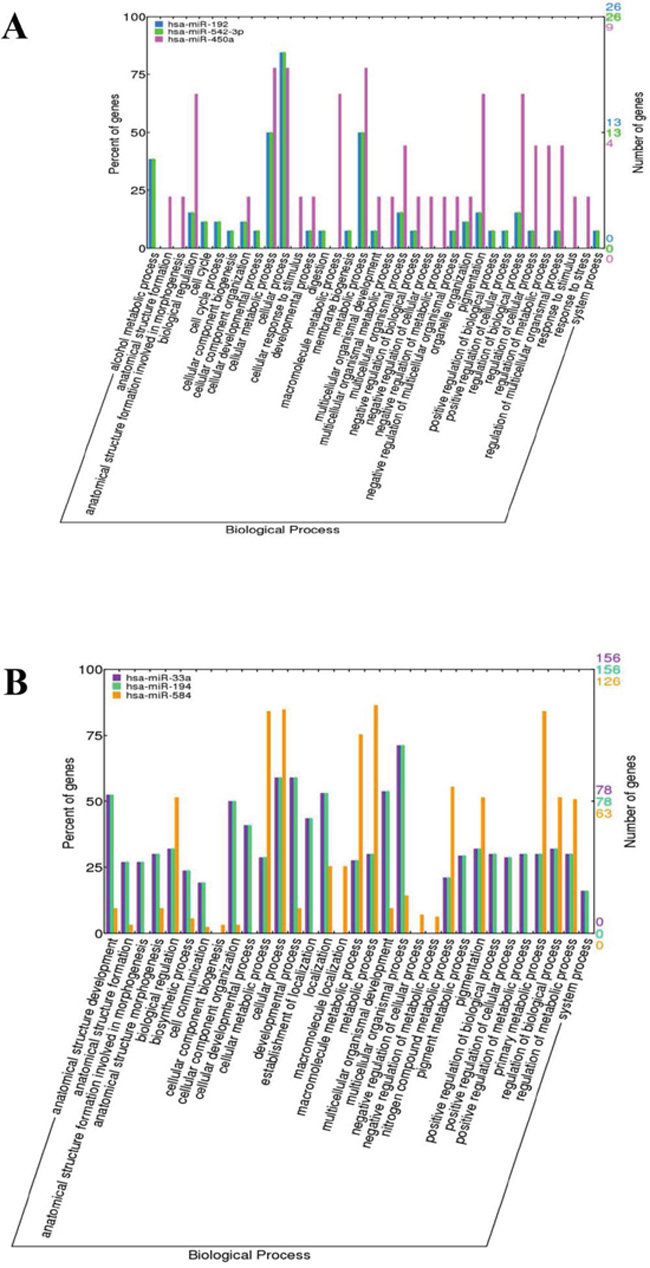
Figure 4: Distribution of the GO terms for the target genes of DE-miRNAs. A. The GO terms for the target genes of hsa-miR-192, hsa-miR-542-3P, and hsa-miR-450a were classified into biological process by WEGO. B. The GO terms for the target genes of hsa-miR-33a, hsa-miR-194, and hsa-miR-584 were classified into biological processes by WEGO.
DISCUSSION
In the study, we employed bioinformatics methods to identify 6 DE-miRNA(has-miR-192, has-miR-194, has-miR-542-3p, has-miR-450a, has-miR-584, and has-miR-33a) in AAN, which could better our understandings of the pathogenesis of AAN and provide potential biomarkers for diagnosis and treatment. miRNAs perform a complex role in cell cycle arrest induced by the increased activation of p53, which is a major feature of AAN [24, 38–40]. The miR-192 and miR-194 affect cell cycle progression, which is induced by p53 [41–44]. Jenkins et al [24]., noted the over-expression of has-miR-192, has-miR-194, has-miR-450a and has-miR-542-3p in AA-induced G2/M arrest in proximal tubular epithelial cells (PTCs), and our study indicated that the has-miR-192over-expression is recapitulated in G2/M arrest in AAN. has-miR-194 was significantly up-regulated in acute kidney impairment and down-regulated in the growth arrest injury in carcinogenesis; for example, miR-194 is often down-regulated in colorectal cancer, renal childhood neoplasms, and multiple myeloma [45–48]. Previous studies have indicated that the expression of has-miR-194 may cause G1 and G2 arrest in a p53-dependent manner [41, 43]. The above studies suggested that altered has-miR-194 expression is most likely to contribute to the progression of AAN. miR-542-3p is a tumour suppressor that improves p53 stability by promoting the regulatory relationship between p53 and MDM2, a negative regulator of p53 [49]. Yoon et al [50]., demonstrated that the enforced expression of miR-542-3p caused the inhibition of cell proliferation and apoptosis by inducing G1 and G2/M cell cycle arrest. has-miR-450a targeted the MAPK signalling pathway in HK-2 cells treated with AA, and the MAPK signalling pathway results in the fibrosis that accompanies the tubular atrophy/loss that is the main lesion in AAN [51–53]. Moreover, has-miR-450a was reported to be significantly decreased in hepatocellular carcinoma as well as ectopically; its expression leads to cell proliferation inhibition through DNMT3a [54]. has-miR-33a is a tumour suppressor that inhibits cell cycle progression [55]. A previous study found that miR-33a transfection led to reduced cell proliferation in colon carcinoma cells, and miR-33a inhibition increased cisplatin-induced apoptosis by antagomir-33a in osteosarcoma cells [56, 57]. Additionally, miR-33a inhibits expression of the insulin receptor substrate 2, which is involved in the insulin-signalling pathway in the liver [58]. miR-584 has been reported to be inhibitd in kidney cancer and breast cancer [59, 60].miR-584 has anti-inflammatory effects that modulate AA functions [61].Subsequently, miR-584 regulated pathways such as MAPK signalling pathways, pathways in cancer, Wnt signalling pathways, p53 signalling pathways, and TGF-beta signalling pathways are associated with AAN [9, 53, 62–66]. Thus, has-miR-33a and has-miR-584 may be biomarkers for the diagnosis and prognosis of AAN, which must be confirmed in further experiments.
In addition, MDM2 is target gene for has-miR-192, has-miR-194, and has-miR-542-3p. It was shown that the expression of MDM2, a negative regulator of p53, was suppressed by AA, while has-miR-192 was significantly up-regulated. The analysis suggested that over-expression of has-miR-192 resulted in G2/M cell cycle arrest by inhibiting MDM2 expression via the increased activity of p53. In AAN, the expression of MDM2 is down-regulated, but has-miR-192 is over-expressed in renal cells after AA treatment, indicating that MDM2 is related to cell cycle and AAN-associated cancers [4]. RB1 is target gene for has-miR-192. RB1 is a tumour suppressor gene and a negative regulator of the cell cycle [67, 68]. A study by Chang [69]., suggested that AA-induced urothelial proliferation in rats via cell-cycle progression was mediated via the induction of cyclin D1/cdk4 and/or cyclinE/cdk2 activity and the increasing phosphorylation of Rb. Volker et al [70]., also confirmed that RB1 is often affected in human urothelial cancer. CDC7 is the G1 and G2 checkpoint protein of has-miR-192 and contains has-miR-194. It encodes a cell division cycle protein with kinase activity that is critical for maintaining genomic stability via the S-phase checkpoint pathways in response to DNA damage, which can be associated with AAN [4].
This study has some limitations. First, the retrieval of miRNA target genes may not have been complete. Although we retrieved target genes of miRNAs based on 10 databases, the identified target genes might include false positives. Second, current material on AAN-related miRNAs is limited, and their regulatory mechanisms in the development of AAN are still largely unclear. Third, the sample sizes for the expression profiling were not large. We hope that additional high-throughput data can be obtained. Despite these limitations, the study still provides insight into miRNA regulation in the developmental process of AAN.
MATERIALS AND METHODS
Microarray analysis
The miRNA expression profile of GSE53911 was downloaded from the GEO database (http://www.ncbi.nlm.nih.gov/geo/), which was amassed by Jenkins et al [24]. A total of six human proximal tubular epithelial cell line (HK-2) samples were selected, including 3 HK-2 cells without AA treatment (used as controls) and 3 HK-2 cells with AA treatment. The annotation information of for the miRNA expression profiles was downloaded from the GPL15446 3D-Gene Human miRNA V17_1.0.0 platform.
Data preprocessing
For the miRNA expression profiles, we first inferred missing values using the K-nearest neighbours method [25].Then, these values were normalised with the limma package in R [26]. The probe level data were finally converted into miRNA names based on the annotation information from the GPL15446 platform.
Differentially expressed miRNAs analysis
To identify the differentially expressed miRNAs (DE-miRNAs), a significance analysis of microarrays (SAM) was used to screen the DE-miRNAs, and miRNAs with a false discovery rate (FDR) threshold of less than 0.05 were designated as differentially expressed.
DE-miRNA target genes prediction
The target genes of DE-miRNA were retrieved from the following miRNA-target databases:DIANA-microT(DT) [27], mirSVR [28], PicTar5(PT5) [29], RNA22 [30], RNAhybrid(Rh) [31], TargetScan (TS) [32], PITA [33], MirTarget2 [34], TargetMiner [35], miRanda (miR) [36]. Target genes were identified in at least 5 out of the 10 databases to decrease the number of false-positive results.
Pathway and functional analysis
We performed pathway and functional analyses for the target genes of each DE-miRNA using the hypergeometric distribution test which can identify significant pathways and GO biological processes. The p value for each pathway or GO term was calculated as follows:
where N denotes the total number of the whole human genome, n is the number of DE-miRNA target genes, M represents the number of genes that are annotated to a pathway or a GO term, and m is the number of annotated DE-miRNA target genes in a pathway or a GO term. Pathways or GO terms with p<0.05 were considered to be significantly enriched and could be regulated by DE-miRNA. We plotted the distribution of the GO annotation using WEGO (http://wego.genomics.org.cn/cgi-bin/wego/index.pl) [37].
CONCLUSIONS
In our study, we first performed differentially expressed miRNA analysis and identified 11 DE-miRNAs associated with AAN. Next, we performed network analysis and functional analysis on the target genes of these DE-miRNAs. Finally, 6 DE-miRNAs (has-miR-192, has-miR-194, has-miR-542-3p, has-miR-450a, has-miR-584, and has-miR-33a) were predicted as potential biomarkers of AAN. has-miR-192 has been shown to be involved in the pathogenesis of AAN. The altered expression of has-miR-194, has-miR-542-3p and has-miR-450a appear to link to the progression of AAN, but how they exert the regulatory effects in AAN progression is still not clear. has-miR-584 and has-miR-33a have known roles in the progression of cancer and can affect the development of AAN. Although these targets were selected, there have not been many reports that have clearly explored their roles, and further studies are need to confirm their roles in AAN progression. Moreover, the genes MDM2 and RB1 were identified as being significantly affected in AAN-urothelial cancer, and the gene CDC7 was identified as being associated with AAN. However, further investigation is required to confirm the function of CDC7.
ACKNOWLEDGMENTS AND FUNDING
This work was supported by a National Natural Science Foundation of China Grant (grant no. 81274175), the Fundamental Research Funds for the Central Universities (grant no. 106112015CDJZR468808), the Start-up Fund for the “Hundred Young-Talent Scheme” Professorship provided by Chongqing University in China (grant no. 0236011104401), a Queensland University of Technology Vice Chancellor’s Research Fellowship Grant (grant no. 150410-0070/08), Open Grants for the Key Instrumental Platform Usage Provided by Chongqing University in China (grant no. 201412150114, 201512150061), the National Natural Science Foundation of China (81173645), Significant New Drugs Creation Special Science and Technology Major of China (2014ZX09201-022-06) and the Program for Innovative Research Team in IMPLAD (PIRTI) as well as Director's Funds provided by Yunnan Branch, Institute of Medicinal Plant Development, Chinese Academy of Medical Sciences and Peking Union Medical College, and a grant (YZYN-14-06) from the Basic Scientific Research Special of the Central Public Welfare Research Institutes of IMPLAD, Chinese Academy of Medical Sciences.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
REFERENCES
1. Vanherweghem JL, Depierreux M, Tielemans C, Abramowicz D, Dratwa M, Jadoul M, Richard C, Vandervelde D, Verbeelen D, Vanhaelen-Fastre R, et al. Rapidly progressive interstitial renal fibrosis in young women: association with slimming regimen including Chinese herbs. Lancet. 1993; 341:387-391.
2. Depierreux M, Van Damme B, Vanden Houte K, Vanherweghem JL. Pathologic aspects of a newly described nephropathy related to the prolonged use of Chinese herbs. American journal of kidney diseases. 1994; 24:172-180.
3. Schmeiser HH, Stiborova M, Arlt VM. Chemical and molecular basis of the carcinogenicity of Aristolochia plants. Current opinion in drug discovery & development. 2009; 12:141-148.
4. Chen YY, Chung JG, Wu HC, Bau DT, Wu KY, Kao ST, Hsiang CY, Ho TY, Chiang SY. Aristolochic acid suppresses DNA repair and triggers oxidative DNA damage in human kidney proximal tubular cells. Oncology reports. 2010; 24:141-153.
5. Baudoux TE, Pozdzik AA, Arlt VM, De Prez EG, Antoine MH, Quellard N, Goujon JM, Nortier JL. Probenecid prevents acute tubular necrosis in a mouse model of aristolochic acid nephropathy. Kidney Int. 2012; 82:1105-1113.
6. Zhou L, Fu P, Huang XR, Liu F, Chung AC, Lai KN, Lan HY. Mechanism of chronic aristolochic acid nephropathy: role of Smad3. American journal of physiology Renal physiology. 2010; 298:F1006-1017.
7. Jankovic Velickovic L, Hattori T, Stefanovic V. Molecular markers in upper urothelial carcinoma associated to Balkan endemic nephropathy. Aristolochic acid as the major risk factor of the worldwide disease. TheScientificWorldJournal. 2009; 9:1360-1373.
8. Zhou L, Fu P, Huang XR, Liu F, Lai KN, Lan HY. Activation of p53 promotes renal injury in acute aristolochic acid nephropathy. Journal of the American Society of Nephrology. 2010; 21:31-41.
9. Karanovic S, Lela IV, Jelakovic B, Dickman KG, Peic AK, Dittrich D, Knezevic M, Matijevic V, Fernandes AS, Miller F. Variation in presentation and presence of DNA adducts and p53 mutations in patients with endemic nephropathy--an environmental form of the aristolochic acid nephropathy. Kidney & blood pressure research. 2013; 37:1-8.
10. Arlt VM, Alunni-Perret V, Quatrehomme G, Ohayon P, Albano L, Gaid H, Michiels JF, Meyrier A, Cassuto E, Wiessler M, Schmeiser HH, Cosyns JP. Aristolochic acid (AA)-DNA adduct as marker of AA exposure and risk factor for AA nephropathy-associated cancer. International journal of cancer. 2004; 111:977-980.
11. Hopkins AL. Network pharmacology: the next paradigm in drug discovery. Nature chemical biology. 2008; 4:682-690.
12. Zhang B, Wang X, Li S. An Integrative Platform of TCM Network Pharmacology and Its Application on a Herbal Formula, Qing-Luo-Yin. Evidence-based complementary and alternative medicine. 2013; 2013:456747.
13. Li S, Zhang B. Traditional Chinese medicine network pharmacology: theory, methodology and application. Chinese journal of natural medicines. 2013; 11:110-120.
14. Li S, Fan TP, Jia W, Lu A, Zhang W. Network pharmacology in traditional chinese medicine. Evidence-based complementary and alternative medicine. 2014; 2014:138460.
15. Liang X, Li H, Li S. A novel network pharmacology approach to analyse traditional herbal formulae: the Liu-Wei-Di-Huang pill as a case study. Molecular bioSystems. 2014; 10:1014-1022.
16. Li H, Zhao L, Zhang B, Jiang Y, Wang X, Guo Y, Liu H, Li S, Tong X. A network pharmacology approach to determine active compounds and action mechanisms of ge-gen-qin-lian decoction for treatment of type 2 diabetes. Evidence-based complementary and alternative medicine. 2014; 2014:495840.
17. Vasudevan S, Tong Y, Steitz JA. Switching from repression to activation: microRNAs can up-regulate translation. Science. 2007; 318:1931-1934.
18. Yates LA, Norbury CJ, Gilbert RJ. The long and short of microRNA. Cell. 2013; 153:516-519.
19. Striteska J, Nekvindova J, Cerny V, Palicka V. [MicroRNAs and kidneys]. Casopis lekaru ceskych. 2014; 153:187-192.
20. Vergho DC, Kneitz S, Kalogirou C, Burger M, Krebs M, Rosenwald A, Spahn M, Loser A, Kocot A, Riedmiller H, Kneitz B. Impact of miR-21, miR-126 and miR-221 as Prognostic Factors of Clear Cell Renal Cell Carcinoma with Tumor Thrombus of the Inferior Vena Cava. PloS one. 2014; 9:e109877.
21. Qiu M, Liu L, Chen L, Tan G, Liang Z, Wang K, Liu J, Chen H. microRNA-183 plays as oncogenes by increasing cell proliferation, migration and invasion via targeting protein phosphatase 2A in renal cancer cells. Biochemical and biophysical research communications. 2014; 452:163-169.
22. Yu G, Li H, Wang J, Gumireddy K, Li A, Yao W, Tang K, Xiao W, Hu J, Xiao H, Lang B, Ye Z, Huang Q, Xu H. miRNA-34a Suppresses Cell Proliferation and Metastasis by Targeting CD44 in Human Renal Carcinoma Cells. The Journal of urology. 2014; 192:1229-1237.
23. Lorenzen JM, Kaucsar T, Schauerte C, Schmitt R, Rong S, Hubner A, Scherf K, Fiedler J, Martino F, Kumarswamy R, Kolling M, Sorensen I, Hinz H, Heineke J, van Rooij E, Haller H, et al. MicroRNA-24 Antagonism Prevents Renal Ischemia Reperfusion Injury. Journal of the American Society of Nephrology. 2014.
24. Jenkins RH, Davies LC, Taylor PR, Akiyama H, Cumbes B, Beltrami C, Carrington CP, Phillips AO, Bowen T, Fraser DJ. miR-192 induces G2/M growth arrest in aristolochic acid nephropathy. The American journal of pathology. 2014; 184:996-1009.
25. Troyanskaya O, Cantor M, Sherlock G, Brown P, Hastie T, Tibshirani R, Botstein D, Altman RB. Missing value estimation methods for DNA microarrays. Bioinformatics (Oxford, England). 2001; 17:520-525.
26. Wettenhall JM, Smyth GK. limmaGUI: a graphical user interface for linear modeling of microarray data. Bioinformatics (Oxford, England). 2004; 20:3705-3706.
27. Maragkakis M, Alexiou P, Papadopoulos GL, Reczko M, Dalamagas T, Giannopoulos G, Goumas G, Koukis E, Kourtis K, Simossis VA, Sethupathy P, Vergoulis T, Koziris N, Sellis T, Tsanakas P, Hatzigeorgiou AG. Accurate microRNA target prediction correlates with protein repression levels. BMC bioinformatics. 2009; 10:295.
28. Betel D, Koppal A, Agius P, Sander C, Leslie C. Comprehensive modeling of microRNA targets predicts functional non-conserved and non-canonical sites. Genome biology. 2010; 11:R90.
29. Krek A, Grun D, Poy MN, Wolf R, Rosenberg L, Epstein EJ, MacMenamin P, da Piedade I, Gunsalus KC, Stoffel M, Rajewsky N. Combinatorial microRNA target predictions. Nature genetics. 2005; 37:495-500.
30. Miranda KC, Huynh T, Tay Y, Ang YS, Tam WL, Thomson AM, Lim B, Rigoutsos I. A pattern-based method for the identification of MicroRNA binding sites and their corresponding heteroduplexes. Cell. 2006; 126:1203-1217.
31. Rehmsmeier M, Steffen P, Hochsmann M, Giegerich R. Fast and effective prediction of microRNA/target duplexes. RNA (New York, NY). 2004; 10:1507-1517.
32. Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005; 120:15-20.
33. Kertesz M, Iovino N, Unnerstall U, Gaul U, Segal E. The role of site accessibility in microRNA target recognition. Nature genetics. 2007; 39:1278-1284.
34. Wang X. miRDB: a microRNA target prediction and functional annotation database with a wiki interface. RNA (New York, NY). 2008; 14:1012-1017.
35. Bandyopadhyay S, Mitra R. TargetMiner: microRNA target prediction with systematic identification of tissue-specific negative examples. Bioinformatics (Oxford, England). 2009; 25:2625-2631.
36. Enright AJ, John B, Gaul U, Tuschl T, Sander C, Marks DS. MicroRNA targets in Drosophila. Genome biology. 2003; 5:R1.
37. Ye J, Fang L, Zheng H, Zhang Y, Chen J, Zhang Z, Wang J, Li S, Li R, Bolund L, Wang J. WEGO: a web tool for plotting GO annotations. Nucleic acids research. 2006; 34:W293-297.
38. Cui X, Witalison EE, Chumanevich AP, Chumanevich AA, Poudyal D, Subramanian V, Schetter AJ, Harris CC, Thompson PR, Hofseth LJ. The induction of microRNA-16 in colon cancer cells by protein arginine deiminase inhibition causes a p53-dependent cell cycle arrest. PloS one. 2013; 8:e53791.
39. He X, Dong Y, Wu CW, Zhao Z, Ng SS, Chan FK, Sung JJ, Yu J. MicroRNA-218 inhibits cell cycle progression and promotes apoptosis in colon cancer by downregulating BMI1 polycomb ring finger oncogene. Molecular medicine (Cambridge, Mass). 2012; 18:1491-1498.
40. Oh J, Kim JW, Lee BE, Jang MJ, Chong SY, Park PW, Hwang SG, Oh D, Kim NK. Polymorphisms of the pri-miR-34b/c promoter and TP53 codon 72 are associated with risk of colorectal cancer. Oncology reports. 2014; 31:995-1002.
41. Pichiorri F, Suh SS, Rocci A, De Luca L, Taccioli C, Santhanam R, Zhou W, Benson DM, Jr., Hofmainster C, Alder H, Garofalo M, Di Leva G, Volinia S, Lin HJ, Perrotti D, Kuehl M, et al. Downregulation of p53-inducible microRNAs 192, 194, and 215 impairs the p53/MDM2 autoregulatory loop in multiple myeloma development. Cancer cell. 2010; 18:367-381.
42. Boni V, Bitarte N, Cristobal I, Zarate R, Rodriguez J, Maiello E, Garcia-Foncillas J, Bandres E. miR-192/miR-215 influence 5-fluorouracil resistance through cell cycle-mediated mechanisms complementary to its post-transcriptional thymidilate synthase regulation. Molecular cancer therapeutics. 2010; 9:2265-2275.
43. Georges SA, Biery MC, Kim SY, Schelter JM, Guo J, Chang AN, Jackson AL, Carleton MO, Linsley PS, Cleary MA, Chau BN. Coordinated regulation of cell cycle transcripts by p53-Inducible microRNAs, miR-192 and miR-215. Cancer Res. 2008; 68:10105-10112.
44. Zhang Q, He XJ, Ma LP, Li N, Yang J, Cheng YX, Cui H. [Expression and significance of microRNAs in the p53 pathway in ovarian cancer cells and serous ovarian cancer tissues]. [Article in Chinese]. Zhonghua zhong liu za zhi. 2011; 33:885-890.
45. Chiang Y, Song Y, Wang Z, Liu Z, Gao P, Liang J, Zhu J, Xing C, Xu H. microRNA-192, -194 and -215 are frequently downregulated in colorectal cancer. Experimental and therapeutic medicine. 2012; 3:560-566.
46. Khella HW, Bakhet M, Allo G, Jewett MA, Girgis AH, Latif A, Girgis H, Von Both I, Bjarnason GA, Yousef GM. miR-192, miR-194 and miR-215: a convergent microRNA network suppressing tumor progression in renal cell carcinoma. Carcinogenesis. 2013; 34:2231-2239.
47. Braun CJ, Zhang X, Savelyeva I, Wolff S, Moll UM, Schepeler T, Orntoft TF, Andersen CL, Dobbelstein M. p53-Responsive micrornas 192 and 215 are capable of inducing cell cycle arrest. Cancer Res. 2008; 68:10094-10104.
48. Wang JF, Zha YF, Li HW, Wang F, Bian Q, Lai XL, Yu G. Screening plasma miRNAs as biomarkers for renal ischemia-reperfusion injury in rats. Medical science monitor. 2014; 20:283-289.
49. Wang Y, Huang JW, Castella M, Huntsman DG, Taniguchi T. p53 is positively regulated by miR-542-3p. Cancer Res. 2014; 74:3218-3227.
50. Yoon S, Choi YC, Lee S, Jeong Y, Yoon J, Baek K. Induction of growth arrest by miR-542-3p that targets survivin. FEBS letters. 2010; 584:4048-4052.
51. Fragiadaki M, Witherden AS, Kaneko T, Sonnylal S, Pusey CD, Bou-Gharios G, Mason RM. Interstitial fibrosis is associated with increased COL1A2 transcription in AA-injured renal tubular epithelial cells in vivo. Matrix biology. 2011; 30:396-403.
52. Novitskaya T, McDermott L, Zhang KX, Chiba T, Paueksakon P, Hukriede NA, de Caestecker MP. A PTBA small molecule enhances recovery and reduces postinjury fibrosis after aristolochic acid-induced kidney injury. American journal of physiology Renal physiology. 2014; 306:F496-504.
53. Tian Y, Yang Y, Gao L, Zhao H, Peng X, Zhang Z, Wu G. Expression of histone deacetylase-1 and p300 in aristolochic acid nephropathy models. Toxicology mechanisms and methods. 2014; 24:377-384.
54. Weng Z, Wang D, Zhao W, Song M, You F, Yang L, Chen L. microRNA-450a targets DNA methyltransferase 3a in hepatocellular carcinoma. Experimental and therapeutic medicine. 2011; 2:951-955.
55. Thomas M, Lange-Grunweller K, Weirauch U, Gutsch D, Aigner A, Grunweller A, Hartmann RK. The proto-oncogene Pim-1 is a target of miR-33a. Oncogene. 2012; 31:918-928.
56. Ibrahim AF, Weirauch U, Thomas M, Grunweller A, Hartmann RK, Aigner A. MicroRNA replacement therapy for miR-145 and miR-33a is efficacious in a model of colon carcinoma. Cancer Res. 2011; 71:5214-5224.
57. Zhou Y, Huang Z, Wu S, Zang X, Liu M, Shi J. miR-33a is up-regulated in chemoresistant osteosarcoma and promotes osteosarcoma cell resistance to cisplatin by down-regulating TWIST. Journal of experimental & clinical cancer research. 2014; 33:12.
58. Davalos A, Goedeke L, Smibert P, Ramirez CM, Warrier NP, Andreo U, Cirera-Salinas D, Rayner K, Suresh U, Pastor-Pareja JC, Esplugues E, Fisher EA, Penalva LO, Moore KJ, Suarez Y, Lai EC, et al. miR-33a/b contribute to the regulation of fatty acid metabolism and insulin signaling. Proceedings of the National Academy of Sciences of the United States of America. 2011; 108:9232-9237.
59. Ueno K, Hirata H, Shahryari V, Chen Y, Zaman MS, Singh K, Tabatabai ZL, Hinoda Y, Dahiya R. Tumour suppressor microRNA-584 directly targets oncogene Rock-1 and decreases invasion ability in human clear cell renal cell carcinoma. British journal of cancer. 2011; 104:308-315.
60. Fils-Aime N, Dai M, Guo J, El-Mousawi M, Kahramangil B, Neel JC, Lebrun JJ. MicroRNA-584 and the protein phosphatase and actin regulator 1 (PHACTR1), a new signaling route through which transforming growth factor-beta Mediates the migration and actin dynamics of breast cancer cells. The Journal of biological chemistry. 2013; 288:11807-11823.
61. Ouhara K, Savitri IJ, Fujita T, Kittaka M, Kajiya M, Iwata T, Miyagawa T, Yamakawa M, Shiba H, Kurihara H. miR-584 expressed in human gingival epithelial cells is induced by Porphyromonas gingivalis stimulation and regulates interleukin-8 production via lactoferrin receptor. Journal of periodontology. 2014; 85:e198-204.
62. Aydin S, Dekairelle AF, Ambroise J, Durant JF, Heusterspreute M, Guiot Y, Cosyns JP, Gala JL. Unambiguous detection of multiple TP53 gene mutations in AAN-associated urothelial cancer in Belgium using laser capture microdissection. PloS one. 2014; 9:e106301.
63. Schmeiser HH, Nortier JL, Singh R, da Costa GG, Sennesael J, Cassuto-Viguier E, Ambrosetti D, Rorive S, Pozdzik A, Phillips DH, Stiborova M, Arlt VM. Exceptionally long-term persistence of DNA adducts formed by carcinogenic aristolochic acid I in renal tissue from patients with aristolochic acid nephropathy. International journal of cancer. 2014; 135:502-507.
64. Hollstein M, Moriya M, Grollman AP, Olivier M. Analysis of TP53 mutation spectra reveals the fingerprint of the potent environmental carcinogen, aristolochic acid. Mutation research. 2013; 753:41-49.
65. Rui HL, Wang YY, Cheng H, Chen YP. JNK-dependent AP-1 activation is required for aristolochic acid-induced TGF-beta1 synthesis in human renal proximal epithelial cells. American journal of physiology Renal physiology. 2012; 302:F1569-1575.
66. Li J, Zhang Z, Wang D, Wang Y, Li Y, Wu G. TGF-beta 1/Smads signaling stimulates renal interstitial fibrosis in experimental AAN. Journal of receptor and signal transduction research. 2009; 29:280-285.
67. Abouzeid H, Munier FL, Thonney F, Schorderet DF. Ten novel RB1 gene mutations in patients with retinoblastoma. Molecular vision. 2007; 13:1740-1745.
68. Storcelova M, Vician M, Reis R, Zeman M, Herichova I. Expression of cell cycle regulatory factors hus1, gadd45a, rb1, cdkn2a and mre11a correlates with expression of clock gene per2 in human colorectal carcinoma tissue. Molecular biology reports. 2013; 40:6351-6361.
69. Chang HR, Lian JD, Lo CW, Chang YC, Yang MY, Wang CJ. Induction of urothelial proliferation in rats by aristolochic acid through cell cycle progression via activation of cyclin D1/cdk4 and cyclin E/cdk2. Food and chemical toxicology. 2006; 44:28-35.
70. Arlt VM, Zuo J, Trenz K, Roufosse CA, Lord GM, Nortier JL, Schmeiser HH, Hollstein M, Phillips DH. Gene expression changes induced by the human carcinogen aristolochic acid I in renal and hepatic tissue of mice. International journal of cancer. 2011; 128:21-32.

