INTRODUCTION
Osteosarcoma, one of the most common types of malignant primary bone tumors, is characterised by the direct formation of immature bone or osteoid tissue by tumor cells, and is most prevalent in children and young people [1, 2]. In the 1970s, adjuvant chemotherapy for osteosarcoma was introduced to supplement standard treatments such as surgery and radiotherapy, and resulted in a high five-year survival rate [3, 4]. However, the survival of patients with initial pulmonary metastases and recurrent disease remained low [5]. Other side effects, such as bone disability, physical dysfunction and drug toxicity of chemotherapy, seriously reduce patients’ quality of life, bringing a heavy medical burden to their families and the society [6]. To date, the pathogenesis of osteosarcoma remains unclear.
Recent molecular studies have suggested that genome dysfunction contributes to tumorigenesis [7, 8]. Mutations and abnormal expression of certain genes, such as those encoding microRNA, interleukin family proteins and cytochrome P450 superfamily members, facilitate the development of tumors and other malignant diseases [9–12]. Angiogenesis is a critical process to promote tumor cell proliferation and metastasis through the formation of new capillaries [13, 14]. Vascular endothelial growth factor (VEGF) is an important stimulator of physiological and pathological angiogenesis that acts on vascular endothelial cells and promotes human blood vessel growth during tumor formation and growth, enabling invasion and metastasis [15–17]. Increased VEGF expression has been detected in tumor tissue, blood and urine samples from patients with esophageal, prostate, lung, and other cancers.[18–21].
VEGF is located at chromosome 6p21.3 and comprises a 14-kb coding region with eight exons and seven introns [22, 23]. Several single-nucleotide polymorphisms (SNPs) have been described for the VEGF gene [24, 25]. Some of these polymorphisms, including those in the promoter and 5’-untranslated region, have been shown to alter the expression and biological activity of VEGF [26]. In 2014, Wang et al reported the first case-control study showing that +936 C/T was associated with a significant increase in osteosarcoma risk [27]. To date, the three most common VEGF polymorphisms, namely +936C/T (rs3025039), +1612G/A (rs10434), and –634G/C (rs2010963), have been investigated for their association with osteosarcoma risk, but the results have been inconsistent. Therefore, we conducted a comprehensive meta-analysis of all published studies to determine the association between VEGF polymorphism and osteosarcoma susceptibility more accurately. Our meta-analysis was performed according to the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) statement [28, 29]. No ethical issues were involved in this study, given that our data were based on published studies.
RESULTS
Study characteristics
Forty studies were identified through literature searching. In accordance with the selection criteria described in the Methods, 23 studies were excluded for duplication in the first step of title screening, and 17 studies were excluded for other reasons (two were review articles, five were not case-control studies, and eight were fundamental biology studies) during the systematic screening. Ultimately, seven articles were included in this meta-analysis (Figure 1, Table 1), all of which involved Chinese populations [27, 30–35]. For the +936C/T polymorphism, the seven studies included a total of 1,350 cases and 1,706 controls [27, 30–35]. For the +1612 G/A polymorphism, the six studies included a total of 1,166 cases and 1,524 controls [27, 30–32, 34, 35]. For the –634 G/C polymorphism, the six studies involved a total of 1,166 cases and 1,524 controls [27, 30–32, 34, 35]. All the included case-control studies used the polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) method. In three studies of the +936C/T polymorphism, the genotype distribution of the controls deviated from Hardy-Weinberg equilibrium (HWE) [30, 33, 34].
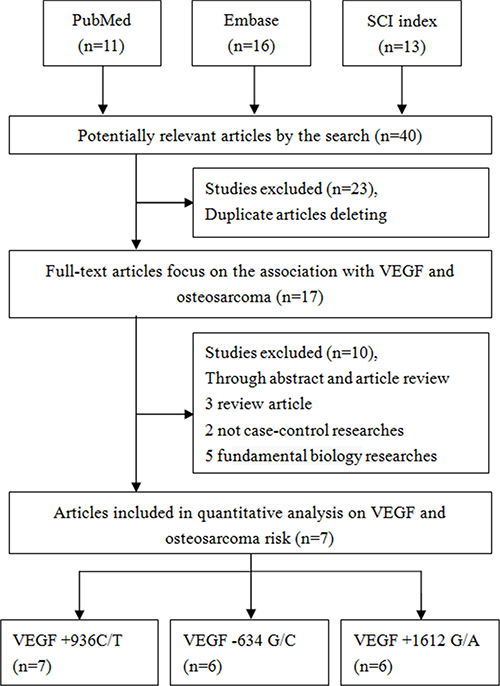
Figure 1: Flow diagram of the study selection process.
Table 1: Characteristics of case-control studies on VEGF polymorphisms and osteosarcoma risk included in the meta-analysis
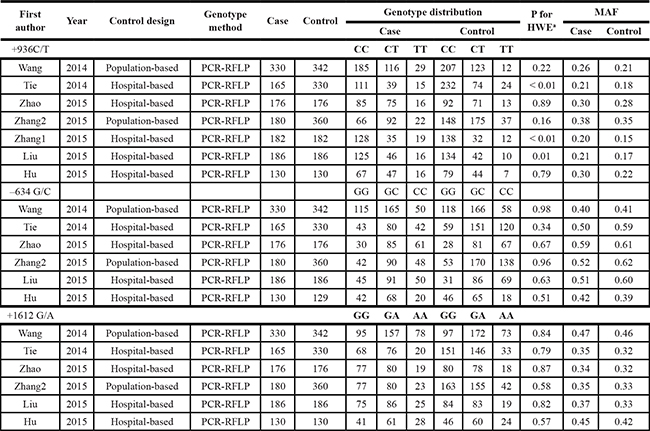
aHWE in control.
MAF: Minor allele frequency.
Quantitative analysis
For the +936C/T polymorphism
In the seven studies examined, a significant increase in osteosarcoma risk was observed in four genetic models (T vs. C: Odds Ratio [OR] = 1.26, 95% Confidence Interval [CI] = 1.12–1.42, P < 0.01, I2 = 0%; TT vs. CC: OR = 1.70, 95% CI = 1.29–2.24, P < 0.01, I2 = 0%; CT+TT vs. CC: OR = 1.23, 95% CI = 1.06–1.44, P < 0.01, I2 = 0% [Figure 2A], TT vs. CC+CT: OR = 1.61, 95% CI = 1.23–2.10, P < 0.01, I2 = 0%) (Table 2). Similarly increased risk also detected in the subgroup analysis of the control from hospital population (T vs. C: OR = 1.28, 95% CI = 1.09–1.50, P < 0.01, I2 = 0%; TT vs. CC: OR = 1.62, 95% CI = 1.15–2.30, P < 0.01, I2 = 0%; CT+TT vs. CC: OR = 1.26, 95% CI = 1.03–1.53, P = 0.02, I2 = 0%, TT vs. CC+CT: OR = 1.55, 95% CI = 1.10–2.19, P = 0.01, I2 = 0%). Sensitivity analyses were performed by excluding each single study one by one, and the results of pooled ORs still presented stability (Figure 2B for the dominant model). Moreover, cumulative analysis with publication date demonstrated that the osteosarcoma risk was increasing gradually and presented positive with the report by Zhang et al. in 2015 (Figure 2C for the dominant model). Funnel plot symmetry was performed to estimate publication bias, and no evidence of asymmetry was obtained (Figure 2D for the dominant model). The results were further validated by Egger’s test (T vs. C, P = 0.26; TT vs. CC, P = 0.28; CT+TT vs. CC, P = 0.11; TT vs. CC+CT, P = 0.24).
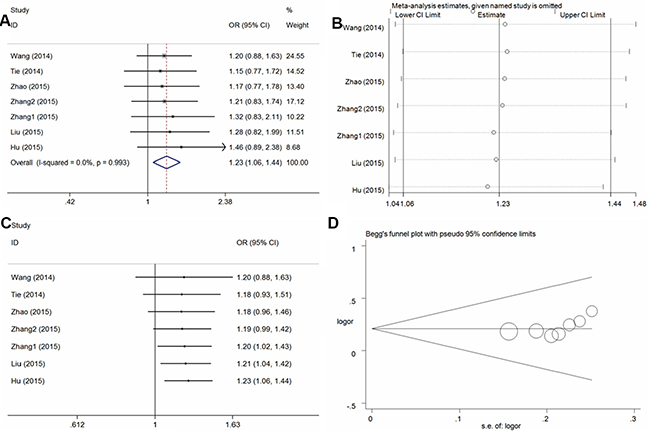
Figure 2: Statistical analysis of the association between the VEGF +936C/T polymorphism and osteosarcoma risk in the CT + TT vs. CC model. (A) ORs and 95% CIs; (B) sensitivity analysis; (C) cumulative analysis; (D) publication bias.
Table 2: Summary ORs and 95% CI of VEGF polymorphisms and osteosarcoma risk
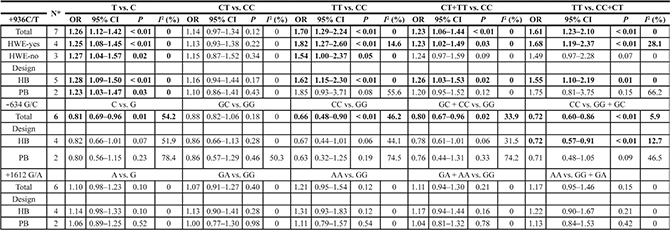
*Numbers of comparisons.
I2 for heterogeneity test.
For the –634 G/C polymorphism
In the six studies examined, overall, significant protective effects of the –634 G/C polymorphism against osteosarcoma risk were found in the four genetic models (C vs. G: OR = 0.81, 95% CI = 0.69–0.96, P = 0.01, I2 = 54.2%; CC vs. GG: OR = 0.66, 95% CI = 0.48–0.90, P < 0.01, I2 = 46.2%; GC+CC vs. GG: OR = 0.80, 95% CI = 0.67–0.96, P = 0.02, I2 = 33.9% (Figure 3A for the dominant model); CC vs. GG+GC: OR = 0.72, 95%CI = 0.60-0.86, P < 0.01, I2 = 5.9%) (Table 2). Sensitivity analysis (Figure 3B for the dominant model) was conducted and the results were stable on the whole. Cumulative analysis (Figure 3C for dominant model) demonstrated a significant protective association from the study of Zhang et al. in 2015 [32]. No publication bias was found in the funnel plot and Egger’s test (C vs. G, P = 0.80; CC vs. GG, P = 0.55; GC+CC vs. GG, P = 0.61; CC vs. GG+GC, P = 0.19) (Figure 3D for the dominant model).
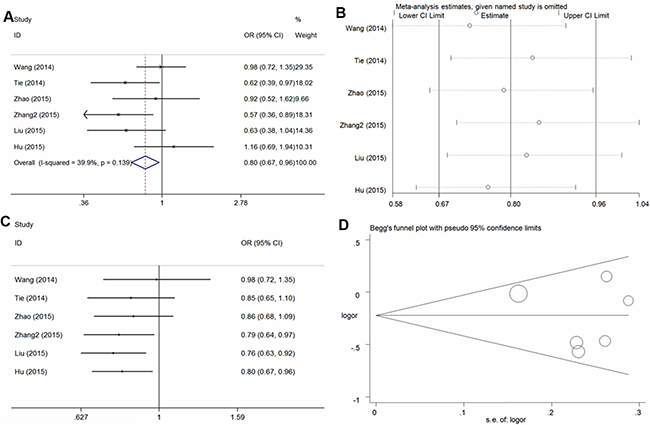
Figure 3: Statistical analysis of the association between the VEGF – 634G/C polymorphism and osteosarcoma risk in the GC + CC vs. GG model. (A) ORs and 95% CIs; (B) sensitivity analysis; (C) cumulative analysis; (D) publication bias.
For the +1612 G/A polymorphism
In the six studies included, no significant association with osteosarcoma risk was found in any model in the total population(A vs. G: OR = 1.10, 95% CI = 0.98–1.23, = 0.10, I2 = 0%; GA vs. GG: OR = 1.07, 95% CI = 0.91–1.27, = 0.40, I2 = 0%; AA vs. GG: OR = 1.21, 95% CI = 0.95–1.54, P = 0.12, I2 = 0%; GA + AA vs. GG: OR = 1.11, 95% CI = 0.94–1.30, P = 0.21, I2 = 0% [Figure 4A for the dominant model]; AA vs. GG + GA: OR = 1.17, 95% CI = 0.95–1.46, P = 0.15, I2 = 0%) and subsequent subgroup analysis (Table 2). Sensitivity analysis (Figure 4B for dominant model) and cumulative analysis (Figure 4C for the dominant model) were conducted, and no substantial changes in the ORs were observed. Furthermore, no publication bias was detected, suggesting that the results are statistically robust (A vs. G, P = 0.17; GA vs. GG, P = 0.25; AA vs. GG, P = 0.19; GA + AA vs. GG, P = 0.21; AA vs. GG + GA, P = 0.54) (Figure 4D for the dominant model).
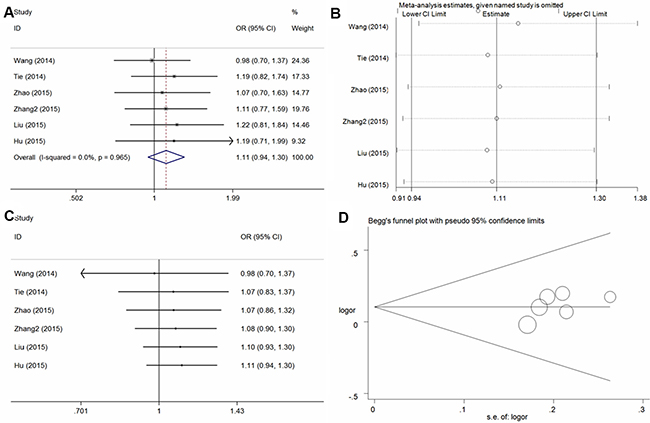
Figure 4: Statistical analysis of the association between the VEGF +1612G/A polymorphism and osteosarcoma risk in the GA + AA vs. GG model. (A) ORs and 95% CIs; (B) sensitivity analysis; (C) cumulative analysis; (D) publication bias.
DISCUSSION
Angiogenesis is a critical cause of proliferation, invasion, and metastasis in cancer [36]. VEGF is one of the most important cytokines in angiogenesis, which can promote the mitosis of vascular endothelial cells and accelerate the formation of new blood vessels [37]. High expression of VEGF in the primary tumor has been found in certain types of malignant tumors with high metastatic rates and poor prognoses, such as breast, esophageal and colorectal cancers [38, 39].
Studies of osteosarcoma have strongly suggested that higher VEGF expression and activity in primary tumor tissue correlates with increased local microvessel density, faster development of pulmonary metastasis, and poorer prognosis for osteosarcoma patients [40, 41]. An increasing number of studies are focusing on the associations among gene mutations (particularly SNPs), protein expression/activity, and tumor occurrence. Wang et al. [27] conducted the first case-control study of the VEGF gene and reported that the +936C/T polymorphism was associated with increased osteosarcoma risk in a Chinese population (T vs. C: OR = 1.31, 95% CI = 1.02–1.68, P = 0.04; TT vs. CC: OR = 2.70, 95% CI = 1.34–5.45, P < 0.01). Since then, a series of case-control studies have been conducted, but the conclusions have been inconsistent. In 2016, Zhang et al. [42] conducted a meta-analysis and observed that VEGF –634 G/C and +936 C/T polymorphisms were significantly associated with osteosarcoma risk. However, their meta-analysis only included three case-control studies. To date, four additional studies have explored the associations between VEGF polymorphisms and osteosarcoma. In our meta-analysis of seven eligible case-control studies, more precise associations between the three most common polymorphisms in VEGF (+936 C/T, –634 G/C, +1612 G/A) and osteosarcoma susceptibility were investigated than in the previous meta-analysis. All the results revealed that the VEGF +936 C/T polymorphism significantly increased the risk of osteosarcoma in the Chinese population. Subgroup analyses based on the HWE status and control design also identified similar risks. Interestingly, our results suggested that the VEGF –634 G/C polymorphism protects against osteosarcoma development. However, no significant association between VEGF +1612 G/A and osteosarcoma susceptibility was observed. These inconsistent findings among the three SNP loci indicat that different polymorphisms exert different effects on gene function, even when they are located at the same unit.
Meta-analysis is an effective method of combining the quantitative results of previous studies in order to derive a pooled summary conclusion through statistical measures [43, 44]. This can reduce the risk of drawing incorrect conclusions based on insufficient methods or small sample sizes. We conducted this meta-analysis to comprehensively investigate the relationships of VEGF polymorphisms (+936C/T, – 634 G/C, and +1612 G/A) with osteosarcoma susceptibility. None of the studies evaluated herein displayed significant heterogeneity or publication bias within the three common polymorphism loci. All the pooled data yielded consistent results, which not only confirms the validity of our results, but also supports our statistical methods.
However, there were some limitations to our meta-analysis. First, only seven eligible studies were collected, and the limited number of studies with small sample sizes may have affected the analysis of the correlation between VEGF polymorphisms and osteosarcoma susceptibility. Second, all the included studies were conducted with Chinese subjects. Ethnicity bias may exist because a single Asian race was examined, and the conclusions may not be applicable to other races. Third, the interactions among various risk factors, such as smoking, drinking, and other genetic factors, are crucial determinants of cancer formation. Thus, the inherent mechanism could not be explained clearly with unadjusted databases in this meta-analysis. Fourth, deviations from HWE were found in the distribution of controls in some of the included studies, possibly due to the insufficient of sample sizes or genotyping errors in those studies.
Our meta-analysis contained a large sample size, new research data and high-quality statistics. The results are credible and reliable as a preliminary exploration of the relationship between the VEGF polymorphisms and osteosarcoma susceptibility, and suggest that analysis of VEGF polymorphisms may be useful for the early clinical diagnosis and treatment of osteosarcoma.
In summary, our meta-analysis suggests that VEGF polymorphisms are associated with osteosarcoma susceptibility in the Chinese population. Nevertheless, additional studies including different ethnicities and larger populations are needed for further exploration of these associations.
MATERIALS AND METHODS
Search strategy
Three online electronic databases (PubMed, Embase, and Science Citation Index) were searched with the following terms: (“osteosarcoma”) AND (“vascular endothelial growth factor” OR “VEGF”) AND (“polymorphism” OR “SNP” OR “single nucleotide polymorphism” OR “variant”), from inception to January 2016. Furthermore, all the references of the collected studies were examined so that additional relevant studies could be identified. Only English language full-text case-controls studies were included.
Study selection
All the selected studies met the following inclusion criteria: (1) case-control studies focused on osteosarcoma; (2) reported on VEGF polymorphisms; (3) evaluated the association between VEGF polymorphisms and osteosarcoma risk; (4) presented adequate genotype data or data necessary to calculate the OR and 95% CI; and (5) described SNP loci which were reported in at least five publications. Studies were excluded if they: (1) were review papers; (2) were animal studies; (3) contained insufficient information for assessment of the association; (4) contained repeated or overlapping results from another study (in such cases, only the study with the largest sample size or the most recent study was included).
Data extraction
Two reviewers (Hu and Du) independently conducted data extraction using a standardized form. The following information was collected: the first author’s name, year of publication, sources of controls, study region or country, ethnicity of subjects (such as Asian or Caucasian), genotyping method and genotype distribution data for cases and controls. Disagreements were resolved by a third author during the analysis.
Statistical analysis
HWE was evaluated with the chi-square test using the genotypes of the controls. The association between the genotype distribution and osteosarcoma was evaluated based on the ORs and corresponding 95% CIs. For example, the pooled ORs were obtained for the allele contrast (T vs. C), co-dominant model (CT vs. CC, TT vs. CC), dominant model (CT + TT vs. CC), and recessive model (TT vs. CC + CT) in the VEGF +936C/T locus. Similar genetic models were also assessed for the other SNPs. Heterogeneity was assessed with Cochran’s Q statistic and the I2 method, and P < 0.10 or I2 > 40% were considered to demonstrate substantial heterogeneity [45]. ORs were estimated with a fixed-effects model (the Mantel-Haenszel method) when there was no considerable heterogeneity [46]; otherwise, a random-effects model (DerSimonian and Laird method) was adopted [47]. Subgroup analyses were conducted based on the classification of the study design and HWE status. Cumulative meta-analyses were conducted so that any potential trends in the pooled estimates over the study years could be identified. Sensitivity analyses were conducted, in which the stability of the results was evaluated as each study was sequentially removed for each locus. Egger’s linear regression and Begg’s funnel plots were used to assess potential publication bias. Statistical analysis was performed with STATA version 12.0 (Stata Corporation, College Station, TX, USA) with two-sided P values, and P < 0.05 was considered statistically significant.
ACKNOWLEDGMENTS AND FUNDING
This study was supported by grants from the Foundation of Ministry of Education of Hubei Province (D20142102) and Taihe Hospital (2013PY02, 2013JJXM011). The funders had no involvement in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
CONFLICTS OF INTEREST
The authors declare no competing financial interests.
REFERENCES
1. Siclari VA, Qin L. Targeting the osteosarcoma cancer stem cell. J Orthop Surg Res. 2010; 5:78.
2. Yang J, Yang D, Sun Y, Sun B, Wang G, Trent JC, Araujo DM, Chen K, Zhang W. Genetic amplification of the vascular endothelial growth factor (VEGF) pathway genes, including VEGFA, in human osteosarcoma. Cancer. 2011; 117:4925–4938.
3. Sutow WW, Sullivan MP, Fernbach DJ, Cangir A, George SL. Adjuvant chemotherapy in primary treatment of osteogenic sarcoma. A Southwest Oncology Group study. Cancer. 1975; 36:1598–1602.
4. Harvei S, Solheim O. The prognosis in osteosarcoma: Norwegian National Data. Cancer. 1981; 48:1719–1723.
5. Fagioli F, Aglietta M, Tienghi A, Ferrari S, Brach del Prever A, Vassallo E, Palmero A, Biasin E, Bacci G, Picci P, Madon E. High-dose chemotherapy in the treatment of relapsed osteosarcoma: an Italian sarcoma group study. J Clin Oncol. 2002; 20:2150–2156.
6. Fan Z, Patel S, Lewis VO, Guadagnolo BA, Lin PP. Should High-grade Extraosseous Osteosarcoma Be Treated With Multimodality Therapy Like Other Soft Tissue Sarcomas? Clin Orthop Relat Res. 2015; 473:3604–3611.
7. Si W, Huang W, Zheng Y, Yang Y, Liu X, Shan L, Zhou X, Wang Y, Su D, Gao J, Yan R, Han X, Li W, et al. Dysfunction of the Reciprocal Feedback Loop between GATA3- and ZEB2-Nucleated Repression Programs Contributes to Breast Cancer Metastasis. Cancer Cell. 2015; 27:822–836.
8. Niu YM, Du XY, Cai HX, Zhang C, Yuan RX, Zeng XT, Luo J. Increased risks between Interleukin-10 gene polymorphisms and haplotype and head and neck cancer: a meta-analysis. Sci Rep. 2015; 5:17149.
9. Niu YM, Du XY, Lu MY, Xu QL, Luo J, Shen M. Significant association between functional microRNA polymorphisms and head and neck cancer susceptibility: a comprehensive meta-analysis. Sci Rep. 2015; 5:12972.
10. Gu Y, Yu C, Miao L, Wang L, Xu C, Xue W, Du J, Yuan H, Dai J, Jin G, Hu Z, Ma H, Shen H. Telomere length, genetic variants and risk of squamous cell carcinoma of the head and neck in Southeast Chinese. Sci Rep. 2016; 6:20675.
11. Zhang Y, Zhu L, Wang R, Miao L, Jiang H, Yuan H, Ma H, Chen N. Genetic variants in let-7/Lin28 modulate the risk of oral cavity cancer in a Chinese Han population. Sci Rep. 2014; 4:7434.
12. Niu YM, Yuan H, Zhou Y. Interleukin-17 gene polymorphisms contribute to cancer risk. Mediators Inflamm. 2014; 2014:128490.
13. Carmeliet P, Jain RK. Molecular mechanisms and clinical applications of angiogenesis. Nature. 2011; 473:298–307.
14. Carmeliet P, Jain RK. Angiogenesis in cancer and other diseases. Nature. 2000; 407:249–257.
15. Kumar B, Chile SA, Ray KB, Reddy GE, Addepalli MK, Kumar AS, Ramana V, Rajagopal V. VEGF-C differentially regulates VEGF-A expression in ocular and cancer cells; promotes angiogenesis via RhoA mediated pathway. Angiogenesis. 2011; 14:371–380.
16. Carmeliet P. VEGF as a key mediator of angiogenesis in cancer. Oncology. 2005; 69:4–10.
17. McMahon G. VEGF receptor signaling in tumor angiogenesis. Oncologist. 2000; 5:3–10.
18. Adams J, Carder PJ, Downey S, Forbes MA, MacLennan K, Allgar V, Kaufman S, Hallam S, Bicknell R, Walker JJ, Cairnduff F, Selby PJ, Perren TJ, et al. Vascular endothelial growth factor (VEGF) in breast cancer: comparison of plasma, serum, and tissue VEGF and microvessel density and effects of tamoxifen. Cancer Res. 2000; 60:2898–2905.
19. Schwaederle M, Lazar V, Validire P, Hansson J, Lacroix L, Soria JC, Pawitan Y, Kurzrock R. VEGF-A Expression Correlates with TP53 Mutations in Non-Small Cell Lung Cancer: Implications for Antiangiogenesis Therapy. Cancer Res. 2015; 75:1187–1190.
20. Green MM, Hiley CT, Shanks JH, Bottomley IC, West CM, Cowan RA, Stratford IJ. Expression of vascular endothelial growth factor (VEGF) in locally invasive prostate cancer is prognostic for radiotherapy outcome. Int J Radiat Oncol Biol Phys. 2007; 67:84–90.
21. Mukherjee T, Kumar A, Mathur M, Chattopadhyay TK, Ralhan R. Ets-1 and VEGF expression correlates with tumor angiogenesis, lymph node metastasis, and patient survival in esophageal squamous cell carcinoma. J Cancer Res Clin Oncol. 2003; 129:430–436.
22. Watson CJ, Webb NJ, Bottomley MJ, Brenchley PE. Identification of polymorphisms within the vascular endothelial growth factor (VEGF) gene: correlation with variation in VEGF protein production. Cytokine. 2000; 12:1232–1235.
23. Vincenti V, Cassano C, Rocchi M, Persico G. Assignment of the vascular endothelial growth factor gene to human chromosome 6p21.3. Circulation. 1996; 93:1493–1495.
24. Krippl P, Langsenlehner U, Renner W, Yazdani-Biuki B, Wolf G, Wascher TC, Paulweber B, Haas J, Samonigg H. A common 936 C/T gene polymorphism of vascular endothelial growth factor is associated with decreased breast cancer risk. Int J Cancer. 2003; 106:468–471.
25. Stevens A, Soden J, Brenchley PE, Ralph S, Ray DW. Haplotype analysis of the polymorphic human vascular endothelial growth factor gene promoter. Cancer Res. 2003; 63:812–816.
26. Awata T, Inoue K, Kurihara S, Ohkubo T, Watanabe M, Inukai K, Inoue I, Katayama S. A common polymorphism in the 5′-untranslated region of the VEGF gene is associated with diabetic retinopathy in type 2 diabetes. Diabetes. 2002; 51:1635–1639.
27. Wang Z, Wen P, Luo X, Fang X, Wang Q, Ma F, Lv J. Association of the vascular endothelial growth factor (VEGF) gene single-nucleotide polymorphisms with osteosarcoma susceptibility in a Chinese population. Tumour Biol. 2014; 35:3605–3610.
28. Zeng X, Zhang Y, Kwong JS, Zhang C, Li S, Sun F, Niu Y, Du L. The methodological quality assessment tools for preclinical and clinical studies, systematic review and meta-analysis, and clinical practice guideline: a systematic review. J Evid Based Med. 2015; 8:2–10.
29. Moher D, Liberati A, Tetzlaff J, Altman DG. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. Ann Intern Med. 2009; 151:264–269, W264.
30. Tie Z, Bai R, Zhai Z, Zhang G, Zhang H, Zhao Z, Zhou D, Liu W. Single nucleotide polymorphisms in VEGF gene are associated with an increased risk of osteosarcoma. Int J Clin Exp Pathol. 2014; 7:8143–8149.
31. Li-Lian Z, Lin W, Lei S, Yao-Nan Z. Investigation on the role of VEGF gene polymorphisms in the risk of osteosarcoma. Pak J Med Sci. 2015; 31:364–368.
32. Zhang G, Bai R, Zhang T, Zhang H, Wen SZ, Jiang DM. Investigation of the role of VEGF gene polymorphisms in the risk of osteosarcoma. Genet Mol Res. 2015; 14:8283–8289.
33. Zhang HF, Yan JP, Zhuang YS, Han GQ. Association between angiogenic growth factor genetic polymorphisms and the risk of osteosarcoma. Genet Mol Res. 2015; 14:10524–10529.
34. Liu JQ, Bai X, Duan DC, Dou AX. Role of five small nucleotide polymorphisms in the VEGF gene on the susceptibility to osteosarcoma and overall survival of patients. Oncol Lett. 2015; 10:1481–1486.
35. Hu GL, Ma G, Ming JH. Impact of common SNPs in VEGF gene on the susceptibility of osteosarcoma. Genet Mol Res. 2015; 14:14561–14566.
36. Zetter BR. Angiogenesis and tumor metastasis. Annu Rev Med. 1998; 49:407–424.
37. Dvorak HF. Vascular permeability factor/vascular endothelial growth factor: a critical cytokine in tumor angiogenesis and a potential target for diagnosis and therapy. J Clin Oncol. 2002; 20:4368–4380.
38. Yang XR, Xu Y, Yu B, Zhou J, Qiu SJ, Shi GM, Zhang BH, Wu WZ, Shi YH, Wu B, Yang GH, Ji Y, Fan J. High expression levels of putative hepatic stem/progenitor cell biomarkers related to tumour angiogenesis and poor prognosis of hepatocellular carcinoma. Gut. 2010; 59:953–962.
39. Seo Y, Baba H, Fukuda T, Takashima M, Sugimachi K. High expression of vascular endothelial growth factor is associated with liver metastasis and a poor prognosis for patients with ductal pancreatic adenocarcinoma. Cancer. 2000; 88:2239–2245.
40. Kaya M, Wada T, Akatsuka T, Kawaguchi S, Nagoya S, Shindoh M, Higashino F, Mezawa F, Okada F, Ishii S. Vascular endothelial growth factor expression in untreated osteosarcoma is predictive of pulmonary metastasis and poor prognosis. Clin Cancer Res. 2000; 6:572–577.
41. Lammli J, Fan M, Rosenthal HG, Patni M, Rinehart E, Vergara G, Ablah E, Wooley PH, Lucas G, Yang SY. Expression of Vascular Endothelial Growth Factor correlates with the advance of clinical osteosarcoma. Int Orthop. 2012; 36:2307–2313.
42. Zhang Z, Xiong J, Wang S, Qiu Y. Associations between VEGF polymorphisms and the risk and prognosis of osteosarcoma: a systematic review and meta-analysis. Int J Clin Exp Med. 2016; 9:515–524.
43. Niu YM, Weng H, Zhang C, Yuan RX, Yan JZ, Meng XY, Luo J. Systematic Review by Multivariate Meta-analyses on the Possible Role of Tumor Necrosis Factor-alpha Gene Polymorphisms in Association with Ischemic Stroke. Neuromolecular Med. 2015; 17:373–384.
44. O’Rourke K, Detsky AS. Meta-analysis in medical research: strong encouragement for higher quality in individual research efforts. J Clin Epidemiol. 1989; 42:1021–1024.
45. Huedo-Medina TB, Sanchez-Meca J, Marin-Martinez F, Botella J. Assessing heterogeneity in meta-analysis: Q statistic or I2 index? Psychol Methods. 2006; 11:193–206.
46. Mantel N, Haenszel W. Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst. 1959; 22:719–748.
47. DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986; 7:177–188.

