INTRODUCTION
Recent large-scale genome wide studies have underscored a complex role of NOTCH1 as a candidate tumor suppressor harboring inactivating mutation and deletions , as well as a driver of tumorigenesis harboring activating missense mutations and amplifications in a context dependent manner in HNSCC, and other cancers [1-10]. In addition, Notch signaling pathway plays a significant role in the maintenance of cancer stem-like population of cells (CSCs) in several human cancers [11-15] . Inhibition of Notch signaling prevents the formation of secondary mammospheres by cell lines derived from primary breast cancer patient samples. However, the biological significance of cancer stem-like cells (CSCs) in HNSCC has not been well characterized.
To understand the role of Notch signaling pathway in early-stage (T1-T2) tongue tumors, we examined the mutational landscape, copy number alterations and differential expression of receptor, ligands, modifiers and target genes of the Notch pathway, along with effect of genetic and pharmacologic perturbation of Notch pathway on cancer stem-like cells (CSCs) features of HNSCC cells.
RESULTS
All the samples with available genomic DNA were tested for the presence of HPV using MY09/11 PCR and E6 transcript PCR primers. 40 of 71 samples analyzed, all were found to be HPV negative. Where exome sequence was available, the absence of HPV was re-confirmed using HPVDetector, as previously described [16]. TSCC samples of Indian origin to be HPV negative is consistent with other studies [17-19].
Notch pathway is activated in early TSCC patients
To characterize somatic alterations across 48 genes of Notch signaling pathway in 29 early-stage (T1-T2) tongue squamous cell carcinoma (TSCC) patient-derived tumors, we analyzed 23 paired whole exome and 10 whole transcriptome tongue cancer tumor sequencing data (unpublished data), as detailed in Supplementary Table S1 and Supplementary Figure S1A. Fourteen mutations were observed in 7 genes across 12 of 22 samples (Supplementary Table S1). Of note, inactivating NOTCH1 mutation (4%) were found at a lower frequency in our sample set than that reported from the Caucasian population [7, 8, 20] but consistent with similar finding from a recent Asian study [21, 22]. In further contrast to Caucasian population, we observed Notch family receptors, ligands, and downstream effector genes were amplified or over expressed in 59% samples (17 of 29 patients) based on copy number variations called from whole- exome and whole- transcriptome data. To extend and validate these findings, we performed real-time quantitative PCR to estimate DNA copy number and transcript levels, along with an immunohistochemical analysis of Notch pathway components in paired tumor-normal samples from tongue cancer patients. We found somatic amplification at NOTCH1 in 12 of 38 tumors (Figure 1A); overexpression of NOTCH1 transcripts was observed in 16 of 45 samples (Figure 1B, 1C)-- consistent with our analysis of the TCGA TSCC data set (n=126) (Supplementary Figure 1B, 1C), not reported earlier. Also, samples harboring amplification at NOTCH1 (P value <0.001) and DLL4 (P value <0.001) showed significantly higher expression of transcript as compared to no amplification. (Supplementary Figure S1D and Supplementary Figure S2). Consistent with amplification and over expression of Notch pathway components, Immunohistochemical analysis for activated NOTCH1 intracellular domain (NICD) in a set of 50 patients indicated strong immunoreactivity for active Notch signaling present in 40% tumor samples (Figure 1D-1E, Supplementary Figure S3A-S3C).
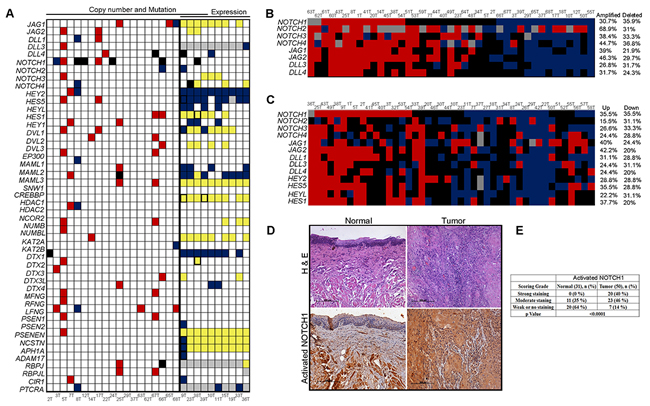
Figure 1: Activation of Notch pathway in early stage tongue squamous cell carcinoma. A. Schematic representation of somatic mutation, copy number changes and expression changes identified in Notch pathway genes (N=48) using whole exome and transcriptome sequencing. Red filled; copy number gains, Yellow; high transcript expression, blue; copy number loss and low transcript expression, black; mutation, white; no events, grey; transcript not detected and black borderline boxes ;any two events. Thick black line denoting separation of samples with exome and transcriptome sequencing. B. Schematic representation of DNA copy number alteration of Notch pathway genes in a cohort of 41 paired tumor samples estimated by quantitative real-time PCR. Red blocks; high copy number, blue; low copy number, black; diploid and grey color; experiment could not be done or data could not be acquired. C. Schematic representation of gene expression of Notch signaling pathway and its downstream targets in the cohort of 44 paired tongue tumor samples. Colors denotes: Red; upregulation, blue; down regulation, black; basal expression and grey color; experiment could not be done or results could not be acquired D. Immunohistochemistry (IHC) was performed for activated NOTCH1 in paired normal and tongue tumor samples (N=50). Brown color indicates positive expression. Representative IHC stained photomicrographs from normal and tongue tumor samples are shown. Scale bar, 100µM; corresponding H&E stained slides are shown in the upper panel. E. Tabular representation for quantification of activated NOTCH1 immunostaining data. Significant differences of IHC staining scores between normal and tumor were estimated using the Chi-square test and p value ≤0.05 was considered as threshold for statistical significance.
Expression of NOTCH1 is required for survival, migration and stemness of TSCC tumor cells
To assess the functional significance of Notch pathway activation, we asked if the expression of NOTCH1 is essential for survival, migration and stem-like feature of HNSCC cells in vitro. First, we checked for the presence of Notch pathway transcript expression by real-time PCR and western analysis of NOTCH1 protein using multiple head and neck cancers cell lines (NT8e, AW13516, CAL27 and DOK) [23]. NT8e and CAL27 cells showed higher expression of NOTCH1 as compared to AW13516 and DOK cells (Supplementary Figure S4A, S4B). Next, we tested a series of shRNA constructs to knockdown NOTCH1 in these cells. The knockdowns were confirmed by western blot analysis for NOTCH1 (Figure 2A) and quantitative real-time PCR for NOTCH1 and its target gene HES1 (Supplementary Figure S4C). We identified two shRNA clones sh1 and sh2 that efficiently knocked down expression of NOTCH1 compared to scrambled (SCR). Knock down of NOTCH1 inhibited cell survival (Figure 2B), anchorage-independent growth (Figure 2C), in NT8e and CAL27, and migration in NT8e (Figure 2D).
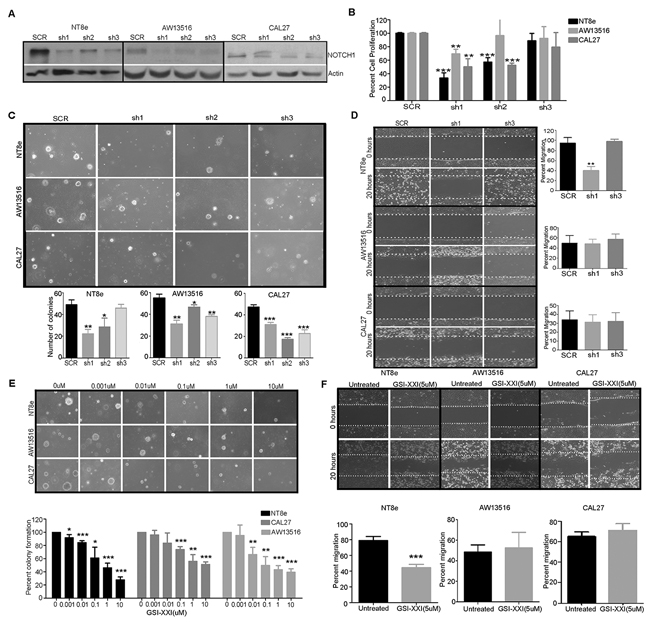
Figure 2: shRNA mediated knockdown and inhibition of NOTCH1 inhibits transformation, survival and migration of HNSCC cells. A. shRNA constructs used to knock down NOTCH1 expression in NT8e, AW13516, and CAL27 cells. Anti-NOTCH1 immunoblot shows that hairpins knock down to varying extents in different cells. Actin is included as a loading control. SCR, scrambled hairpin used as a negative control. B. Infection with 2 of 3 independent hairpins (shNOTCH1#1 and shNOTCH1#2) inhibited cell survival of NT8e and CAL27 cells expressing higher NOTCH1 levels as compared to the AW13516 cells-- as assessed by plotting total cell count on day 6 compared to day 2, normalized against cells infected with SCR. C. Infection with independent hairpins inhibit soft agar colony formation by the NT8e and CAL27 cells expressing higher NOTCH1 levels compared to the AW13516 cells (upper panel). Colonies were photographed after 3 weeks (Magnification: ×10). Bar graph representation of soft agar colony formation (lower panel). D. Wound healing assay of knockdown clones of NT8e, CAL27 and AW13516 cells. NT8e cells with highest migration potential was most significantly inhibited following infection with shNOTCH1 constructs. Percent inhibition of migration was calculated after 20 hours of wound incision. E. Representative images of soft agar colony formation (upper panel) and bar graph representation of soft agar colony formation post gamma secretase inhibitor (GSI-XXI) treatment in HNSCC cell lines F. Wound healing assay of NT8e, CAL27 and AW13516 cells were performed post GSI-XXI inhibitor as indicated concentration and % migration was calculated after 20 hours of wound healing. Experiment was performed in triplicate and colonies were counted and shown as mean ± SD and P value is denoted as *; P < 0.01, **; P < 0.001, ***; P < 0.0001 versus non-targeting shRNA. Experiments were repeated two times independently.
Expression of NOTCH1 and its pathway genes maintains cancer stem-like cells (CSCs) in various tumors, as determined by their ability to form spheroids and expression of molecular markers ALDH1, CD133 and CD44 [12, 24]. An in vitro spheroid formation assay was performed to examine the cancer stem cell population (CSCs) in HNSCC cell lines (NT8e, CAL27, AW13516, and DOK) expressing a variable level of NOTCH1 expression (Supplementary Figure S4B). As shown in Figure 3A, following 10 days of incubation in undifferentiating stem cell media, NOTCH1 over expressing NT8e cells showed a higher number of oralspheres with 32% and 0.21% NT8e cells for cancer stem-like cells molecular marker such as ALDH and CD133, respectively. Similarly, CAL27 cells also showed a significantly higher number of oralspheres with 13.5% and 1.59% CAL27 cells positive for ALDH and CD133 (Figure 3B, 3C). In contrast, AW13516 cells expressing comparatively lower NOTCH1 levels showed a reduced spheroid formation capacity with 0.34% ALDH positive and 0.12% CD133 positive while DOK cells did not show any oralsphere formation. To test whether a high fraction of the NT8e population constitutes the stem-like cells, we sorted NT8e cells in ALDH positive and ALDH negative fraction and assessed the sphere-forming efficiency (Supplementary Figure S5A). With subsequent passaging, the cells form ALDH negative population could not maintain their spheroid formation capacity while the ALDH positive population retained their self-renewal capacity demonstrating that indeed NT8e possess high ALDH positive cells are showing cancer stem-like cells features (Supplementary Figure S5B, S5C). NOTCH1 knockdown clones showed significant reduction in oralsphere formation ability with concomitant decrease in ALDH positive cells in NT8e and AW13516 cells as compared to scrambled (SCR) cells (Figure 3D, 3E), highlighting their dependency on NOTCH1 expression with concomitant decrease in ALDH positive population of cells, thus regulating and promoting the survival of HNSCC CSCs. Next, we attempted to overexpress activated NOTCH1 and full-length NOTCH1 in AW13516 cells and checked for the sphere forming efficiency. Activated NOTCH1 form more number of spheres as compared to vector control cells post 5 days (Supplementary Figure S6, S6C). However, given that AW13516 cells are HPV negative [25] and harbor wild-type p16INK4A and mutant Tp53 [23], ectopic expression of full-length NOTCH1 or NICD led to continuous cell death and senescence mediated growth arrest (Supplementary Figure S6, S6G), as described earlier [26-28].
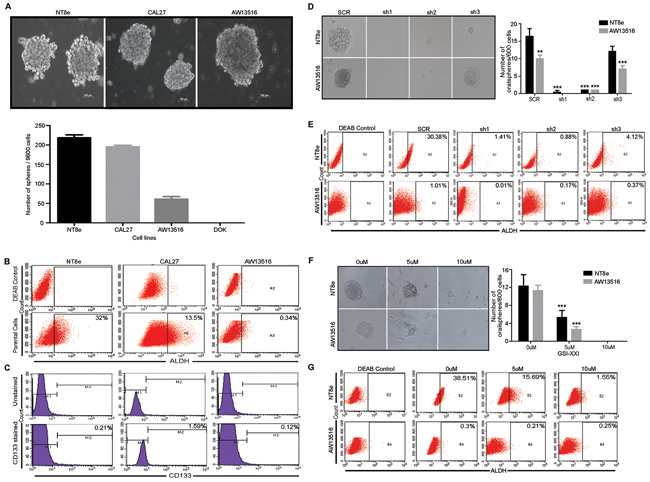
Figure 3: Notch pathway is essential for cancer stem-like property of HNSCC cells. A. Oralsphere formation capacity of HNSCC cells. Representative images of oralsphere are shown in HNSCC cells. Oralsphere (>75µm size) were counted manually in triplicate via visualization under microscope and data was represented as Mean± SD. B. and C. Analysis of cancer stem-like cells (CSCs) marker ALDH and CD133 in HNSCC cells. Percentage ALDH positive cells were calculated against DEAB control. D. Representative images of oralsphere are shown in scrambled (SCR) and different shRNA clones (sh1, sh2 and sh3) of NT8e and AW13516 cells. Oralsphere formation assay was performed in triplicate and counting was done by observing under phase contrast microscope and data was represented as Mean± SD. E. ALDH staining for shRNA mediated knockdown clones and GSI-XXI treatment in HNSCC cells, respectively. Percentage ALDH positive cells were calculated against DEAB control. F. and G. Representative images of oralsphere post GSI-XXI treatment. NT8e and AW13516 cells post respective concentration treatment and ALDH positive cells. Number of oralsphere were counted and represented as Mean± SD. ALDH staining for shRNA mediated knockdown clones and GSI-XXI treatment in HNSCC cells, respectively. Percentage ALDH positive cells were calculated against DEAB control P-value *; ≥ 0.05 was considered as threshold for significance. All the above experiment were performed by at least two times independently by separate individuals.
Notch pathway inhibitors block stem-like feature, proliferation, and survival of HNSCC cells over expressing NOTCH1
Finally, we investigated whether pharmacological inhibition of Notch pathway activation would be effective against HNSCC cell lines over expressing NOTCH1. Treatment of the NT8e, CAL27, AW13516 and DOK HNSCC cell lines with gamma secretase inhibitor (GSI-XXI) that abolished the presence of activated NOTCH1 (Supplementary Figure S4D) that resulted in significant reduction in soft agar colony formation (Figure 2E) and cell survival (Supplementary Figure S4E) as compared to vehicle treated in NT8e and CAL27 cells but not AW13516 cells. Additionally, the migration potential of NT8e cells was significantly inhibited by GSI-XXI, consistent with our observation using shRNA knockdown based approach (Figure 2F). Similarly, marked decrease in spheroid forming ability and ALDH expression in NT8e and AW13516 cells was observed post GSI-XXI treatment (Figure 3F, 3G).
Activation of notch pathway correlates with node positive and non-smoker TSCC patients
Of particular significance is the correlation between clinicopathological characteristics and overall Notch pathway activation: immuno-histochemical based expression of activated NOTCH1 intracellular domain NICD (χ2=7.10, P=0.029), amplification at DLL4 (χ2=7.5, P=0.023), and transcript over expression of Notch pathway effector genes HEY2 (χ2=9.8, P=0.007) and HES5 (χ2=5.71 P=0.057) significantly correlated with lymph node metastases (Table 1) and poor prognosis (Supplementary Figure S7A). Interestingly, this was consistent also with our analysis of the TCGA tongue cancer patient dataset (Supplementary Figure S7B), and with other cancers [24, 29, 30].
Table 1: Clinical correlation analysis of Notch pathway alterations
Clinicopathologic features |
Variable |
N (% along column) |
DLL3 Copy Number (N=26) N (% along row) |
P- value |
N, % along column |
HEY2 Expression (N=34) N (% along row) |
P- value |
||||
|---|---|---|---|---|---|---|---|---|---|---|---|
Gain (N=8) |
Diploid (N=16) |
Deleted (N=2) |
Up (N=15) |
Basal (N=12) |
Down (N=7) |
||||||
Nodal Status |
Node positive |
19 (73%) |
5 (26%) |
14 (74%) |
0 (0%) |
0.023 |
22 (65%) |
12 (55%) |
9 (41%) |
1 (5%) |
0.007 |
Node negative |
7 (27%) |
3 (43%) |
2 (29%) |
2 (29%) |
12 (35%) |
3 (25%) |
3 (25%) |
6 (50%) |
|||
Clinicopathologic features |
Variable |
N (% along column) |
HES5 Expression (N=34) N (% along row) |
P- value |
N, % along column |
Activated NOTCH1 IHC (N=49) N (% along row) |
P- value |
||||
|---|---|---|---|---|---|---|---|---|---|---|---|
Up (N=15) |
Basal (N=12) |
Down (N=7) |
Strong (N=15) |
Moderate (N=12) |
Weak or No (N=7) |
||||||
Nodal Status |
Node positive |
23 (66%) |
9 (39%) |
12 (52%) |
2 (9%) |
0.057+ |
23 (66%) |
13 (43%) |
15 (50%) |
2 (7%) |
0.029 |
Node negative |
12 (34%) |
4 (33%) |
3 (25%) |
5 (42%) |
12 (34%) |
6 (32%) |
6 (32%) |
7 (37%) |
|||
Clinicopathologic features |
Variable |
N (% along column) |
NOTCH1 transcript expression (N=35), % along row |
P- value |
||
|---|---|---|---|---|---|---|
Up (N=14) |
Basal (N=9) |
Down (N=12) |
||||
Smoking |
Smoker |
11 (32%) |
2 (18%) |
6 (55%) |
3 (27%) |
0.026 |
Non-smoker |
24 (68%) |
12 (50%) |
3 (13%) |
9 (38%) |
||
All clinical correlation analysis were performed in SPSS and significant alterations has been presented in the table. Patient’s samples showing strong and moderate staining of activated NOTCH1 was considered as having activated Notch signaling. N; Number of samples, Up; upregulation, Down; downregulation. Chi-square test was used to calculate statistical significance. Significant P-value are highlighted in bold font. P-value ≤ 0.05 was considered as threshold for significance. + denotes; marginally significant.
In addition, NOTCH1 expression significantly correlated with a non-smoking habit of patients (χ2=7.325, P=0.026), where 12 of 24 non-smokers patients derived tumors showed upregulation of NOTCH1 transcript, consistent with previously described NOTCH1 upregulation in non-smokers in other diseases including lung adenocarcinoma [31-33]. We also observed a significant correlation with AJCC (American Joint committee on Cancer) TNM tumor staging wherein stage III-IVA showed increases activation of NOTCH pathway (χ2=7.84, P=0.02). However, no statistically significant correlation was observed between the activated NOTCH1 expression with the sex, age, alcohol and tobacco consumption in the cohort, as represented in Supplementary Table S2. Next, we performed an interim analysis and assessed disease-free survival (DFS) by IHC defined activated NOTCH1 (strong and moderate staining) status vs non-activated NOTCH1 (weak or no staining) status. DFS was defined as time interval between the date of registration and the date of first documented evidence of relapse at any site (local, regional, metastatic, or secondary primary) or death from any cause, whichever earlier. There was no statistically significant difference was observed in tumors with activated NOTCH1 compared to those with non-activated NOTCH1 tumors, as shown in Supplementary Figure S7C.
Taken together, we present a novel clinicopathological correlation such that expression of Notch pathway components and activated NOTCH1 levels predispose TSCC patients to lymph node metastasis, and that non-smokers TSCC patients tend to have higher NOTCH1 levels as compared to smokers. Clinically, determination of NICD by immuno-histochemistry could be a good predictor of nodal status. This could be a biomarker to predict lymph node metastasis for therapeutic utility among early stage tongue cancer patient to help patient stratification for treatment [34, 35].
DISCUSSION
We demonstrate that 40% TSCC tumors have strong Notch pathway activation and that this property may be important in the maintenance of stem cell component in these tumors. Genetic or chemical perturbation of NOTCH pathway using shRNA and GSI-XXI showed decrease soft agar colony formation, migration potential and cancer stem-like features of HNSCC cells, highlighting their dependency on NOTCH1 expression. Thus, targeted elimination of these cells may provide a new lead in treatment of head and neck cancer. These findings are consistent with reports where Notch signaling has been shown to be required for stem cell-like features in several cancer types [4, 13, 37]. Interestingly, genetic determinants of cancer stem cells share features with their role in the development of tumorigenesis [38]. These findings are consistent with reports where Notch signaling has been shown to be required for stem cell-like features in several cancer types, including HNSCC [14, 15].
Clinically, NOTCH1 transcript expression significantly correlate with non-smoking habit of patients, consistent with previous reports in other pathological conditions including lung adenocarcinoma [31-33]; lymph node metastasis in tongue cancer correlate with poor prognosis and survival of the patients, thus activated NOTCH1 could serve as a reliable marker to predict lymph node metastasis [35]. Moreover, AJCC TNM tumor stage III-IVA significantly correlates with activation of NOTCH pathway as compared to stage I-II, consistent with reports in HNSCC [39]. The sample size in this study, however, is underpowered to reach the statistical significance for survival data. No significant difference was observed in disease-free survival of the patients with IHC defined activated NOTCH1 tumors as compared to non-activated NOTCH1, as shown in Supplementary Figure S7C.
In conclusion, we demonstrate that a considerable fraction of TSCC tumors has upregulated Notch pathway and that this property may be important for the maintenance of stem cell component in these tumors. And that, NOTCH1 could be a potential therapeutic target in these patients.
MATERIALS AND METHODS
Patient Samples
Tumor-normal paired samples were collected at Tata Memorial Hospital and Advanced Centre for Treatment, Research and Education in Cancer (ACTREC), Mumbai. Sample set and study protocols were approved by (ACTREC-TMC) Internal Review Board (IRB) and most of the patients were recruited from 2010-2013 with a predefined inclusion criteria of early (pT1 and pT2) stage. Percent tumor content was determined using hematoxylin and eosin based staining by two independent pathologists which varied from 60 to 90%. Patient samples and characteristics are provided in the Supplementary Table S3.
DNA and RNA extraction
DNA from tongue primary paired normal-tumor tissue samples were extracted using DNeasy Blood and tissue DNA extraction kit (Qiagen) according to manufacturer’s instructions. DNA was quantified using Nanodrop 2000c Spectrophotometer (Thermo Fisher Scientific Inc.) and DNA quality was checked by resolving on 0.8% agarose gel. Total RNA was extracted from tongue primary paired normal-tumor samples and cell lines using RNeasy RNA isolation kit (Qiagen) and Trizol reagent (Invitrogen) based methods and later resolved on 1.2% Agarose gel to confirm the RNA integrity.
Exome capture and NGS DNA sequencing
Two different Exome capture kits were used to capture exome for different samples. The TruSeq Exome Enrichment kit (Illumina) was used to capture 62Mb region of human genome comprising of 201,121 exons representing 20,974 gene sequences, including 5’UTR, 3’UTR, microRNAs and other non-coding RNA and NimbleGen SeqCap EZ Exome Library v3.0 was also used to capture 64 Mb region of the human genome. Exome library preparation and sequencing was performed as per manufacturer’s instructions. Briefly, 2 µg genomic DNA was sheared using Covaris (Covaris Inc) for generating the fragment size of 200-300bp size. DNA libraries were prepared from both the kits were quantified by qPCR using KAPA Library Quant Kit (Kapa Biosystems) in ABI 7,900HT system (Life Technologies). Seven pmol of 6-plex DNA library pool was loaded per lane on flow cell (Flow Cell v3) to generate clusters using TruSeq PE (Paired-End) Cluster Kit v3-cBot-HS kit and clustered flow was sequenced for 201 cycles on HiSeq-1,500 System (Illumina) using TruSeq SBS Kit v3 (Illumina) at in-house core NGS facility.
Identification of somatic mutations from exome Sequencing
Paired-end raw sequence reads generated were mapped to the human reference genome (build hg19) using BWA v. 0.6.2 [40]. Mapped reads were then used to identify and remove PCR duplicates using Picard tools v1.100 (http:broadinstitute.github.io/picard/). Base quality score recalibration and indel re-alignment performed and variants were called from each sample separately using GATK 2.5-2 Unified Genotyper and MuTect v. 1.0.27783 [41]. Post subtraction of variants from its paired normal, remaining variants was taken for further analysis if they were having ≥5 altered reads. Furthermore, all samples variants were further filtered against pooled normal variants database (N=62) to reduce the possibility of the germline variation We further annotated variants using Oncotator v1.1.6.0 [42] and dbSNP v142 [43] and COSMIC database v68 [44] using an in-house developed script. Later, we performed functional prediction tool based analysis for somatic non-synonymous variants using nine different tools such as: dbNSFP v2.0 (includes SIFT, Polyphen2_HDIV, Polyphen2_HVAR, LRT, Mutation Taster, Mutation Accessor and FATHMM) [45], CanDRA v1.0 [46] and Provean v.1.1 [47]. Variants called deleterious in nature by at least one software was taken for further analysis. We confirmed the identity of mutations by manual visualization in IGV [48, 49].
Somatic copy number analysis from exome sequencing data
BAM files prepared for variant calling were used for copy number analysis using Control-FREEC [50]. Paired tumor-normal samples BAM files were fed into Control-FREEC along with target region for Illumina and Nimblegen exome kits as bed file. Read count were generated and normalized for GC content for each of the target region followed by computation of ratio of read count in a tumor to normal. Read count ratio was converted to copy numbers followed by segmentation using lasso method. Segmented copy number data generated by control-FREEC was further used for annotation and post-processing using R programming.
Transcriptome sequencing and data analysis
Transcriptome libraries for sequencing were constructed according to the TruSeq RNA library protocol (Illumina). Briefly, mRNA was purified from 4 µg of intact total RNA using oligodT beads and library preparation was done as per manufacturer’s instructions (TruSeq RNA Sample Preparation Kit, Illumina). 7pmol of quantified cDNA libraries were loaded on Illumina flow cell (v3) to generate clusters using TruSeq PE (Paired-End) Cluster Kit v3-cBot-HS kit and clustered flow was sequenced for 201 cycles on HiSeq-1500 System (Illumina) using TruSeq PE Cluster Kit v3 and TruSeq SBS Kit v3 (Illumina) to generate at least 30 million reads per sample. Post sequencing, de-multiplexing was carried out on the basis of index sequences using CASAVA (version 1.8.4, Illumina). Transcriptome data analysis was performed using Tuxedo-suite pipeline [51]. In brief, alignment of short reads was done against reference genome (hg19) using TOPHAT2 v. 2.0.8b [52] in which 95-99% of reads were mapped to the reference genome. Cufflinks v.2.0.2 was used to find the expressed transcripts in the data and quality control steps was performed using CummeRbund package v2.0. All the actively expressed transcripts per samples were then binned by log10(FPKM+1) to differentiate the significantly expressed transcripts from the background noise and transcripts represented by <0.1 log10(FPKM+1) were filtered out from further analysis. Since paired normal of these tumors cannot be obtained, we defined a significant change in expression for those genes whose expression is higher (>80%) or lower (<20%) than the median expression as suggested [53].
Analysis of the cancer genome atlas tongue cancer data
The Cancer Genome Atlas (TCGA) dataset of HNSCC including DNA copy number dataset (gistic2 threshold) from 452 HNSCC tumor tissue, RNA seq expression (Illumina HiSeq) dataset from 541 HNSCC was downloaded from UCSC Cancer genome browser on 20th June 2014. Later, tongue cancer patient data for DNA copy number (n=126) and gene expression (n=129 has been taken for further analysis. For expression and DNA copy number the median centered RSEM counts and gene-level copy number estimates have been used, respectively (n=126). Notch pathway genes (n=13) data has been retrieved and heatmaps were generated using MeV 4.9.0. The RNAseq gene expression data has been retrieved for Notch pathway genes (n=13) and raw data has been median centered using Cluster 3.0 software. The median centered RSEM counts for each gene has been used to generate heatmap using MeV 4.9.0. Fold change criteria was ≥1.5 fold change for upregulation, ≤1.499 fold change for down-regulation and in between -1.5 to 1.499 fold change was denoted as a basal expression. DNA copy number and Expression correlation analysis and clinical correlation analysis have been performed using SPSS. P=value <0.05 was criteria for statistical significance.
Tissue processing
Surgically resected oral tumor tissues and matched nonmalignant (cut margins) adjacent tissues were obtained from patients with informed consent after IRB approval from ACTREC. These tissues were processed for paraffin embedding and sectioned at 4µm for H/E staining for evaluation of tumor.
Immunohistochemistry
Immunohistochemistry was done following the standard protocol of DAKO Envision Flex. Briefly, the slides were microwaved by incubating them for 10 minutes in high pH antigen Retrieval Solution (DAKO;DM828), then allowed to cool to room temperature before rinsing with Tris-buffered saline wash buffer (DAKO;DM831). Endogenous peroxidase activity was blocked by incubating the slides for 20 minutes in 3% hydrogen peroxide (EnVision/HRP, Dako). After rinsing in wash buffer, the sections were incubated for 3hours at room temperature with the monoclonal human anti-activated Notch1 antibody (Cat.ab8925; dilution 1:50) in Tris-HCl buffer antibody diluent (Dako; K8016). Slides were rinsed in wash buffer (DAKO; DM831) and incubated for 90 minutes with peroxidase-labeled polymer conjugated to goat anti-rabbit immunoglobulins (EnVision/HRP, Dako; SM801). The chromogenic reaction was carried out with 3,3′-diaminobenzidine chromogen solution for 5 minutes, resulting in the expected brown-colored signal. Finally, after rinsing with deionized water, the slides were counterstained with hematoxylin, dehydrated, mounted with toluene-based mounting medium (Thermo Scientific Richard-Allan) and cover slip.
Immunohistochemical staining analysis
Evaluation of immunohistochemical staining of activated Notch1 expression was scored as 0, 1+, 2+ and 3+. The percentage of cells with positive staining was scored from 0 to 4 (0=0% positive cells; 1: <10% positive cells; 2: 10-49% positive cells; 3: 50-80% positive cells; 4: >80% positive cells) and staining intensity was scored from 0 to 3 (0, negative; 1, weak; 2, moderate; 3, strong). The two scores were then multiplied. Final scores of 0-2 were scored as 0, 3-5 as 1, 6-8 as 2 and 9-12 as 3.6
Quantitative real-time PCR for copy number analysis
Primers details used for copy number study has been provided in Supplementary Table S4. All primers used have been tested for their specificity by performing evaluative PCR as well as melt curve analysis during quantitative real-time PCR. Amplification efficiency for all primer was tested with series of dilutions (0.625 ng, 1.25 ng, 2.5 ng, 5 ng, 10 ng) of genomic DNA and PCR amplification efficiency was ~97%(~R2=0.979) (Supplementary Figure S8A, S8B). Based on above quality control, 10 ng of genomic DNA per 10 µl reaction volume in triplicates were run on Light cycler 480 (Roche, Mannheim, Germany) twice independently and relative copy number analysis was performed as described previously [54]. The threshold for calling high and low copy number was ≥2.5 and ≤1.5, respectively and ≤2.5 and 1.5≤ ; diploid.
Quantitative real-time RT-PCR for expression analysis
Prepared cDNA was diluted 1:10 and reaction were performed in 10µl volume in triplicate. The melt curve analysis was performed to check the primer dimer or non-specific amplifications. Real-time PCR was carried out using KAPA master mix (KAPA SYBR® FAST Universal qPCR kit) in 10 µl volume in triplicate on Light cycler 480 (Roche, Mannheim, Germany) machine. All the experiments were repeated at least twice independently. The data was normalized with internal reference GAPDH, and analyzed by using delta-delta Ct method described previously The criteria were ≥2 fold change for upregulation, ≤0.5 fold change for down-regulation and in between 1.99-0.501 fold change as a basal expression. The details of all the primers used for expression analysis have been provided in Supplementary table S4.
Cell culture
Cell lines established from different sub-sites of head and neck cancer: AW13516 from tongue, NT8e from upper aero-digestive tract, CAL27 cells from tongue [55] and partially transformed cell line DOK (tongue) [56] were used in this study. AW13516 and NT8e were acquired from Tata Memorial Hospital while CAL27 and DOK cells were procured from ATCC and Sigma, respectively. All cells were grown in Dulbecco’s Modified Eagle Medium (Pan biotech, Germany). Culture media was supplemented with 10% FBS (Gibco, US), 1% Penicillin-Streptomycin solution (Sigma) and maintained at 37°C in an incubator with 5% CO2. DOK cells were grown with 5ug/ml hydrocortisone (Sigma) as a supplement. Trypsinization was performed using 0.25% Trypsin-EDTA (Invitrogen) and freezing of cells performed in 90 % FBS (Gibco, US) and 10% DMSO (Sigma) and were stored in liquid Nitrogen for long term storage. All the cell lines were authenticated using a short tandem repeat (STR) analysis kit (Gene Print v10, Promega, USA). The results are shown in Supplementary Table S5.
Retrovirus production, infection and drug selection
Retroviral shRNA constructs were purchased from TransOMIC technologies, USA. Target sequences of NOTCH1 shRNA constructs: sh1 5’-CAGTGAGCGATGACTGCACAGAGAGCTCCTAT-3’, sh2 5’-CAGTGAGCGATGGACGGACCCAACACTTACAT-3’, and sh3 5’-CAGTGAGCGAGACGAGGACCTGGAGACCAAAT-3’. 293T cells were seeded in 6 well plates one day before transfection and each construct (pMLP Retroviral-puro) along with pCL-ECO and pVSVG helper vector were transfected using Lipofectamine LTX reagent (Invitrogen). The viral soup was collected 48 and 72 hours post transfection, passed through 0.45µM filter and stored at 4OC. Respective cells for transduction were seeded one day before infection in a six-well plate and allowed to grow to reach 50-60% confluency. One ml of the virus soup (1:5 dilution) and 8µg/ml of polybrene (Sigma) was added to cells and incubated for six hours. Cells were maintained under puromycin (Sigma) selection.
Overexpression of NOTCH1 and selection
The human full-length NOTCH1 (pcDNA-NOTCH1) [57] was obtained from Artavanis-Tsakonas Laboratory (Havard Medical School) and activated NOTCH1 (pEGFP-NICD) [58] constructs was obtained from Annapoorni Rangarajan (Indian Institute of Sciences (IISc), Bangalore, India). Cells expressing pcDNA-NOTCH1 or pEGFP-NICD were generated by transfection with 10µg of DNA using Lipofectamine 3000 (Invitrogen) as per manufacturer’s instructions. After 48hours, cells were cultures for 8-10 days in complete medium supplemented with 1mg/ml of G418 for antibiotics selection of transfected cells or cells were sorted based on GFP expression using BD FACSAria II. Pooled GFP sorted or antibiotics selected cells were later used for oralsphere assay. In case of 293T cells, post 48hours transfection cells were taken for RNA extraction and protein extraction for quantitative real-time PCR and western blot analysis, respectively.
Western blotting
Cells were lysed in RIPA buffer (Sigma) and protein concentration was estimated using BCA (MP biomedicals) method [59]. Forty microgram protein was separated on 10% SDS-PAGE gel, transferred to nitrocellulose membrane and transfer was verified using Ponceau S (Sigma). Later the blots were blocked in Tris-buffered saline containing 5% BSA (Sigma) and 0.01% Tween-20(Sigma) and were probed with full-length NOTCH1 (sc-6014-R, Santacruz biotechnology), anti-activated NOTCH1 antibody (Abcam; ab8925) and anti-actin (A5316, Sigma) antibody. The membranes were then incubated with corresponding secondary HRP-conjugated antibodies (Santa Cruz Biotechnology, USA) and the immune complexes were visualized by Pierce ECL (Thermo Scientific, USA) according to manufacturer’s protocol. Western blot experiments were performed in triplicate.
Anchorage-independent growth assay
For analysis of growth in soft agar, 5 × 103 cells were seeded in triplicate onto a six-well dish (Falcon) in 4 ml of complete medium containing 0.33% agar solution along with respective treatments of GSI-XXI at 37°C in CO2 incubator. Ten images per well were photographed after 21 days using inverted phase contrast microscope and colonies were counted manually.
MTT assay
A Thousand cells per well (six replicate per concentration) were seeded in 96-well plate followed by incubation with the drug for 72 hours and subsequently incubated with MTT (0.5 mg/ml) for 4 hours. Later, MTT assay was performed and data was acquired at 570nm using Microplate reader. Percentage cell viability was calculated against vehicle treated control.
Wound healing assay
The cells were grown in 6 well plates to 95% confluency and were replaced with fresh medium containing 5/ml mitomycin C (Sigma). After 2 hours incubation, the medium was discarded and wounds were scratched with the help of sterile 10µl pipette tip. Cells were washed with PBS to remove the detached cells during creating a wound. The cells were fed with fresh medium and observed by time-lapse microscopy, and images were taken every 10 min for 20 hr. Migration was measured using Image J software.
Oralsphere formation assay
Ninety-six hundred cells were seeded in 1.2 % agar coated 6-well plates supplemented with stem cell media (recombinant EGF (20 ng/ml), human basic FGF (20 ng/ml), L-glutamine (2 mM), B-27 supplement and N2 supplement) and allowed to grow for 10 days. After every five days media, additional media was supplemented. Five hundred cells from NT8e, AW13516 and CAL27 shRNA clones were seeded on an ultra-low adherent 96-well plate in stem cell medium. Oralspheres were then cultured and maintained in low adherent 24-well plates. Additionally, the parent NT8e, AW13516, and CAL27 cells were also checked for the spheroid formation capacity upon 5 µM and 10 µM GSI-XXI administration using the same conditions.
ALDH activity and CD133 staining
The ALDH activity was checked using ALDEFLUOR™ detection kit (StemCell Technology, 01700) following the kit protocol and data was acquired on FACS Caliber and analysis was carried out using CellQuest software. For CD133 staining was performed using CD133 (AC133) antibody (MACS Miltenyi Biotech) in FACS buffer for 15 min in dark at 4 °C. The cells were then washed twice with staining buffer and acquired on FACS Caliber, BD Biosciences.
β-Galactosidase activity staining
Ten thousand cells were seeded in 12 well plates in triplicates and next day, AW13516 cells, vector control and overexpressing full length NOTCH1 were washed with 1X PBS and fixed with 0.5ml of fixative solution in the Abcam Senescence detection kit (Ab65351) for 10–15min at 25°C. Fixed cells then washed twice with 1X PBS and incubated for 8 hours with 0.5ml of staining solution containing 20mg/ml of X-gal. Stained cells were microscopically analyzed using Olympus IX-71. Images were analyzed using Image J and percentage β-Galactosidase positive cells were plotted.
Survival and Statistical analysis
The relative impact of Notch pathway alterations on disease free survival (DFS) of TSCC patients was analyzed using Kaplan-Meier method [60] and was compared using the log-rank test for statistical significance. Data are expressed as mean ± standard deviation (SD) or standard error (SE). Significant differences between selected two groups were estimated using unpaired Student t-test using Graph Pad prism version 5. Statistical significance was set at p ≤ 0.05. Pearson correlation analysis and chi-square tests were performed in IBM SPSS statistics software version 21 for correlation analysis.
ACKNOWLEDGMENTS
All members of the Dutt laboratory for critically reviewing the manuscript. Sandor Proteomics Pvt. Ltd. and Scigenome Labs, for providing Exome and Transcriptome library preparation services. A.D. is supported by an Intermediate Fellowship from the Wellcome Trust/DBT India Alliance (IA/I/11/2500278), by a grant from DBT (BT/PR2372/AGR/36/696/2011), and intramural grants (IRB project 92 and 55). P.U. is supported by a senior research fellowship from CSIR. E.K. is supported by a senior research fellowship from UGC. P.C. and M.G are supported by a senior research fellowship from ACTREC. N.G is supported by a junior research fellowship from Tata Memorial hospital. The funders had no role in study design, data collection, and analysis, decision to publish, or preparation of the manuscript.
CONFLICTS OF INTEREST
The authors declare no competing financial interests.
FUNDING
This work was supported by Wellcome Trust/DBT India Alliance (IA/I/11/2500278) and intramural grants (IRB project 92 and 55).
REFERENCES
1. Ntziachristos P, Lim JS, Sage J and Aifantis I. From fly wings to targeted cancer therapies: a centennial for notch signaling. Cancer cell. 2014; 25:318-334.
2. Egloff AM and Grandis JR. Molecular pathways: context-dependent approaches to Notch targeting as cancer therapy. Clin Cancer Res. 2012; 18:5188-5195.
3. Izumchenko E, Sun K, Jones S, Brait M, Agrawal N, Koch W, McCord CL, Riley DR, Angiuoli SV, Velculescu VE, Jiang WW and Sidransky D. Notch1 mutations are drivers of oral tumorigenesis. Cancer Prev Res (Phila). 2015; 8:277-286.
4. Sun W, Gaykalova DA, Ochs MF, Mambo E, Arnaoutakis D, Liu Y, Loyo M, Agrawal N, Howard J, Li R, Ahn S, Fertig E, Sidransky D, et al. Activation of the NOTCH pathway in head and neck cancer. Cancer research. 2014; 74:1091-1104.
5. Pickering CR, Zhang J, Yoo SY, Bengtsson L, Moorthy S, Neskey DM, Zhao M, Ortega Alves MV, Chang K, Drummond J, Cortez E, Xie TX, Zhang D, et al. Integrative genomic characterization of oral squamous cell carcinoma identifies frequent somatic drivers. Cancer Discov. 2013; 3:770-781.
6. Rettig EM, Chung CH, Bishop JA, Howard JD, Sharma R, Li RJ, Douville C, Karchin R, Izumchenko E, Sidransky D, Koch W, Califano J, Agrawal N, et al. Cleaved NOTCH1 Expression Pattern in Head and Neck Squamous Cell Carcinoma Is Associated with NOTCH1 Mutation, HPV Status, and High-Risk Features. Cancer Prev Res (Phila). 2015; 8:287-295.
7. Stransky N, Egloff AM, Tward AD, Kostic AD, Cibulskis K, Sivachenko A, Kryukov GV, Lawrence MS, Sougnez C, McKenna A, Shefler E, Ramos AH, Stojanov P, et al. The mutational landscape of head and neck squamous cell carcinoma. Science. 2011; 333:1157-1160.
8. Agrawal N, Frederick MJ, Pickering CR, Bettegowda C, Chang K, Li RJ, Fakhry C, Xie TX, Zhang J, Wang J, Zhang N, El-Naggar AK, Jasser SA, et al. Exome sequencing of head and neck squamous cell carcinoma reveals inactivating mutations in NOTCH1. Science. 2011; 333:1154-1157.
9. Jemal A, Bray F, Center MM, Ferlay J, Ward E and Forman D. Global cancer statistics. CA: a cancer journal for clinicians. 2011; 61:69-90.
10. Rothenberg SM and Ellisen LW. The molecular pathogenesis of head and neck squamous cell carcinoma. The Journal of clinical investigation. 2012; 122:1951-1957.
11. Suman S, Das TP and Damodaran C. Silencing NOTCH signaling causes growth arrest in both breast cancer stem cells and breast cancer cells. Br J Cancer. 109:2587-2596.
12. Wang J, Sullenger BA and Rich JN. Notch signaling in cancer stem cells. Advances in experimental medicine and biology. 2012; 727:174-185.
13. Shrivastava S, Steele R, Sowadski M, Crawford SE, Varvares M and Ray RB. Identification of molecular signature of head and neck cancer stem-like cells. Scientific reports. 2015; 5:7819.
14. Zhao ZL, Zhang L, Huang CF, Ma SR, Bu LL, Liu JF, Yu GT, Liu B, Gutkind JS, Kulkarni AB, Zhang WF and Sun ZJ. NOTCH1 inhibition enhances the efficacy of conventional chemotherapeutic agents by targeting head neck cancer stem cell. Scientific reports. 2016; 6:24704.
15. Lee SH, Do SI, Lee HJ, Kang HJ, Koo BS and Lim YC. Notch1 signaling contributes to stemness in head and neck squamous cell carcinoma. Lab Invest. 2016; 96:508-516.
16. Chandrani P, Kulkarni V, Iyer P, Upadhyay P, Chaubal R, Das P, Mulherkar R, Singh R and Dutt A. NGS-based approach to determine the presence of HPV and their sites of integration in human cancer genome. British journal of cancer. 2015; 112:1958-1965.
17. Laprise C, Madathil SA, Allison P, Abraham P, Raghavendran A, Shahul HP, ThekkePurakkal AS, Castonguay G, Coutlee F, Schlecht NF, Rousseau MC, Franco EL and Nicolau B. No role for human papillomavirus infection in oral cancers in a region in southern India. Int J Cancer. 2015.
18. Patel KR, Vajaria BN, Begum R, Desai A, Patel JB, Shah FD, Shukla SN and Patel PS. Prevalence of high-risk human papillomavirus type 16 and 18 in oral and cervical cancers in population from Gujarat, West India. Journal of oral pathology & medicine. 2014; 43:293-297.
19. Pathare SM, Gerstung M, Beerenwinkel N, Schaffer AA, Kannan S, Pai P, Pathak KA, Borges AM and Mahimkar MB. Clinicopathological and prognostic implications of genetic alterations in oral cancers. Oncology letters. 2011; 2:445-451.
20. Mutational landscape of gingivo-buccal oral squamous cell carcinoma reveals new recurrently-mutated genes and molecular subgroups. Nat Commun. 2013; 4:2873.
21. Vettore AL, Ramnarayanan K, Poore G, Lim K, Ong CK, Huang KK, Leong HS, Chong FT, Lim TK, Lim WK, Cutcutache I, McPherson JR, Suzuki Y, et al. Mutational landscapes of tongue carcinoma reveal recurrent mutations in genes of therapeutic and prognostic relevance. Genome medicine. 2015; 7:98.
22. Izumchenko E, Sun K, Jones S, Brait M, Agrawal N, Koch W, McCord CL, Riley DR, Angiuoli SV, Velculescu VE, Jiang WW and Sidransky D. Notch1 mutations are drivers of oral tumorigenesis. Cancer prevention research. 2015; 8:277-286.
23. Pratik Chandrani PU, Prajish Iyer, Mayur Tanna, Madhur Shetty, Gorantala Venkata Raghuram, Ninad Oak, Ankita Singh, Rohan Chaubal and Manoj Ramteke SG, Amit Dutt. Integrated genomics approach to identify biologically relevant alterations in fewer samples. BMC genomics. 2015; 16:936.
24. Takebe N, Harris PJ, Warren RQ and Ivy SP. Targeting cancer stem cells by inhibiting Wnt, Notch, and Hedgehog pathways. Nat Rev Clin Oncol. 2011; 8:97-106.
25. Chandrani P, Kulkarni V, Iyer P, Upadhyay P, Chaubal R, Das P, Mulherkar R, Singh R and Dutt A. NGS-based approach to determine the presence of HPV and their sites of integration in human cancer genome. British journal of cancer. 2015; 112:1958-1965.
26. Kagawa S, Natsuizaka M, Whelan KA, Facompre N, Naganuma S, Ohashi S, Kinugasa H, Egloff AM, Basu D, Gimotty PA, Klein-Szanto AJ, Bass AJ, Wong KK, et al. Cellular senescence checkpoint function determines differential Notch1-dependent oncogenic and tumor-suppressor activities. Oncogene. 2015; 34:2347-2359.
27. Lefort K, Mandinova A, Ostano P, Kolev V, Calpini V, Kolfschoten I, Devgan V, Lieb J, Raffoul W, Hohl D, Neel V, Garlick J, Chiorino G and Dotto GP. Notch1 is a p53 target gene involved in human keratinocyte tumor suppression through negative regulation of ROCK1/2 and MRCKalpha kinases. Genes Dev. 2007; 21:562-577.
28. Kunnimalaiyaan M, Vaccaro AM, Ndiaye MA and Chen H. Overexpression of the NOTCH1 intracellular domain inhibits cell proliferation and alters the neuroendocrine phenotype of medullary thyroid cancer cells. The Journal of biological chemistry. 2006; 281:39819-39830.
29. Alniaimi AN, Demorest-Hayes K, Alexander VM, Seo S, Yang D and Rose S. Increased Notch1 expression is associated with poor overall survival in patients with ovarian cancer. International journal of gynecological cancer. 2015; 25:208-213.
30. Zhou L, Yu L, Ding G, Chen W, Zheng S and Cao L. Overexpressions of DLL4 and CD105 are Associated with Poor Prognosis of Patients with Pancreatic Ductal Adenocarcinoma. Pathology oncology research : POR. 2015; 21:1141-1147.
31. Huang J, Song H, Liu B, Yu B, Wang R and Chen L. Expression of Notch-1 and its clinical significance in different histological subtypes of human lung adenocarcinoma. Journal of experimental & clinical cancer research : CR. 2013; 32:84.
32. Tilley AE, Harvey BG, Heguy A, Hackett NR, Wang R, O’Connor TP and Crystal RG. Down-regulation of the notch pathway in human airway epithelium in association with smoking and chronic obstructive pulmonary disease. American journal of respiratory and critical care medicine. 2009; 179:457-466.
33. Wang G, Xu Z, Wang R, Al-Hijji M, Salit J, Strulovici-Barel Y, Tilley AE, Mezey JG and Crystal RG. Genes associated with MUC5AC expression in small airway epithelium of human smokers and non-smokers. BMC medical genomics. 2012; 5:21.
34. Ren ZH, Xu JL, Li B, Fan TF, Ji T and Zhang CP. Elective versus therapeutic neck dissection in node-negative oral cancer: Evidence from five randomized controlled trials. Oral Oncol. 2015; 51:976-981.
35. D’Cruz AK, Vaish R, Kapre N, Dandekar M, Gupta S, Hawaldar R, Agarwal JP, Pantvaidya G, Chaukar D, Deshmukh A, Kane S, Arya S, Ghosh-Laskar S, et al. Elective versus Therapeutic Neck Dissection in Node-Negative Oral Cancer. The New England journal of medicine. 2015; 373:521-529.
36. Hijioka H, Setoguchi T, Miyawaki A, Gao H, Ishida T, Komiya S and Nakamura N. Upregulation of Notch pathway molecules in oral squamous cell carcinoma. Int J Oncol. 2010; 36:817-822.
37. Zhang T, Liu H, Liang Y, Liang L, Liao G, Wu J and Huang H. [The expression and significance of the Notch signaling pathway molecules in tongue squamous cell carcinoma]. Hua Xi Kou Qiang Yi Xue Za Zhi. 2013; 31:303-309.
38. Singh SK, Clarke ID, Terasaki M, Bonn VE, Hawkins C, Squire J and Dirks PB. Identification of a cancer stem cell in human brain tumors. Cancer Res. 2003; 63:5821-5828.
39. Yoshida R, Nagata M, Nakayama H, Niimori-Kita K, Hassan W, Tanaka T, Shinohara M and Ito T. The pathological significance of Notch1 in oral squamous cell carcinoma. Lab Invest. 2013; 93:1068-1081.
40. Li H and Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009; 25:1754-1760.
41. Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C, Gabriel S, Meyerson M, Lander ES and Getz G. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nature biotechnology. 2013; 31:213-219.
42. Ramos AH, Lichtenstein L, Gupta M, Lawrence MS, Pugh TJ, Saksena G, Meyerson M and Getz G. Oncotator: cancer variant annotation tool. Human mutation. 2015; 36:E2423-2429.
43. Sherry ST, Ward MH, Kholodov M, Baker J, Phan L, Smigielski EM and Sirotkin K. dbSNP: the NCBI database of genetic variation. Nucleic acids research. 2001; 29:308-311.
44. Forbes SA, Bhamra G, Bamford S, Dawson E, Kok C, Clements J, Menzies A, Teague JW, Futreal PA and Stratton MR. The Catalogue of Somatic Mutations in Cancer (COSMIC). Current protocols in human genetics / editorial board, Jonathan L Haines [et al]. 2008; Chapter 10:Unit 10 11.
45. Liu A, Yu X and Liu S. Pluripotency transcription factors and cancer stem cells: small genes make a big difference. Chinese journal of cancer. 2013; 32:483-487.
46. Mao Y, Chen H, Liang H, Meric-Bernstam F, Mills GB and Chen K. CanDrA: cancer-specific driver missense mutation annotation with optimized features. PloS one. 2013; 8:e77945.
47. Choi Y and Chan AP. PROVEAN web server: a tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics. 2015; 31:2745-2747.
48. Thorvaldsdottir H, Robinson JT and Mesirov JP. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform. 2013; 14:178-192.
49. Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G and Mesirov JP. Integrative genomics viewer. Nature biotechnology. 2011; 29:24-26.
50. Boeva V, Popova T, Bleakley K, Chiche P, Cappo J, Schleiermacher G, Janoueix-Lerosey I, Delattre O and Barillot E. Control-FREEC: a tool for assessing copy number and allelic content using next-generation sequencing data. Bioinformatics. 2012; 28:423-425.
51. Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL and Pachter L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc. 2012; 7:562-578.
52. Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R and Salzberg SL. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013; 14:R36.
53. Barbieri CE, Baca SC, Lawrence MS, Demichelis F, Blattner M, Theurillat JP, White TA, Stojanov P, Van Allen E, Stransky N, Nickerson E, Chae SS, Boysen G, et al. Exome sequencing identifies recurrent SPOP, FOXA1 and MED12 mutations in prostate cancer. Nature genetics. 2012; 44:685-689.
54. Livak KJ and Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001; 25:402-408.
55. Gioanni J, Fischel JL, Lambert JC, Demard F, Mazeau C, Zanghellini E, Ettore F, Formento P, Chauvel P, Lalanne CM and et al. Two new human tumor cell lines derived from squamous cell carcinomas of the tongue: establishment, characterization and response to cytotoxic treatment. European journal of cancer & clinical oncology. 1988; 24:1445-1455.
56. Chang SE, Foster S, Betts D and Marnock WE. DOK, a cell line established from human dysplastic oral mucosa, shows a partially transformed non-malignant phenotype. Int J Cancer. 1992; 52:896-902.
57. Jin G, Zhang F, Chan KM, Xavier Wong HL, Liu B, Cheah KS, Liu X, Mauch C, Liu D and Zhou Z. MT1-MMP cleaves Dll1 to negatively regulate Notch signalling to maintain normal B-cell development. The EMBO journal. 2011; 30:2281-2293.
58. Castel D, Mourikis P, Bartels SJ, Brinkman AB, Tajbakhsh S and Stunnenberg HG. Dynamic binding of RBPJ is determined by Notch signaling status. Genes Dev. 2013; 27:1059-1071.
59. Walker JM. The bicinchoninic acid (BCA) assay for protein quantitation. Methods Mol Biol. 1994; 32:5-8.
60. Dinse GE and Lagakos SW. Nonparametric estimation of lifetime and disease onset distributions from incomplete observations. Biometrics. 1982; 38:921-932.

