INTRODUCTION
Breast cancer is the most common tumor and the second leading cause of cancer deaths in women with over 1,400,000 new cases diagnosed every year worldwide. From the clinical point of view, breast cancer is characterized by wide heterogeneity. Gene expression profiling, as well as immuno-histochemical analysis of estrogen receptor (ER) α, progesterone receptor (PR) and human epidermal growth factor receptor 2 (HER2) can classify human breast cancers in four major histopatological/therapeutical subtypes: luminal A, luminal B, HER2 and basal-like [1, 2]. Each category differs in terms of prognosis and response to therapy. The ER positive group (luminal A and B) is amenable to hormone therapy, with several genomic tests to support in predicting outcomes [3]. HER2 group includes patients responsive to trastuzumab therapy, which has led to a great clinical success [3]. Basal cancers, also known as triple-negative breast cancers, are characterized by the lack of expression ER, PR and HER2 [3]. This phenotype makes basal tumors difficult to treat, more aggressive and with poor prognosis compared to the other subtypes [4].
During the last few years, new evidences estimated that approximately 95% of the human genome transcripts are non-coding RNAs (ncRNAs) [5, 6, 7]. Long non-coding RNAs (lncRNAs) are a subtype of ncRNAs molecules longer than 200nt involved in several biological and pathological processes, such as differentiation [8], immune response [9], metabolism [10] and cancer development and progression [11, 12, 13]. Interestingly, many cancer specific lncRNAs have been identified as biomarkers for metastasis or prognosis, for example metastasis associated long antisense transcript-1 (MALAT-1) in lung adenocarcinoma [14], HOX transcript antisense RNA (HOTAIR) in pancreatic and breast cancer [15, 11] and colon cancer associated transcripts (CCAT2) [16].
Recently, a new class of lncRNAs has been characterized, called transcribed-ultraconserved regions (T-UCRs). T-UCRs are encoded by a subset of ultraconserved regions (UCRs) in the DNA, which are absolutely conserved between orthologous loci of the human, rat, and mouse genomes [17, 18]. Interestingly, the expression of several T-UCRs is altered in tumorigenesis [18, 22] and it occurs in different ways. Frequently, fragile sites and cancer-associated genomic regions (CAGRs) contain T-UCRs, thus affecting their transcription when rearranged [18]. Furthermore, interactions with miRNAs have been reported to regulate T-UCRs levels in chronic lymphocytic leukemia (CLL) [18] and in neuroblastoma [19], whereas in prostate cancer and in other cancer cell lines (breast, colorectal, lung, lymphoma and leukemia) methylation of CpG islands in the promoter has been found involved in T-UCRs silencing [20, 21]. Finally, Ferdin et al. [23] discovered that a subset of T-UCRs (uc.63, uc.73, uc.106, uc.134 and uc.475) is regulated by hypoxia, a typical feature of tumor aggressiveness, and showed that uc.475 has a key role in supporting cancer cell proliferation in low-oxygen conditions [23]. Taken together, all these evidences underline the importance of T-UCRs in cancer biology and suggest the relevance of T-UCRs-expression profiles as useful tools to differentiate human cancer types and correlate with diagnosis and prognosis.
Here, we characterized a specific hypoxia-induced T-UCRs, named uc.63, in breast cancer. The data available on this T-UCRs reveal high levels of the transcript in colorectal cancer but there are no evidences about its biological role and prognostic significance in tumors. We found that uc.63 overexpression is correlated with poor prognosis in a luminal A subgroup of breast cancer patients and we discovered a key role in controlling cancer cells survival.
RESULTS
Long non-coding RNA uc.63 overexpression is associated with poor prognosis in breast cancer
uc.63 ultraconserved region (278bp) is localized on the chromosome 2p15 (chr2:61752501-61752778, GRCh37/hg19), in the third intron of XPO1 gene (Exportin-1, CRM1) (Figure 1A). The sequence is conserved in a broad spectrum of species and in mouse is localized in the fourth intron of Xpo1 orthologous gene (Supplementary Table S1)
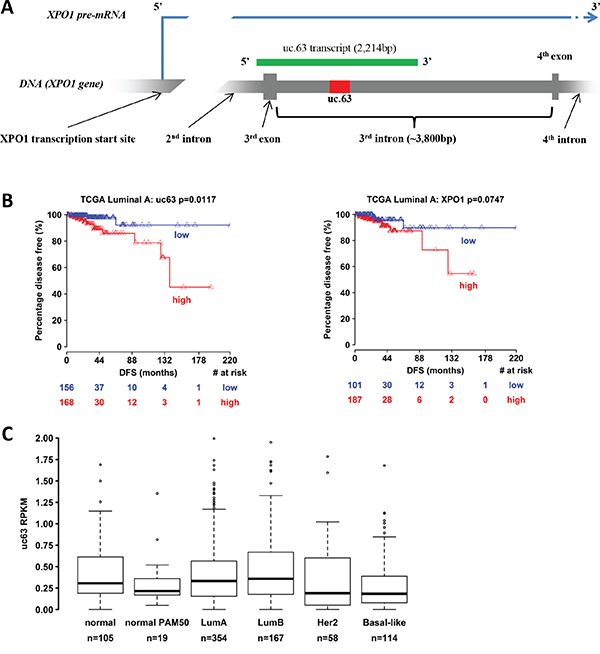
Figure 1: Bionformatic analysis of uc.63 expression in breast cancer patients. A. Scheme of uc.63 transcript and locus in chromosome 2 (grey: XPO1 gene / red: uc.63 ultraconserved sequence / green: uc.63 transcript). B. Disease-free survival information of breast cancer patients were downloaded from TCGA portal (http://tcga-data.nci.nih.gov/) [3, 87]. Kaplan-Meier method was used to generate percentage disease-free curves. C. Expression analysis of uc.63 transcript in PAM50 breast cancer subgroups. PAM50 gene signature relies on 50 discriminatory genes to segregate tumors into luminal A, luminal B, HER2–enriched and basal-like [88]. Samples labelled with “normal PAM50” are tumor subtype with no corresponding clinical-pathologic category (normal-like). Samples categorized as “normal” are normal breast tissue.
In MCF-7 breast cancer cell line, the transcription of this region leads to a longer transcript, approximately of 2,200nt [23], thus overlapping the third exon of XPO1 gene (Figure 1A). Furthermore, cellular fractionation experiment revealed a predominant nuclear localization of uc.63 [23]. Overexpression of uc.63 transcript has been found in patients with colorectal cancer and its upregulation is induced by hypoxic conditions [23] in vitro. Despite these data, the clinical relevance of uc.63 is still unknown.
In order to investigate the prognostic significance of uc.63 overexpression in breast cancer, we performed a bionformatic analysis by using TCGA (The Cancer Genome Atlas) portal [3, 87], which contains expression profiles data of over 2,000 breast cancer specimens. We found that the increased expression of uc.63 is associated with reduced disease-free survival in luminal A subtype of breast cancer patients (Figure 1B, left panel). Interestingly, XPO1 overexpression is not correlated with the free survival, thus demonstrating that high levels of XPO1 are not linked to relapse in this group of tumors (Figure 1B, right panel). Despite these data, analysis of uc.63 expression in breast cancer subgroups of PAM50 gene signature does not reveal any significant upregulation of uc.63 transcript both in luminal A and in all of other subtypes (Figure 1C), suggesting that the prognostic value of uc.63 is limited to a cohort of patients with more aggressive luminal A tumors which develop a relapse of the disease.
Taken together, these data show a correlation between uc.63 upregulation and aggressiveness in luminal A breast cancers, identifying a potentially useful molecular signature for the prognosis of this neoplasia.
Analysis of uc.63 transcript
In order to analyze in vitro the uc.63 transcript, we designed specific primers targeting uc.63 ultraconserved sequence. First, we evaluated uc.63 expression in a panel of breast cancer cell lines by RT-qPCR, using as a reference sample HMEC cells (Human Mammary Epithelial Cells), which resemble the normal breast epithelium (Figure 2A). The analyzed cell lines showed high variability in terms of uc.63 expression. For the sake of simplicity, we discriminated cell lines with low (MCF-7, T-47D, MDA MB 231, MDA MB 468, BT-20, BT-549) and high (MDA MB 453, ZR-75-1, BT-474, SUM 149 PT, HCC1937, HCC1954) uc.63 levels by calculating the median of all RQ values in tumor cells (Figure 2A). For following analysis, we used total RNA from MDA MB 453 breast cancer cell line derived from metastatic breast carcinoma and showing high expression of uc.63 (Figure 2A).
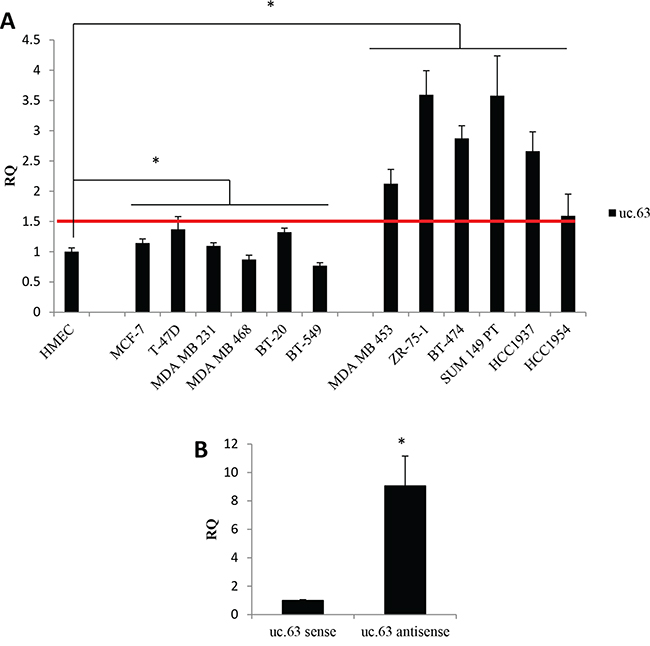
Figure 2: uc.63 expression in human breast cancer lines. A. Expression analysis of uc.63 in breast cancer cell lines. 500,000 cells were plated in complete medium and uc.63 level was evaluated by RT-qPCR after 48h. HMECs were used as reference sample and TBP was used as endogenous control. Median of all RQ values in tumor cells (RQ=1.50) is represented by a red line. Cell lines with low (left) and high (right) uc.63 levels were separated. *p < 0.01 vs HMEC. B. Strand-specific RT-qPCR. Total RNA isolated from MDA MB 453 cells was used to perform RT-qPCR with strand-specific primers complementary to uc.63 antisense or uc.63 sense transcripts. TBP was used as endogenous control. uc.63 sense was used as reference sample. *p < 0.01 vs uc.63 sense.
Given that uc.63 is localized inside XPO1 gene, we asked whether uc.63 is transcribed in the same orientation of host gene in these cells. To address this point, we carried out a strand-specific RT-qPCR making two reverse transcription reaction mix, each one containing the primer complementary to the different forms of the transcript. We found that antisense transcript is the predominant form of uc.63 expressed in MDA MB 453, being sense transcript poorly detectable (Figure 2B, Supplementary Figure S1A). As shown in Supplementary Figure S1B, uc.63 antisense has the same sequence of XPO1 primary transcript, completely matching the region between the end of the second intron and a part of the third intron and containing the third exon (Figure 1A). Thus, we clearly demonstrated that uc.63 RNA is transcribed in the same orientation of XPO1 mRNA. Our results are in agreement with previously published data [23], which reported the same transcription orientation in MCF-7 breast cancer cell line.
Although uc.63 promoter has not been characterized yet, these data demonstrate that uc.63 transcript has the same orientation of host gene’s mRNA. Furthermore, the presence in the putative uc.63 promoter region at 5' of XPO1 gene [23] of hypoxia-inducible factor (HIF) candidate binding sites, which may drive uc.63 expression in hypoxic conditions, further confirm the same transcription orientation.
Biological effects of uc.63
Given that there are no evidences about the biological role of uc.63 overexpression in breast cancer, we used high-expressing uc.63 MDA MB 453 cell line and a siRNA-based approach by designing two different specific siRNAs targeting uc.63 ultraconserved sequence, in order to investigate the phenotypic effect of uc.63 knockdown (KD). We found that uc.63 downregulation leads to cell death in MDA MB 453 cells (Figure 3A, 3C), associated with an increase in G0/G1 cells and a reduction in G2/M events (Figure 3B). Of note, the same results were obtained by using uc.63 siRNAs alone or in combination. uc.63 siRNAs also cause a slight but significant decrease in XPO1 mRNA level, probably by targeting the primary transcript (the third intron) of host gene (Figure 3C). However, uc.63 siRNAs do not affect XPO1 protein level (Figure 3D, Supplementary Figure S2), thus excluding the possibility that XPO1 downregulation leads to the phenotypic effect. Finally, we asked whether apoptosis is involved in inducing cell death of MDA MB 453 cells in vitro. Thus, we analyzed the activation status of apoptotic markers PARP-1 and Caspase-3. Western blot analysis revealed both PARP-1 and Caspase-3 activated forms (cleaved) after uc.63 KD, thus confirming our hypothesis (Figure 3E). Taken together, these findings show that uc.63 plays an important role in controlling survival in breast cancer cells.
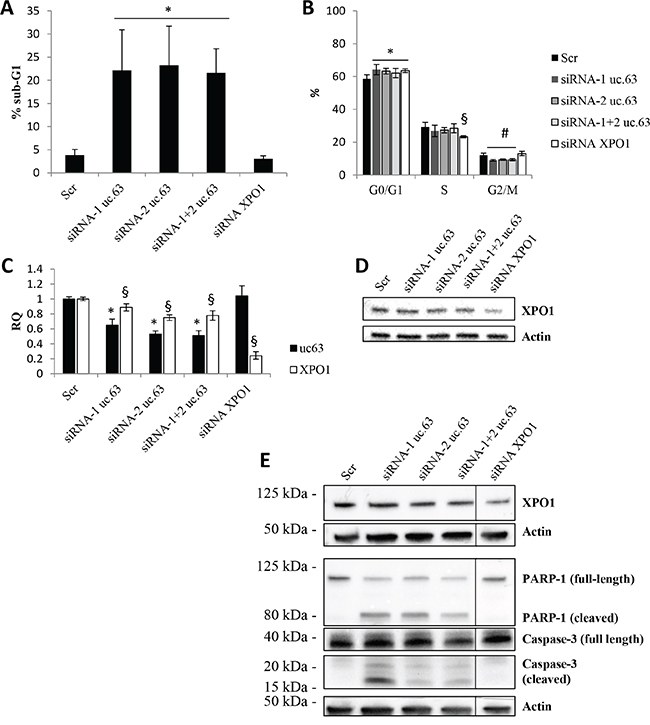
Figure 3: uc.63 knockdown induces apoptosis in MDA MB 453 cell line. A. sub-G1 events were quantified by using flow cytometry analysis after propidium iodide staining of MDA MB 453 cell line. 500,000 cells were plated and then silenced with siRNA (20nM) targeting uc.63 or XPO1 for 48h. *p < 0.01 vs Scr. B. Cell cycle analysis following uc.63 or XPO1 knockdown. Analysis was carried out by using ModFit software. *p < 0.05 vs G0/G1 Scr; §p < 0.01 vs S Scr; #p < 0.05 vs G2/M Scr. C. RT-qPCR for uc.63 and XPO1 after uc.63 or XPO1 knockdown. TBP was used as endogenous control. *p < 0.01 vs Scr, §p < 0.01 vs Scr. D. XPO1 immunoblot analysis following uc.63 or XPO1 knockdown. β-actin was used as endogenous control. E. Immunoblot analysis of apoptotic markers PARP-1 and caspase-3. β-actin was used as endogenous control.
XPO1 mRNA and uc.63 are two independent transcripts
In order to further demonstrate that XPO1 is not involved in the biological effects induced by uc.63 downregulation, we carried out XPO1 KD by using a pool of four specific siRNAs. As expected, XPO1 downregulation does not affect survival of MDA MB 453 cells (Figure 3A, 3C, 3D) and we only detected a reduction in S phase associated with increased G1/G0 events (Figure 3B). In agreement, we did not find any activation of apoptotic pathway, as indicated by PARP-1 and Caspase-3 immunoblot analysis (Figure 3E). Altogether, this functional analysis strongly suggests that uc.63 and XPO1 mRNA are two independent transcripts and only uc.63 is able to modulate apoptosis.
DISCUSSION
Breast cancer is a heterogeneous disease with different histopathological, genetic and genomic variations, and clinical outcomes, which make difficult to define therapies and prognostic factors. Genetic and epigenetic changes, as well as aberrant interactions with tumor microenvironment, drive a multistep process characterized by progressive deregulation of proliferation, survival, differentiation and metabolism, finally giving rise to aggressive and metastatic breast tumor [26, 27]. Despite novel therapies, ~30% of treated patients later relapse and over 450,000 die yearly [25]. For all these reasons, gene-expression profiles of breast carcinomas are useful tools in order to identify prognostic markers and predict response to treatments.
A relevant player in breast cancer, as well as in cancer in general, is the entire p53 family. The p53 gene family of transcription factors includes three very similar genes that codify for the p53, p73 and p63 proteins [28, 29, 30, 31]. p53 shows a very complex transcription activation program ranging from metabolism [32, 33, 34, 35], mitochondria and ROS regulation [36, 37], DNA damage repair [38, 39, 40, 41], autophagy [42, 43], stemness and lineage determination [44, 45]. At the molecular level, this complexity can be studied from different aspects; indeed, detailed analysis are under way on its splicing isoforms [46, 47], its connection and regulation to miRNAs [48, 49, 50, 51] as well as its stability and degradation [24, 52, 53, 54, 55, 56, 57, 58]. Furthermore, in keeping with the progress in understanding p53 function, there is also a strong effort in investigating innovative therapeutic cues [59, 60, 61, 62, 63]. Of note, the other members of the family, p63 and p73, which were discovered only ten years ago, have an evident complex interplay with p53 itself [64, 65, 66, 67, 68]. In the case of p63, its function is crucial for skin development and homeostasis, as well as for cancer [69, 70, 66, 71, 72, 73, 74], whereas p73 is a critical player in brain development and homeostasis, as well as in tumors [75, 33, 76, 77, 78, 79].
In the last few years, ncRNAs are emerging as new actors in breast cancer biology [80]. In this regard, miRNAs play a key role in proliferation, metastasis and drug resistance in breast tumors [81]. For example, miR-34 family members, which are directly regulated by p53, control cell cycle and apoptosis by modulating the levels of proteins such as CDK4, CDK6, cyclin D1, cyclin E2 and BCL-2 [81]. Furthermore, miR-125a and miR-125b are able to suppress HER2 expression, thus decreasing both growth and invasiveness [82], while miR-200 family members are directly involved in regulating epithelial-mesenchymal transition [83]. Lastly, miR-221 and miR-222 contribute to tamoxifen resistance, downregulating ER-α level [84]. Interestingly, among ncRNAs, lncRNAs have recently appeared as relevant drivers of breast cancer. The long non-coding transcript HOTAIR has been described to remodel chromatin through polycomb repressive complex 2 (PRC2), thus increasing cancer invasiveness in vitro and in vivo, and its overexpression is an independent predictor of overall survival and progression-free survival [11]. On the other hand, GAS5 lncRNA acts as an oncosuppressor, being downregulated in breast tumors and leading to apoptosis and block of proliferation in vitro [85]. Despite these data, the new class of lncRNAs T-UCRs is not characterized in breast cancer and our knowledge about their role in breast tumor biology is very limited.
Dysregulation of T-UCRs expression is emerging as a common feature of human cancers. Here, we discovered that aberrant expression of uc.63 in breast cancer cells is important in controlling survival. In fact, uc.63 KD leads to apoptosis in vitro and this effect is specific for the T-UCR. Although uc.63 is localized inside the XPO1 gene and is transcribed in the same transcriptional orientation, exportin-1 KD does not recapitulate the phenotypic effect of uc.63 downregulation, suggesting that uc.63 and XPO1 mRNA are two independent transcripts with two independent promoters. However, even though our functional analysis strongly suggests an independent transcription, further studies will be needed in order to better characterize uc.63 RNA and its regulation. Of note, hypoxia has been already demonstrated to transcriptionally induce uc.63 [23] and this feature strongly supports the link between cancer aggressiveness and its expression.
As for uc.73 in colorectal cancer cells [18], we found that uc.63 is able to modulate apoptosis in breast cancer cells. This data suggests that uc.63 acts as an oncogene and its overexpression is important in promoting cell survival and tumor growth. At the moment, we have no evidences about the proper mechanism by which uc.63 controls survival in tumor cells and further analysis are necessary to deeply understand the cellular pathways involved in this process.
Several evidences demonstrated that, compared to normal counterpart, neoplastic cells show a unique expression profile of T-UCRs, suggesting a significant role of T-UCRs in the malignant process [18]. Variations in ultraconserved non-coding transcript levels offer a new strategy for tumor diagnosis and prognosis. In our study we found that, in the specific context of luminal A breast cancer tumors, uc.63 overexpression is associated with reduced disease-free survival, thus correlating this T-UCR with aggressiveness. Of note, XPO1 overexpression is not correlated with relapse in luminal A subtype, suggesting that exportin-1 does not affect disease-free survival in this particular group of tumors. Moreover, this data further confirms the independence in terms of expression between XPO1 and uc.63. Therefore, we propose uc.63 expression as a potential tool useful for prognosis in breast cancer patients
In summary, our study revealed an important role of uc.63 in breast cancer and could potentially offer the perspective of identification of T-UCRs' signatures associated with prognosis of the disease.
MATERIALS AND METHODS
Cell cultures and growth conditions
MDA MB 453, MDA MB 231, MDA MB 468, BT-20, BT-549, BT-474, MCF-7 and T-47D cells were cultured in Dulbecco's Modified Eagle's Medium DMEM (LONZA) supplemented with 10% FBS (Gibco) and Penicillin-Streptomycin (Gibco), respectively 100 U/mL and 100 μg/mL. SUM 149 PT were cultured in HAMs F12 medium supplemented with 5% FBS, 2mM glutamine (Gibco), 5μg/mL insulin, 10mM HEPES, 1μg/mL hydrocortisone and Penicillin-Streptomycin. HCC 1937 and HCC 1954 were cultured in RPMI medium supplemented with 10% FBS, 2mM glutamine and Penicillin-Streptomycin. ZR-75-1 were cultured in RPMI medium supplemented with 10% FBS, 1mM sodium pyruvate, 10mM HEPES, 2mM glutamine and Penicillin-Streptomycin. HMEC were obtained from Gibco and were cultured in HuMEC medium consists of HuMEC Basal Serum-Free Medium (Gibco, cat. 12753018) supplemented with HuMEC Supplement Kit (Gibco, cat. 12755013). The HuMEC Supplement Kit includes 5mL of a supplement mix containing epidermal growth factor, hydrocortisone, isoproterenol, transferrin, and insulin, and 25mg of bovine pituitary extract.
siRNA transfection
MDA MB 453 were transfected using Lipofectamine RNAiMAX transfection reagent (Invitrogen) by following manufacturer's protocol. 500,000 cells were plated one day before transfection. uc.63 and XPO1 were silenced using 20nM siRNA for 48h. uc.63 siRNAs target the following sequences: uc.63 siRNA-1, GACATTACTAATGTTTAAGTTGA; uc.63 siRNA-2, TTCTTCAATTTACATAAATTACA. Both siRNAs were provided by Qiagen. XPO1 knockdown was performed by using a pool of four different siRNA (Gene Solution siRNA XPO1, Qiagen, cat. 1027416 - 2991182).
Primers
uc.63 ultraconserved sequence was downloaded from Bejerano et al. [17]. This sequence was used both for primers and siRNAs design. For the sake of simplicity, we used the complementary sequence of uc.63: uc.63 forward primer, CAGTGTTTGCCTGTTTGCTTGC; uc.63 reverse primer, CCTGTTGCTTTCTTTCTGTTCCTC. XPO1 mRNA was detected using the following primer: XPO1 forward primer, CTCATTGTTTCCCAGCATTCCTTG; XPO1 reverse primer, CCCGTATCTGCGACATTCCTCATAG. TATA-box binding protein (TBP) was used as reference gene in all RT-qPCR analysis: TBP forward primer, TCAAACCCAGAATTGTTCTCCTTAT; TBP reverse primer, CCTGAATCCCTTTAGAATAGGGTAGA.
RNA extraction and reverse transcription (RT)
Total RNA was extracted from cultured cells using RNAeasy Mini Kit by following manufacturer’s protocol (Qiagen). RNA concentrations were measured with spectrophotometer NanoDrop ND-1000 instrument (NanoDrop Tachnologies, Termo Scientific). Reverse transcription of RNA to total cDNA was performed on 1μg of RNA using GoTaq Reverse Transcription System (Promega) by following manufacturer's protocol, in a final reaction of 40μl (1X Reaction Buffer 5X, 5mM MgCl2, 0.5mM dNTPs, 1μg random primers, 320U reverse transcriptase, 20U ribonuclease inhibitor).
Strand specific RT was made in 20μL where 100ng of RNA were transcribed to strand specific cDNA (1X Reaction Buffer 5X, 5mM MgCl2, 0.5mM dNTPs, 0.5μM strand-specific primers, 160U reverse transcriptase, 20U ribonuclease inhibitor). Two reverse transcription reaction mix were made for strand-specific cDNA synthesis: one to detect uc.63 sense oriented transcript (using forward primer for uc.63 and reverse primer for TBP) and one for uc.63 antisense oriented transcript (using reverse primers both for uc.63 and TBP). Of note, we used only TBP reverse primer in both reaction mix because it can recognize TBP's mRNA. Before use in qPCR analysis, strand-specific cDNAs were diluted 5-times with nuclease-free water.
Quantitative polymerase chain reaction (qPCR)
Real time PCR was performed using GoTaq qPCR Master Mix (Promega) by following manufacturer's protocol, in a final volume of 25μL (1X qPCR Master Mix 2X, 0.4μM forward and reverse primer mix, 1mM MgCl2). 2-ΔΔCt method was used for relative quantifications. TBP was used as reference gene in all reactions.
Western blot
Cells lysis was performed in RIPA buffer (1% NP-40, 0.1% SDS, 150mM NaCl, 50mM Tris-HCl pH=7.5, 0.5% sodium deoxycholate) supplemented with protease inhibitors and 1mM dithiothreitol. Samples were homogenized by vortexing, kept on ice for 20 minutes and then clarified by centrifugation at 13,000rpm for 10 minutes at 4°C. 20-30μg of total proteins were separated by SDS-PAGE and then transferred onto PVDF membrane (GE Healthcare). Immunoblotting was performed using standard protocols with the following primary antibodies: rabbit anti-XPO1 1:200 (Santa Cruz Biotech., cat. sc-5595), mouse anti-PARP-1 (C-2-10) 1:1000 (Enzo Lifescience, cat. BML-SA250), rabbit anti-Caspase-3 (8G10) 1:500 (Cell Signaling, cat. 9665), mouse anti-actin 1:50,000 (Sigma Aldrich, cat. A5441). Caspase-3 antibody detects endogenous levels of full-length (35KDa) and large fragment (17/19 KDa) of caspase-3 resulting from cleavage at aspartic acid 175. PARP-1 antibody detects intact PARP (~116KDa) and apoptosis-induced cleavage fragment (~80KDa).
Bionformatic analysis
We downloaded RNA-seq BAM files from UCSC Cancer Genomics Hub (CGHub, https://cghub.ucsc.edu/) for Breast invasive carcinoma (BRCA), respectively normal cases from TCGA (The Cancer Genome Atlas). TCGA BAM files were generated based on Mapsplice2 algorithm for alignment against the hg19 reference genome using default parameters. We quantified the expression of uc.63 as RPKM (reads per kilobase per million mapped reads) by extracting the number of reads overlapped with uc.63 and normalized by the total mappable reads of each sample as previously described [86]. We downloaded RNASeq (RPKM) Level3 data publicly available from the TCGA (http://tcga-data.nci.nih.gov/) for XPO1 in patients with BRCA. Patient survival information was retrieved from cbioPortal (http://www.cbioportal.org/). Patient PAM50 classification results were retrieved from the Associated Data Files of the Cancer Genome Atlas Research Network [87]. Data are available per request. Analyses were carried out in R statistical environment (version 3.0.1) (http:///www.r-project.org/). All tests were two-sided and considered statistical significant at the 0.05 level. We performed Cox regression analysis for associations between survival and uc.63 and XPO1 levels. For the Lumina A subgroup, the disease-free survival analysis yielded for uc.63 a hazard ratio of 3.49 (CI(95%)=(1. 17, 10.34), Wald test p-value =0.0243). We than used the log-rank test to find the point (cut-off) with the most significant (lowest p-value) split in high vs low uc.63 mRNA level groups. The Kaplan-Meier plots were generated for this cut-off (0.48). The numbers of patients at risk in low and high uc.63 groups at different time points are presented at the bottom of the graph. The same analysis revealed that XPO1 expression does not associate with survival in this group (hazard ratio= 1.58, CI(95%)=(0.45, 5.9), Wald test p-value = 0.475). We used same methods as for uc63 to generate Kaplan-Meier plots for XPO1 (cut-off=0.35). The Shapiro-Wilk test was applied and verified that uc.63 expression does not follow a normal distribution in each PAM50 subtype, and normal group. Accordingly, the nonparametric Kruskal-Wallis test was applied to assess the relationship between uc.63 expression and subtype. A box-and-whisker plot (Box plot represents first (lower bound) and third (upper bound) quartiles, whiskers represent 1.5 times the interquartile range) was used to visualize the data (log2(x+1)).
Flow cytometry assay
Propidium iodide (PI) staining of fixed cells was used to evaluate apoptosis and to analyze the distribution in the different cell cycle phases of the cell population. Cells in G1/G0 phases have a diploid DNA content, whereas cells in G2/M phases have a double DNA content. Cells in S phase have an intermediate DNA content. Apoptotic cells are detectable as G1 sub-population (sub-G1) with hypo-diploid DNA content caused by DNA fragmentation. Briefly, cells were harvested along with their medium, centrifuged at 1,200rpm for 10 minutes and washed in ice-cold PBS. Cells were centrifuged again at 1,200rpm for 10 minutes and then fixed in 500μL methanol-acetone 4:1 at 4°C for 30 minutes. After fixing, 2mL of PBS was added to cells. The suspension was centrifuged at 1,200rpm for 10 minutes and the pellet was then incubated in 50μL of 10kU/mL RNAse (Sigma Aldrich) at room temperature for 15 minutes. Finally, 200μL of 60μg/mL PI (Sigma Aldrich) was added to fixed cells, incubating the suspension for 20 minutes. Analysis of samples was carried out acquiring 10,000-15,000 events/sample on FACS Calibur (BD). ModFit software was used to analyze cell cycle phases.
Statistical analysis
Data are presented as means ± SD. Statistical analysis of the data was performed by Student’s t-test. p-values of ≤ 0.05 were considered statistically significant.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
GRANT SUPPORT
This work has been mainly supported by “Associazione Italiana per la Ricerca contro il Cancro” (AIRC) 2012 IG13387 (to E.C.) and partially supported by the Medical Research Council, UK and Istituto Dermopatico dell’Immacolata/Istituto di Ricovero e Cura a Carattere Scientifico (IDI-IRCCS).
Abbreviations
T-UCRs, transcribed-ultraconserved regions; lncRNAs, long non-coding RNAs; XPO1, exportin-1; TCGA The Cancer Genome Atlas; HMEC, human mammary epithelial cells; PARP-1, Poly [ADP-ribose] polymerase 1; siRNA, small interfering RNA.
REFERENCES
1. Perou CM, Jeffrey SS, van de Rijn M, Rees CA, Eisen MB, Ross DT, Pergamenschikov A, Williams CF, Zhu SX, Lee JC, Lashkari D, Shalon D, Brown PO, Botstein D. Distinctive gene expression patterns in human mammary epithelial cells and breast cancers. Proc Natl Acad Sci USA. 1999; 96:9212–17.
2. Perou CM, Sørlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Pergamenschikov A, Williams C, et al. Molecular portraits of human breast tumours. Nature. 2000; 406:747–52.
3. Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature. 2012; 490:61–70.
4. Holliday DL, Speirs V. Choosing the right cell line for breast cancer research. Breast Cancer Res. 2011; 13:215.
5. Dermitzakis ET, Reymond A, Antonarakis SE. Conserved non-genic sequences - an unexpected feature of mammalian genomes. Nat Rev Genet. 2005; 6:151–57.
6. Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, Stadler PF, Hertel J, Hackermüller J, Hofacker IL, Bell I, Cheung E, Drenkow J, et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science. 2007; 316:1484–88.
7. Mattick JS. The genetic signatures of noncoding RNAs. PLoS Genet. 2009; 5:e1000459.
8. Kretz M, Siprashvili Z, Chu C, Webster DE, Zehnder A, Qu K, Lee CS, Flockhart RJ, Groff AF, Chow J, Johnston D, Kim GE, Spitale RC, et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature. 2013; 493:231–35.
9. Wang P, Xue Y, Han Y, Lin L, Wu C, Xu S, Jiang Z, Xu J, Liu Q, Cao X. The STAT3-binding long noncoding RNA lnc-DC controls human dendritic cell differentiation. Science. 2014; 344:310–13.
10. Hung CL, Wang LY, Yu YL, Chen HW, Srivastava S, Petrovics G, Kung HJ. A long noncoding RNA connects c-Myc to tumor metabolism. Proc Natl Acad Sci USA. 2014; 111:18697–702.
11. Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, Wang Y, Brzoska P, Kong B, et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature. 2010; 464:1071–76.
12. Yang F, Huo XS, Yuan SX, Zhang L, Zhou WP, Wang F, Sun SH. Repression of the long noncoding RNA-LET by histone deacetylase 3 contributes to hypoxia-mediated metastasis. Mol Cell. 2013; 49:1083–96.
13. Xing Z, Lin A, Li C, Liang K, Wang S, Liu Y, Park PK, Qin L, Wei Y, Hawke DH, Hung MC, Lin C, Yang L. lncRNA directs cooperative epigenetic regulation downstream of chemokine signals. Cell. 2014; 159:1110–25.
14. Ji P, Diederichs S, Wang W, Böing S, Metzger R, Schneider PM, Tidow N, Brandt B, Buerger H, Bulk E, Thomas M, Berdel WE, Serve H, Müller-Tidow C. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene. 2003; 22:8031–41.
15. Kim K, Jutooru I, Chadalapaka G, Johnson G, Frank J, Burghardt R, Kim S, Safe S. HOTAIR is a negative prognostic factor and exhibits pro-oncogenic activity in pancreatic cancer. Oncogene. 2013; 32:1616–25.
16. Qiu M, Xu Y, Yang X, Wang J, Hu J, Xu L, Yin R. CCAT2 is a lung adenocarcinoma-specific long non-coding RNA and promotes invasion of non-small cell lung cancer. Tumour Biol. 2014; 35:5375–80.
17. Bejerano G, Pheasant M, Makunin I, Stephen S, Kent WJ, Mattick JS, Haussler D. Ultraconserved elements in the human genome. Science. 2004; 304:1321–25.
18. Calin GA, Liu CG, Ferracin M, Hyslop T, Spizzo R, Sevignani C, Fabbri M, Cimmino A, Lee EJ, Wojcik SE, Shimizu M, Tili E, Rossi S, et al. Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas. Cancer Cell. 2007; 12:215–29.
19. Scaruffi P, Stigliani S, Moretti S, Coco S, De Vecchi C, Valdora F, Garaventa A, Bonassi S, Tonini GP. Transcribed-Ultra Conserved Region expression is associated with outcome in high-risk neuroblastoma. BMC Cancer. 2009; 9:441.
20. Lujambio A, Portela A, Liz J, Melo SA, Rossi S, Spizzo R, Croce CM, Calin GA, Esteller M. CpG island hypermethylation-associated silencing of non-coding RNAs transcribed from ultraconserved regions in human cancer. Oncogene. 2010; 29:6390–401.
21. Hudson RS, Yi M, Volfovsky N, Prueitt RL, Esposito D, Volinia S, Liu CG, Schetter AJ, Van Roosbroeck K, Stephens RM, Calin GA, Croce CM, Ambs S. Transcription signatures encoded by ultraconserved genomic regions in human prostate cancer. Mol Cancer. 2013; 12:13.
22. Braconi C, Valeri N, Kogure T, Gasparini P, Huang N, Nuovo GJ, Terracciano L, Croce CM, Patel T. Expression and functional role of a transcribed noncoding RNA with an ultraconserved element in hepatocellular carcinoma. Proc Natl Acad Sci USA. 2011; 108:786–91.
23. Ferdin J, Nishida N, Wu X, Nicoloso MS, Shah MY, Devlin C, Ling H, Shimizu M, Kumar K, Cortez MA, Ferracin M, Bi Y, Yang D, et al. HINCUTs in cancer: hypoxia-induced noncoding ultraconserved transcripts. Cell Death Differ. 2013; 20:1675–87.
24. Deisenroth C, Thorner AR, Enomoto T, Perou CM, Zhang Y. Mitochondrial Hep27 is a c-Myb target gene that inhibits Mdm2 and stabilizes p53. Mol Cell Biol. 2010; 30:3981–93.
25. Siegel R, Naishadham D, Jemal A. Cancer statistics, 2012. CA Cancer J Clin. 2012; 62:10–29.
26. Weigelt B, Peterse JL, van ’t Veer LJ. Breast cancer metastasis: markers and models. Nat Rev Cancer. 2005; 5:591–602.
27. Morandi A, Chiarugi P. Metabolic implication of tumor: stroma crosstalk in breast cancer. J Mol Med (Berl). 2014; 92:117–26.
28. Tomasini R, Mak TW, Melino G. The impact of p53 and p73 on aneuploidy and cancer. Trends Cell Biol. 2008; 18:244–52.
29. Zambetti GP. Expanding the reach of the p53 tumor suppressor network. Cell Death Differ. 2014; 21:505–06.
30. Solomon H, Sharon M, Rotter V. Modulation of alternative splicing contributes to cancer development: focusing on p53 isoforms, p53β and p53γ. Cell Death Differ. 2014; 21:1347–49.
31. Soussi T, Wiman KG. TP53: an oncogene in disguise. Cell Death Differ. 2015; 22:1239–49.
32. Xu J, Wang J, Hu Y, Qian J, Xu B, Chen H, Zou W, Fang JY. Unequal prognostic potentials of p53 gain-of-function mutations in human cancers associate with drug-metabolizing activity. Cell Death Dis. 2014; 5:e1108.
33. Amelio I, Cutruzzolá F, Antonov A, Agostini M, Melino G. Serine and glycine metabolism in cancer. Trends Biochem Sci. 2014; 39:191–98.
34. Amelio I, Antonov AA, Catani MV, Massoud R, Bernassola F, Knight RA, Melino G, Rufini A. TAp73 promotes anabolism. Oncotarget. 2014; 5:12820–934. doi: 10.18632/oncotarget.2667.
35. Antonov A, Agostini M, Morello M, Minieri M, Melino G, Amelio I. Bioinformatics analysis of the serine and glycine pathway in cancer cells. Oncotarget. 2014; 5:11004–13. doi: 10.18632/oncotarget/2668.
36. Ci Y, Shi K, An J, Yang Y, Hui K, Wu P, Shi L, Xu C. ROS inhibit autophagy by downregulating ULK1 mediated by the phosphorylation of p53 in selenite-treated NB4 cells. Cell Death Dis. 2014; 5:e1542.
37. Evstafieva AG, Garaeva AA, Khutornenko AA, Klepikova AV, Logacheva MD, Penin AA, Novakovsky GE, Kovaleva IE, Chumakov PM. A sustained deficiency of mitochondrial respiratory complex III induces an apoptotic cell death through the p53-mediated inhibition of pro-survival activities of the activating transcription factor 4. Cell Death Dis. 2014; 5:e1511.
38. Nicolai S, Rossi A, Di Daniele N, Melino G, Annicchiarico-Petruzzelli M, Raschellà G. DNA repair and aging: the impact of the p53 family. Aging (Albany NY). 2015; 7:1050–65.
39. Nair BC, Krishnan SR, Sareddy GR, Mann M, Xu B, Natarajan M, Hasty P, Brann D, Tekmal RR, Vadlamudi RK. Proline, glutamic acid and leucine-rich protein-1 is essential for optimal p53-mediated DNA damage response. Cell Death Differ. 2014; 21:1409–18.
40. Phesse TJ, Myant KB, Cole AM, Ridgway RA, Pearson H, Muncan V, van den Brink GR, Vousden KH, Sears R, Vassilev LT, Clarke AR, Sansom OJ. Endogenous c-Myc is essential for p53-induced apoptosis in response to DNA damage in vivo. Cell Death Differ. 2014; 21:956–66.
41. Dashzeveg N, Taira N, Lu ZG, Kimura J, Yoshida K. Palmdelphin, a novel target of p53 with Ser46 phosphorylation, controls cell death in response to DNA damage. Cell Death Dis. 2014; 5:e1221.
42. Simon HU, Yousefi S, Schmid I, Friis R. ATG5 can regulate p53 expression and activation. Cell Death Dis. 2014; 5:e1339.
43. Garufi A, Pucci D, D’Orazi V, Cirone M, Bossi G, Avantaggiati ML, D’Orazi G. Degradation of mutant p53H175 protein by Zn(II) through autophagy. Cell Death Dis. 2014; 5:e1271.
44. Kim J, Nakasaki M, Todorova D, Lake B, Yuan CY, Jamora C, Xu Y. p53 Induces skin aging by depleting Blimp1+ sebaceous gland cells. Cell Death Dis. 2014; 5:e1141.
45. Li L, Ng DS, Mah WC, Almeida FF, Rahmat SA, Rao VK, Leow SC, Laudisi F, Peh MT, Goh AM, Lim JS, Wright GD, Mortellaro A, et al. A unique role for p53 in the regulation of M2 macrophage polarization. Cell Death Differ. 2015; 22:1081–93.
46. Marcel V, Fernandes K, Terrier O, Lane DP, Bourdon JC. Modulation of p53β and p53γ expression by regulating the alternative splicing of TP53 gene modifies cellular response. Cell Death Differ. 2014; 21:1377–87.
47. Slatter TL, Hung N, Bowie S, Campbell H, Rubio C, Speidel D, Wilson M, Baird M, Royds JA, Braithwaite AW. Δ122p53, a mouse model of Δ133p53α, enhances the tumor-suppressor activities of an attenuated p53 mutant. Cell Death Dis. 2015; 6:e1783.
48. Fiori ME, Barbini C, Haas TL, Marroncelli N, Patrizii M, Biffoni M, De Maria R. Antitumor effect of miR-197 targeting in p53 wild-type lung cancer. Cell Death Differ. 2014; 21:774–82.
49. Hoffman Y, Bublik DR, Pilpel Y, Oren M. miR-661 downregulates both Mdm2 and Mdm4 to activate p53. Cell Death Differ. 2014; 21:302–09.
50. Fortunato O, Boeri M, Moro M, Verri C, Mensah M, Conte D, Caleca L, Roz L, Pastorino U, Sozzi G. Mir-660 is downregulated in lung cancer patients and its replacement inhibits lung tumorigenesis by targeting MDM2-p53 interaction. Cell Death Dis. 2014; 5:e1564.
51. Ren ZJ, Nong XY, Lv YR, Sun HH, An PP, Wang F, Li X, Liu M, Tang H. Mir-509-5p joins the Mdm2/p53 feedback loop and regulates cancer cell growth. Cell Death Dis. 2014; 5:e1387.
52. Liu J, Zhang C, Wang XL, Ly P, Belyi V, Xu-Monette ZY, Young KH, Hu W, Feng Z. E3 ubiquitin ligase TRIM32 negatively regulates tumor suppressor p53 to promote tumorigenesis. Cell Death Differ. 2014; 21:1792–804.
53. Zhang HH, Li SZ, Zhang ZY, Hu XM, Hou PN, Gao L, Du RL, Zhang XD. Nemo-like kinase is critical for p53 stabilization and function in response to DNA damage. Cell Death Differ. 2014; 21:1656–63.
54. Peuget S, Bonacci T, Soubeyran P, Iovanna J, Dusetti NJ. Oxidative stress-induced p53 activity is enhanced by a redox-sensitive TP53INP1 SUMOylation. Cell Death Differ. 2014; 21:1107–18.
55. Sane S, Abdullah A, Boudreau DA, Autenried RK, Gupta BK, Wang X, Wang H, Schlenker EH, Zhang D, Telleria C, Huang L, Chauhan SC, Rezvani K. Ubiquitin-like (UBX)-domain-containing protein, UBXN2A, promotes cell death by interfering with the p53-Mortalin interactions in colon cancer cells. Cell Death Dis. 2014; 5:e1118.
56. Landré V, Rotblat B, Melino S, Bernassola F, Melino G. Screening for E3-ubiquitin ligase inhibitors: challenges and opportunities. Oncotarget. 2014; 5:7988–8013.
57. Grigoreva TA, Tribulovich VG, Garabadzhiu AV, Melino G, Barlev NA. The 26S proteasome is a multifaceted target for anti-cancer therapies. Oncotarget. 2015; 6:24733–49. doi: 10.18632/oncotarget.4619.
58. Lezina L, Aksenova V, Fedorova O, Malikova D, Shuvalov O, Antonov AV, Tentler D, Garabadgiu AV, Melino G, Barlev NA. KMT Set7/9 affects genotoxic stress response via the Mdm2 axis. Oncotarget. 2015; 6:25843–55. doi: 10.18632/oncotarget.4584.
59. Weilbacher A, Gutekunst M, Oren M, Aulitzky WE, van der Kuip H. RITA can induce cell death in p53-defective cells independently of p53 function via activation of JNK/SAPK and p38. Cell Death Dis. 2014; 5:e1318.
60. Cheng J, Fan YH, Xu X, Zhang H, Dou J, Tang Y, Zhong X, Rojas Y, Yu Y, Zhao Y, Vasudevan SA, Zhang H, Nuchtern JG, et al. A small-molecule inhibitor of UBE2N induces neuroblastoma cell death via activation of p53 and JNK pathways. Cell Death Dis. 2014; 5:e1079.
61. Becker MS, Schmezer P, Breuer R, Haas SF, Essers MA, Krammer PH, Li-Weber M. The traditional Chinese medical compound Rocaglamide protects nonmalignant primary cells from DNA damage-induced toxicity by inhibition of p53 expression. Cell Death Dis. 2014; 5:e1000.
62. Rossi M, Rotblat B, Ansell K, Amelio I, Caraglia M, Misso G, Bernassola F, Cavasotto CN, Knight RA, Ciechanover A, Melino G. High throughput screening for inhibitors of the HECT ubiquitin E3 ligase ITCH identifies antidepressant drugs as regulators of autophagy. Cell Death Dis. 2014; 5:e1203.
63. Melino G. Antidepressants synergize with chemotherapy against cancer stem cells. Aging (Albany NY). 2015; 7:1024–25.
64. Adamovich Y, Adler J, Meltser V, Reuven N, Shaul Y. AMPK couples p73 with p53 in cell fate decision. Cell Death Differ. 2014; 21:1451–59.
65. Fatt MP, Cancino GI, Miller FD, Kaplan DR. p63 and p73 coordinate p53 function to determine the balance between survival, cell death, and senescence in adult neural precursor cells. Cell Death Differ. 2014; 21:1546–59.
66. Yallowitz AR, Alexandrova EM, Talos F, Xu S, Marchenko ND, Moll UM. p63 is a prosurvival factor in the adult mammary gland during post-lactational involution, affecting PI-MECs and ErbB2 tumorigenesis. Cell Death Differ. 2014; 21:645–54.
67. Bunjobpol W, Dulloo I, Igarashi K, Concin N, Matsuo K, Sabapathy K. Suppression of acetylpolyamine oxidase by selected AP-1 members regulates DNp73 abundance: mechanistic insights for overcoming DNp73-mediated resistance to chemotherapeutic drugs. Cell Death Differ. 2014; 21:1240–49.
68. Kostecka A, Sznarkowska A, Meller K, Acedo P, Shi Y, Mohammad Sakil HA, Kawiak A, Lion M, Królicka A, Wilhelm M, Inga A, Zawacka-Pankau J. JNK-NQO1 axis drives TAp73-mediated tumor suppression upon oxidative and proteasomal stress. Cell Death Dis. 2014; 5:e1484.
69. Vanbokhoven H, Melino G, Candi E, Declercq W. p63, a story of mice and men. J Invest Dermatol. 2011; 131:1196–207.
70. Wu J, Liang S, Bergholz J, He H, Walsh EM, Zhang Y, Xiao ZX. ΔNp63α activates CD82 metastasis suppressor to inhibit cancer cell invasion. Cell Death Dis. 2014; 5:e1280.
71. Giacobbe A, Compagnone M, Bongiorno-Borbone L, Antonov A, Markert EK, Zhou JH, Annicchiarico-Petruzzelli M, Melino G, Peschiaroli A. p63 controls cell migration and invasion by transcriptional regulation of MTSS1. Oncogene. 2015.
72. Celardo I, Antonov A, Amelio I, Annicchiarico-Petruzzelli M, Melino G. p63 transcriptionally regulates the expression of matrix metallopeptidase 13. Oncotarget. 2014; 5:1279–89. doi: 10.18632/oncotarget.1778.
73. Memmi EM, Sanarico AG, Giacobbe A, Peschiaroli A, Frezza V, Cicalese A, Pisati F, Tosoni D, Zhou H, Tonon G, Antonov A, Melino G, Pelicci PG, Bernassola F. p63 Sustains self-renewal of mammary cancer stem cells through regulation of Sonic Hedgehog signaling. Proc Natl Acad Sci USA. 2015; 112:3499–504.
74. Melino G, Memmi EM, Pelicci PG, Bernassola F. Maintaining epithelial stemness with p63. Sci Signal. 2015; 8:re9.
75. Amelio I, Inoue S, Markert EK, Levine AJ, Knight RA, Mak TW, Melino G. TAp73 opposes tumor angiogenesis by promoting hypoxia-inducible factor 1α degradation. Proc Natl Acad Sci USA. 2015; 112:226–31.
76. Billon N, Terrinoni A, Jolicoeur C, McCarthy A, Richardson WD, Melino G, Raff M. Roles for p53 and p73 during oligodendrocyte development. Development. 2004; 131:1211–20.
77. Agostini M, Tucci P, Killick R, Candi E, Sayan BS, Rivetti di Val Cervo P, Nicotera P, McKeon F, Knight RA, Mak TW, Melino G. Neuronal differentiation by TAp73 is mediated by microRNA-34a regulation of synaptic protein targets. Proc Natl Acad Sci USA. 2011; 108:21093–98.
78. Agostini M, Tucci P, Steinert JR, Shalom-Feuerstein R, Rouleau M, Aberdam D, Forsythe ID, Young KW, Ventura A, Concepcion CP, Han YC, Candi E, Knight RA, et al. microRNA-34a regulates neurite outgrowth, spinal morphology, and function. Proc Natl Acad Sci USA. 2011; 108:21099–104.
79. He Z, Agostini M, Liu H, Melino G, Simon HU. p73 regulates basal and starvation-induced liver metabolism in vivo. Oncotarget. 2015; 6:33178–90. doi: 10.18632/oncotarget.5090.
80. Prensner JR, Chinnaiyan AM. The emergence of lncRNAs in cancer biology. Cancer Discov. 2011; 1:391–407.
81. Yahya SM, Elsayed GH. A summary for molecular regulations of miRNAs in breast cancer. Clin Biochem. 2015; 48:388–96.
82. Scott GK, Goga A, Bhaumik D, Berger CE, Sullivan CS, Benz CC. Coordinate suppression of ERBB2 and ERBB3 by enforced expression of micro-RNA miR-125a or miR-125b. J Biol Chem. 2007; 282:1479–86.
83. Gregory PA, Bert AG, Paterson EL, Barry SC, Tsykin A, Farshid G, Vadas MA, Khew-Goodall Y, Goodall GJ. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol. 2008; 10:593–601.
84. Zhao JJ, Lin J, Yang H, Kong W, He L, Ma X, Coppola D, Cheng JQ. MicroRNA-221/222 negatively regulates estrogen receptor alpha and is associated with tamoxifen resistance in breast cancer. J Biol Chem. 2008; 283:31079–86.
85. Mourtada-Maarabouni M, Pickard MR, Hedge VL, Farzaneh F, Williams GT. GAS5, a non-protein-coding RNA, controls apoptosis and is downregulated in breast cancer. Oncogene. 2009; 28:195–208.
86. Han L, Yuan Y, Zheng S, Yang Y, Li J, Edgerton ME, Diao L, Xu Y, Verhaak RG, Liang H. The Pan-Cancer analysis of pseudogene expression reveals biologically and clinically relevant tumour subtypes. Nat Commun. 2014; 5:3963.
87. Ciriello G, Gatza ML, Beck AH, Wilkerson MD, Rhie SK, Pastore A, Zhang H, McLellan M, Yau C, Kandoth C, Bowlby R, Shen H, Hayat S, et al, and TCGA Research Network. Comprehensive Molecular Portraits of Invasive Lobular Breast Cancer. Cell. 2015; 163:506–19.
88. Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T, Davies S, Fauron C, He X, Hu Z, Quackenbush JF, Stijleman IJ, Palazzo J, et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol. 2009; 27:1160–67.

