INTRODUCTION
Human Herpesvirus 6 (HHV-6) exists as two related herpes viruses, HHV-6A and HHV-6B, that infect almost all human beings, especially the children [1, 2]. HHV-6 has a life-long latency and can become reactivated infection later [3]. HHV-6 reactivation has been linked with many clinical appearances throughout the body, including the lungs, kidney, heart, brain, and gastrointestinal tract [4, 5]. HHV-6 can infect various CNS cells in vitro [6-10]. HHV-6 has been involved in the progress of various range of neurologic disorders, including encephalitis, seizures, chronic fatigue syndrome, mesial temporal lobe epilepsy (MTLE), Alzheimer’s disease, and multiple sclerosis [11, 12]. The diverse pathology may due to the viral sequence variations and differences in antigenic specificity between the HHV-6A and HHV-6B [11]. More research is needed to understand the important disease associations that have been suggested.
Recently, mRNA-seq has been increasingly used to explore the genetic and environmental factors of virus infection and diseases occurrence. Here, we undertook a genome-wide survey to map cellular genes of human astrocyte HA1800 that are infected by HHV-6A GS. In this study, we report the identification and comparative analysis of the differentially expressed genes that occurred during the virus infection phenotype conversion process. Therefore, this work is the first attempt at evaluating, genome-wide, the genotype-to-transcriptome-to-clinical phenotype associations in HHV-6A GS infection diseases.
RESULTS
Analysis of differentially expressed genes (DEGs)
The potential DEGs (16430 genes, and 249 genes of FDR < 0.1) between different groups are displayed in Supplementary Table S1. The potential DEGs with FDR < 0.1 (HA1800-control expression > -1 and HA1800-HHV6-GS expression > 0) between libraries are presented in Figure 1A and Supplementary Table S2. Totals of 66 significant DEGs (only 8 genes are downregulated) were identified during the HHV-6A GS virus infection human astrocyte HA1800.
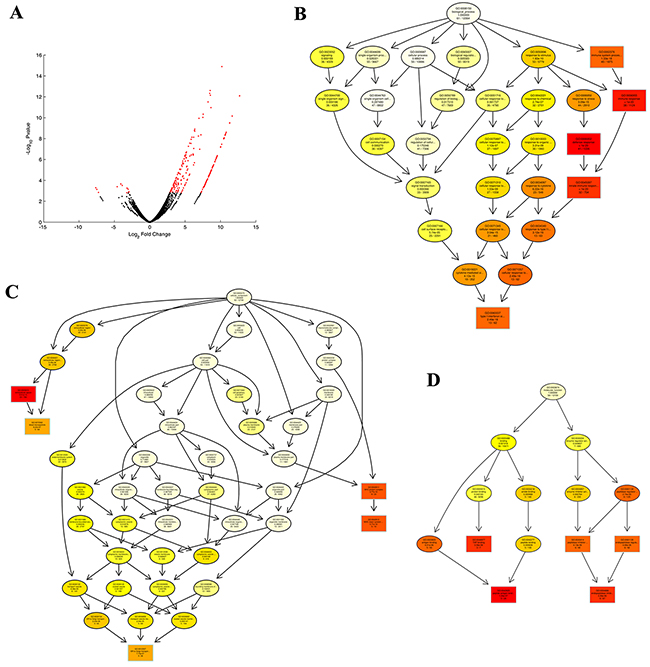
Figure 1: Differentially expressed genes (DEGs) enriched and identified by GO analyses. A. The expression of cellular genes in two enriched populations of HA1800-control and HA1800-HHV6GS cells for 24 h were assessed using mRNA-seq. The distribution of genes with a change in expression of false discovery rate (FDR) < 0.1 is shown in red on the MA plot (log total counts versus log fold-change). B. The biological processes of the DEGs were identified by GO analyses. C. The cellular components of the DEGs were identified by GO analyses. D. The molecular functions of the DEGs were identified by GO analyses.
Gene ontology analysis of potential DEGs
In order to obtain further understanding of the biological functions of the DEGs, Gene Ontology (GO) analyses were performed. We chose significant GO categories listed in Supplementary Table S3, S4, and S5. The biological processes, cellular component, and molecular function are presented in Figure 1B, 1C, and 1D, respectively. And, potential DEGs were enriched for GO categories of defense response (GO:0006952), immune response (GO:0006955), innate immune response (GO:0045087), immune system process (GO:0002376), type I interferon signaling pathway (GO:0060337) in biological processes; extracellular space (GO:0005615), MHC class I protein complex (GO:0042612), blood microparticle (GO:0072562) in cellular component; peptide antigen binding (GO:0042605), endopeptidase inhibitor activity (GO:0004866) in molecular function.
Signaling pathway analysis of potential DEGs
All signaling pathways of DEGs were showed in Figure 2, and the important signaling pathway classes were listed in Supplementary Table S5. According to the results of the KEGG and GO pathway analysis, we focus on the DEGs involved in viral carcinogenesis, viral myocarditis, HTLV-1 infection, Epstein-Barr virus infection, influenza A, herpes simplex infection, TNF signaling pathway, RIG-I-like receptor signaling pathway, NF-kappa B signaling pathway, graft-versus-host disease, complement and coagulation cascades, autoimmune thyroid disease, antigen processing and presentation, allograft rejection, measles and phagosome signaling pathways.
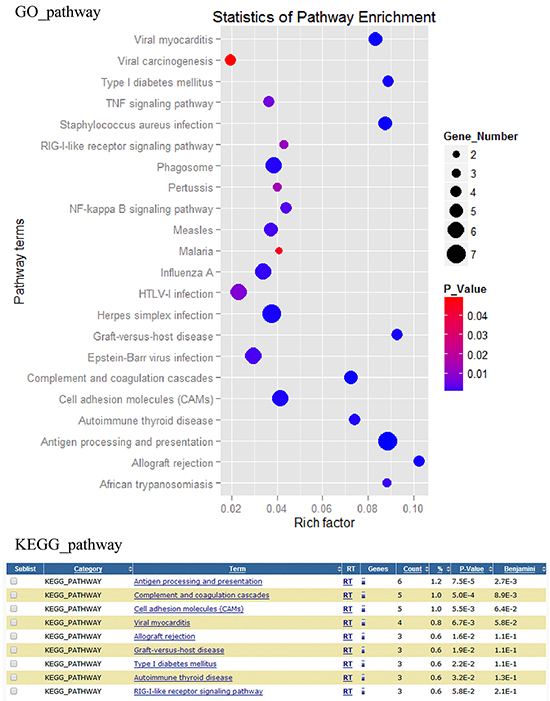
Figure 2: Differentially expressed genes associated pathways analysis. Differentially expressed genes associated pathways were analyzed by GO and KEGG pathway tools.
CNS diseases association analysis of potential DEGs
All disease classes of DEGs were analyzed by functional annotation chart tool (https://david.ncifcrf.gov/home.jsp) [13, 14] in Figure 3 (and Supplementary Figure S1). And the differentially expressed genes involved in infection, immune, neurological, and cardiovascular disease classes. We then further analyzed which of the potential DEGs associated with antivirus, Alzheimer’s Disease, glioma, and multiple sclerosis following HHV-6A GS virus infection human astrocyte HA1800. Of these genes, 12 were associated with antivirus function; 7 were associated with Alzheimer’s Disease; 11 were associated with glioma; 9 were associated with multiple sclerosis (Table 1). And more importantly, CTSS, PTX3, CHI3L1, Mx1, CXCL16, BIRC3, and BST2 genes exhibited significant correlation with more than two CNS diseases. Subsequently, these genes were further recognized by real-time PCR assay in cells at 24 hours and 72 hours (Supplementary Figure S2A and S2B). And the expression of CTSS, Mx1, and BIRC3 genes were further validated by western blot assay at 72hours (Supplementary Figure S2C). The main genes revealing a positive association with viral infection by using the STRING database (i.e., increasing gene expression with viral infection) was shown in Figure 4 (and Supplementary Figure S3). The STRING database (http://string-db.org) designs to supply an important tool for studying protein–protein interactions, including direct and indirect correlations.
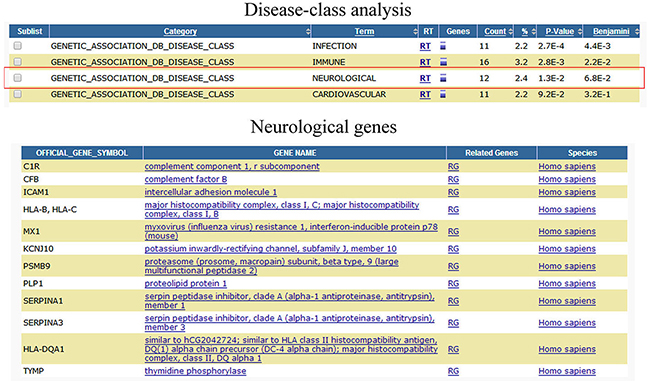
Figure 3: Differentially expressed genes associated CNS diseases analysis. Differentially expressed genes associated CNS diseases were analyzed by DAVID functional annotation chart tool.
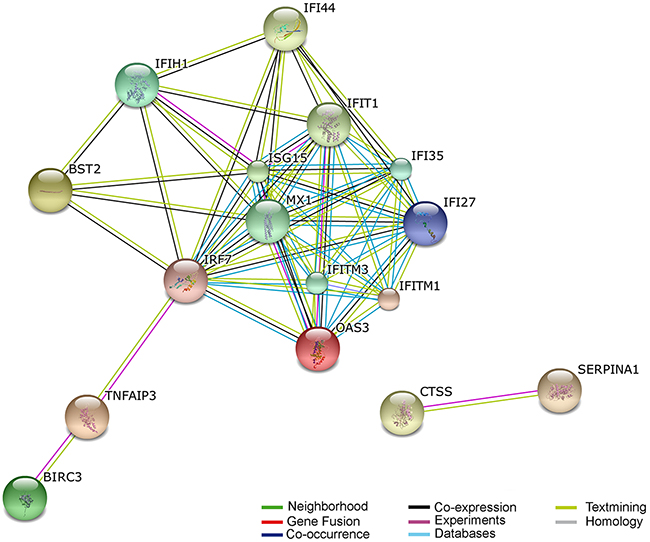
Figure 4: Predicted interaction networks of genes differentially expressed during HHV-6A infection. Differentially expressed genes are depicted: links have been predicted using STRING (http://string.embl.de/). Predicted interactions are depicted according to the type of available evidence. The interactions (see color labels) include direct (physical) and indirect (functional) associations; they are derived from four sources: genomic context, high-throughput experiments, conserved coexpression, and previous knowledge from literature.
Table 1: the differentially expressed genes associated with antivirus, alzheimer’s disease, glioma, and multiple sclerosis
CNS diseases |
Representative DEGs in HA1800-HHV6 GS/control |
|---|---|
Antivirus |
OAS3, LGALS9, IFI35, BIRC3, IFI44, IFITM3, IRF7, IFIT1, IFITM1, IL32, IFI27, ISG15 |
Alzheimer’s disease |
CTSS, SERPINA1, NPTX1, PTX3, CHI3L1, SERPINA3, Mx1 |
Glioma |
CTSS, IRF7, CXCL16, IFITM3, PTX3, CHI3L1, TNFAIP3, BIRC3, BST2, IFIT1, IFITM1 |
Multiple sclerosis |
CHI3L1, IFIH1, KCNJ10, SERPINA1, CXCL16, IFITM3, PTX3, BST2, Mx1 |
DISCUSSION
Genome-wide gene expression profiling is an impartial method to ascertain the pathogenic mechanism underlying virus infection. Our study shows many differentially expressed genes, which related to pathogenic mechanism, are upregulated in HA1800 cells with HHV-6A infection as compared with uninfected controls. The mRNA-Seq technology has no hybridization bias and almost no batch effects on gene expression [15].
Antiviral genes are activated in HA1800 cells with HHV-6A infection. In addition, new effects of anti-viral can be disclosed via comparative analysis of genome-wide transcriptional profiles. OAS3 p100 employs host antiviral effect against Chikungunya virus [16], dengue virus infection [17], and HCV [18]. Galectin-9 (LGALS9) is rapidly released during acute HIV-1 infection [19]. IFN-inducible protein 35 (IFI35) plays an important role in the type I interferon response induced by foot-and-mouth disease virus protein 2C [20] and in the maintenance of foamy virus latency [21] and vesicular stomatitis virus replication [22]. Baculoviral IAP repeat containing 3 (BIRC3) inhibited hepatitis B virus replication [23]. BIRC3 upregulated by E6 oncoprotein confers resistance to cisplatin in human papillomavirus 16/18-infected lung cancer [24]. Interferon-induced protein 44 (IFI44) inhibits HIV-1 LTR promoter activity [25]. IFITM3 restricts influenza A virus entry [26] and has susceptibility to respiratory viral infection [27]. IFITM3 also restricts reovirus cell entry [28] and morbidity and mortality associated with influenza [29]. Lysine residues of interferon regulatory factor (IRF7) affect the replication of KSHV [30]. IFIT1 (ISG56) recognizes 5′-triphosphate RNA [31]. Interferon induced transmembrane protein 1 (IFITM1) restrict membrane fusion [32]. Interferon, alpha-inducible protein 27 (IFI27, ISG12a) is IFN-induced protein that impact cellular apoptosis [33]. High basal ISG12a may inhibit NDV replication and oncolysis [34]. ISG15 inhibits the replication of influenza A virus [35] and the Japanese encephalitis virus [36] and controls the proinflammatory response against viral infection [37]. The consecutive study of these genes in astrocytes can provide new clue for the elucidating of viral antagonism in HHV-6A infection.
HHV-6 showed 23% positivity in peripheral blood leukocytes samples from Alzheimer’s disease (AD) and 4% from controls. HHV-6 may be environmental risk factors for cognitive deterioration and progression to AD in elderly persons [38-41]. We identified a number of AD related genes in astrocyte HA1800 cells with HHV-6A GS infection. Cathepsins S (CTSS) was evaluated as therapeutic target to develop disease modifying drugs to treat AD [42]. Serpin peptidase inhibitor, clade A, member 1 (SERPINA1, ATT) associates with AD-related phenotypes [43] and is recognized as biomarker [44] and potential indicator [45] for AD. Neuronal pentraxin 1 (NPTX1) is overexpressed in dystrophic neurites in AD [46]. Pentraxin-3 (PTX3) is an inflammatory marker [47] and its plasma levels are increased in patients with Parkinson’s disease [48]. PTX3 is also a putative AD biomarker and pharmacological therapeutic target [49]. Cerebrospinal fluid level of chitinase 3-like 1 (CHI3L1, YKL-40) protein is elevated in AD [50-54] and could track the inflammatory processes in AD [55]. YKL-40 has potential prognostic utility as a biomarker for preclinical AD [56]. SERPINA3 (ACT) polymorphism may affect age-at-onset and disease duration of AD [57]. The appearance of MX dynamin-like GTPase 1 (Mx1, MxA) protein in reactive microglia contributes to AD pathology [58]. Therefore, it is conceivable that these genes, combined with previously known mechanisms, may contribute to discovering the correlation between HHV-6A infection and the progression of AD.
HHV-6 DNA was detected in 86% of Nodular Sclerosis Hodgkin lymphoma (NSHL) cases. It suggests that HHV-6 may play an important role in NSHL pathogenesis [59] High percentages of HHV-6 DNA and protein were found in glioma tissue. Additionally, a strain of HHV-6A was isolated from the fluid specimens from glioma cysts. Our previous studies strongly show that HHV-6 infection is involved in the pathogenesis of glioma [60]. Activation of HHV-6 may lead to decrease of lymphocytes total count and develop immunosuppression in patients with gastrointestinal cancer [61]. Cathepsin S (CTSS) expression is linked with tumor progression and poor outcome in glioblastomas [62]. Interferon regulatory factor 7 (IRF7) can enhance glioma cell invasion, chemoresistance, and radioresistance [63]. CXCL16 is highly expressed by glial tumor and stroma cells in glioma [64]. Interferon induced transmembrane protein 3 (IFITM3) plays an important role in glioma cell growth and migration [65]. Pentraxin 3 (PTX3) was significantly associated with the presence of a high-grade glioma tumor [66]. Elevated expression of chitinase 3-like 1 (CHI3L1, YKL-40) in glioma was correlated with decreases in disease survival [67, 68]. YKL-40 serum values were markedly higher in glioma patients than in healthy subjects [69], and as potential serum biomarker for patients with high-grade glioma [70]. TNF alpha induced protein 3 (TNFAIP3, A20) is a tumor enhancer in glioma [71] and inhibits apoptosis in glioblastoma [72]. A20 may serve as a future therapeutic target [73]. BIRC3 (c-IAP2) facilitates cancer cell survival [74]. Increased c-IAP2 expression was found to enhance IκB-α phosphorylation in GBM cells [75, 76]. BST2 expression is upregulated in high grade human astrocytoma [77-79]. And BST-2 expression was increased once oncogenesis is initiated [80]. IFIT1 (ISG56) expression increases in U373MG human astrocytoma cells. [81]. IFITM1 expression significantly inhibited proliferation, migration, and invasion of glioma [82-84].
HHV-6 has been suggested in several autoimmune diseases, including multiple sclerosis (MS). HHV-6 (especially HHV-6A) could participate in neuroinflammation in MS via promoting inflammatory processes through CD46 binding [85]. Viral load and IgGs reacting with HHV-6 U94/REP protein were significantly higher in MS patients. [86]. In addition, anti-HHV-6 IgG was found in CSF of MS patients [87]. HHV-6 may have a role in long-term infection with demyelination in progressive neurological diseases [88]. Cerebrospinal fluid level of chitinase 3-like 1 (CHI3L1, YL-40) is induced in astrocytes in a variety of neurological diseases [89]. YKL-40 has been proposed as a biomarker of multiple sclerosis [90]. CSF level of YKL-40 is increased in MS [91] and is a prognostic marker in MS [92]. Enhanced skin expression of IFIH1 (MDA5) in dermatomyositis and related autoimmune diseases [93]. KCNJ10 (KIR4.1) is expressed in oligodendrocytes and astrocytes in the adult human brain. [94]. Significant expression differences of SERPINA1 (AAT) were identified as potential disease signatures for MS patients [95] and elevates in the cerebrospinal fluid of patients with MS [96]. CXCL16 could be a novel biomarker and potential predictor of disease activity in MS [97]. IFITM3 leads to neuropathological impairments and brain dysfunction in astrocytes [98]. Pentraxin 3 (PTX3) is a novel biomarker of inflammatory in MS [99]. Bone marrow stromal cell antigen 2(BST2) associated statistically with the risk of getting MS. [100]. The appearance of MX dynamin-like GTPase 1 (Mx1, MxA)mRNA is related to clinical exacerbations of MS. [101].
HHV-6 is a global virus in the adult population and correlated with several neurologic diseases, including Alzheimer’s disease, glioma, and multiple sclerosis in the CNS. In conclusion, based upon the results of our comprehensive analysis of HHV-6A infected HA1800 cells, we revealed several genes correlated with neurologic disorders, especially CTSS, PTX3, CHI3L1, Mx1, CXCL16, BIRC3, and BST2 genes. Our studies highlight the human astrocyte HA1800 infected with HHV-6A GS virus and may enhance the understanding of the HHV-6A pathogenicity. The next challenge is to conduct further studies in revealing the role of these genes under HHV-6A infection.
MATERIALS AND METHODS
Cell culture
Cord blood mononuclear cells (CBMCs) were purified from the cord blood samples obtained from the Affiliated Women and Children Hospital of Nanjing Medical University. These studies were approved by the local ethics committee and institutional review board. All samples were obtained with consent from patients and volunteers. HSB-2 cell line (ATCC, USA) was cultured in 1640 medium (Gibco, USA) containing 10% fetal calf serum (FCS, Gibco, USA). Primary human fetal astrocyte HA1800 were purchased from the Sciencell company (Carlsbad, CA,USA) and cultured in DEME/F12 medium (Gibco, USA) supplemented with 10% FCS.
Infection of astrocyte by the isolated HHV-6A GS
HA1800 (2 × 105/well) were cultured in 6-well plates and then infected with the HHV-6A GS at a multiplicity of infection of 100. The culture HA1800 cells were collected for mRNA-seq at 24 hours after infection. The detail experiments were performed as previous described [60, 102-106].
RNA sequencing and data analysis
The six samples (three HA1800-CTL and three HA1800-HHV6AGS samples) were shipped to the GENEWIZ Company (www.genewiz.com) for library construction and mRNA-Seq. Sequencing library construction included these steps: RNA quality checking (Agilent 2100, Agilent Eukaryote Total RNA Nano Kit), library construction (Illumina TruSeq RNA Sample Pre Kit), library purification (Beckman AMPure XP beads), insert fragments test (Agilent 2100,Agilent High Sensitivity DNA Kit), quantitative analysis of library (ABI 7500 real time PCR instrument; KAPA SYBR green fast universal 2×9 qPCR master mix, GRN), and cBOT automatic cluster (TruSeq PE Cluster Kit v3-cBotHS). High-throughtput sequencing was performed with Illumina HiSeq 2000. mRNA-Seq data analysis consisted of the following steps: data quality checking using the Fastqc software (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and removing excess adaptors to get high-quality and clean reads; mapping the high-quality reads to the poplar tree reference genome (http://www.ncbi.nlm.nih.gov/genome/51?genome_assembly_id=273342), using the TopHat software (version 2.0.9) (Trapnell et al.2012); transcript assembling and expression quantificationusing Cufflinks (version 2). Gene expression was expressed as fragments per kilo-base transcript per million mapped reads (FPKM).
Real-time PCR
RNAs were extracted from cells using TRIzol (Invitrogen, California, USA) kit according to the manufacturer’s instructions. Subsequently, total RNA was reverse transcribed using SuperScript III reverse transcriptase (Invitrogen, California, USA). Real-time PCRs were then performed in ABI PRISM7500 system (Applied Biosystems, California, USA), according to the manufacturer’s instructions. The expression level of each gene was normalized by GAPDH and reported as relative levels. The primers for real-time PCR were shown in Supplementary Table S7.
Western blot
Whole cells were washed in PBS and lysed in RIPA lysis buffer supplemented with protease inhibitor cocktail (Roche, Mannheim, Germany). Total protein was quantified using a BCA Protein Assay Kit (Beyotime, Jiangsu, China), and equal amounts of whole cell lysates were resolved by SDS-polyacrylamide gel electrophoresis (PAGE) and transferred to a polyvinylidene difluoride (PVDF) membrane (Millipore, Eschborn, Germany). The blots were blocked with BSA (5% w/v in PBS) for 1 h at room temperature. The following primary antibodies were applied according to the manufacturer’s instructions. Anti-Cathepsin S (CTSS, ab135651), Anti-MX1 (Mx1, ab95926), Anti-cIAP2 antibody (BIRC3, ab32059), Anti-GAPDH antibody (ab8245) were purchased from Abcam (Cambridge, MA, USA). The appropriate secondary antibodies were used at 1:2,000-1:5,000 (v/v) dilutions in PBS + 0.1% Tween 20 for 1 h at room temperature, and the signals were revealed using ECL kit (Thermo Scientific Pierce, Rockford, USA).
Pathway and network analyses
The Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) (http://string.embl.de/) was used to identify known and predicted interactions (derived from four sources: genomic context, high-throughput experiments, co-expression, and previous knowledge). DAVID Bioinformatic resources (http://david.abcc.ncifcrf.gov/) using the annotation sources GOTERM-BP (biological process), and GOTERM-MF (molecular function) identified functional categories.
ACKNOWLEDGMENTS
This work was supported by grants from the National Natural Science Foundation (81272322 to YC), the Qing Lan Project, the Six Talent Peaks Project (JY-018), and the Student Innovation Training Program of Jiangsu Province.
CONFLICTS OF INTEREST
None of the authors have any conflict of interest.
REFERENCES
1. Zerr DM, Meier AS, Selke SS, Frenkel LM, Huang M-L, Wald A, Rhoads MP, Nguy L, Bornemann R, Morrow RA. A population-based study of primary human herpesvirus 6 infection. New Engl J Med. 2005; 8: 768-776.
2. Dockrell DH. Human herpesvirus 6: molecular biology and clinical features. J Med Microbiol. 2003; Pt1: 5-18.
3. Boehmer P, Nimonkar A. Herpes virus replication. IUBMB life. 2003; 1: 13-22.
4. Flamand L, Komaroff AL, Arbuckle JH, Medveczky PG, Ablashi DV. Review, part 1: Human herpesvirus-6-basic biology, diagnostic testing, and antiviral efficacy. J Med Virol. 2010; 9: 1560-1568.
5. Chen T, Hudnall SD. Anatomical mapping of human herpesvirus reservoirs of infection. Modern pathol. 2006; 5: 726-737.
6. Donati D, Martinelli E, Cassiani-Ingoni R, Ahlqvist J, Hou J, Major EO, Jacobson S. Variant-specific tropism of human herpesvirus 6 in human astrocytes. J Virol. 2005; 15: 9439-9448.
7. Ahlqvist J, Fotheringham J, Akhyani N, Yao K, Fogdell-Hahn A, Jacobson S. Differential tropism of human herpesvirus 6 (HHV-6) variants and induction of latency by HHV-6A in oligodendrocytes. J Neurovirol. 2005; 4: 384-394.
8. Ward KN. Human herpesviruses-6 and-7 infections. Curr Opin Infect Dis. 2005; 3: 247-252.
9. Yao K, Honarmand S, Espinosa A, Akhyani N, Glaser C, Jacobson S. Detection of human herpesvirus 6 in cerebrospinal fluid of patients with encephalitis. Ann Neurol. 2009; 3: 257-267.
10. Harberts E, Yao K, Wohler JE, Maric D, Ohayon J, Henkin R, Jacobson S. Human herpesvirus-6 entry into the central nervous system through the olfactory pathway. P Natl Acad Sci USA. 2011; 33: 13734-13739.
11. Yao K, Crawford JR, Komaroff AL, Ablashi DV, Jacobson S. Review part 2: Human herpesvirus-6 in central nervous system diseases. J Med Virol. 2010; 10: 1669-1678.
12. Crawford JR, Santi MR, Thorarinsdottir HK, Cornelison R, Rushing EJ, Zhang H, Yao K, Jacobson S, MacDonald TJ. Detection of human herpesvirus-6 variants in pediatric brain tumors: association of viral antigen in low grade gliomas. J Clin Virol. 2009; 1: 37-42.
13. Chi J, Wang F, Li L, Feng D, Qin J, Xie F, Zhou F, Chen Y, Wang J, Yao K. The role of MAPK in CD4+ T cells toll-like receptor 9-mediated signaling following HHV-6 infection. Virology. 2012; 1: 92-98.
14. Li L, Gu B, Zhou F, Chi J, Wang F, Liu G, Ding C, Xie F, Qing J, Guo Y. Human herpesvirus 6A infects human embryonic fibroblasts and induces G2/M arrest and cell death. J Med Virol. 2012; 4: 657-663.
15. Li L, Gu B, Zhou F, Chi J, Wang F, Peng G, Xie F, Qing J, Feng D, Lu S. Human herpesvirus 6 suppresses T cell proliferation through induction of cell cycle arrest in infected cells in the G2/M phase. J Virol. 2011; 13: 6774-6783.
16. Wang F, Chi J, Peng G, Zhou F, Wang J, Li L, Feng D, Xie F, Gu B, Qin J. Development of virus-specific CD4+ and CD8+ regulatory T cells induced by human herpesvirus 6 infection. J Virol. 2014; 2: 1011-1024.
17. Gu B, Zhang G-F, Li L-Y, Zhou F, Feng D-J, Ding C-L, Chi J, Zhang C, Guo D-D, Wang J-F. Human herpesvirus 6A induces apoptosis of primary human fetal astrocytes via both caspase-dependent and-independent pathways. Virol J. 2011; 1: 530.
18. Chi J, Gu B, Zhang C, Peng G, Zhou F, Chen Y, Zhang G, Guo Y, Guo D, Qin J, Wang J, Li L, Wang F, et al. Human herpesvirus 6 latent infection in patients with glioma. J Infect Dis. 2012; 9: 1394-1398.
19. Huang DW, Sherman BT, Tan Q, Kir J, Liu D, Bryant D, Guo Y, Stephens R, Baseler MW, Lane HC. DAVID Bioinformatics Resources: expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 2007; suppl 2: W169-W175.
20. Huang DW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2008; 1: 44-57.
21. Hwang Y, Kim J, Shin JY, Kim JI, Seo JS, Webster MJ, Lee D, Kim S. Gene expression profiling by mRNA sequencing reveals increased expression of immune/inflammation-related genes in the hippocampus of individuals with schizophrenia. Transl Psychiatry. 2013: e321.
22. Brehin AC, Casademont I, Frenkiel MP, Julier C, Sakuntabhai A, Despres P. The large form of human 2',5'-Oligoadenylate Synthetase (OAS3) exerts antiviral effect against Chikungunya virus. Virology. 2009; 1: 216-222.
23. Lin RJ, Yu HP, Chang BL, Tang WC, Liao CL, Lin YL. Distinct antiviral roles for human 2',5'-oligoadenylate synthetase family members against dengue virus infection. J Immunol. 2009; 12: 8035-8043.
24. Kwon YC, Kang JI, Hwang SB, Ahn BY. The ribonuclease L-dependent antiviral roles of human 2',5'-oligoadenylate synthetase family members against hepatitis C virus. FEBS Lett. 2013; 2: 156-164.
25. Tandon R, Chew GM, Byron MM, Borrow P, Niki T, Hirashima M, Barbour JD, Norris PJ, Lanteri MC, Martin JN, Deeks SG, Ndhlovu LC. Galectin-9 is rapidly released during acute HIV-1 infection and remains sustained at high levels despite viral suppression even in elite controllers. AIDS Res Hum Retroviruses. 2014; 7: 654-664.
26. Zheng W, Li X, Wang J, Cao H, Wang Y, Zeng Q, Zheng SJ. A critical role of interferon-induced protein IFP35 in the type I interferon response in cells induced by foot-and-mouth disease virus (FMDV) protein 2C. Arch Virol. 2014; 11: 2925-2935.
27. Tan J, Qiao W, Wang J, Xu F, Li Y, Zhou J, Chen Q, Geng Y. IFP35 is involved in the antiviral function of interferon by association with the viral tas transactivator of bovine foamy virus. J Virol. 2008; 9: 4275-4283.
28. Das A, Dinh PX, Panda D, Pattnaik AK. Interferon-inducible protein IFI35 negatively regulates RIG-I antiviral signaling and supports vesicular stomatitis virus replication. J Virol. 2014; 6: 3103-3113.
29. Wang Z, Ni J, Li J, Shi B, Xu Y, Yuan Z. Inhibition of hepatitis B virus replication by cIAP2 involves accelerating the ubiquitin-proteasome-mediated destruction of polymerase. J Virol. 2011; 21: 11457-11467.
30. Wu HH, Wu JY, Cheng YW, Chen CY, Lee MC, Goan YG, Lee H. cIAP2 upregulated by E6 oncoprotein via epidermal growth factor receptor/phosphatidylinositol 3-kinase/AKT pathway confers resistance to cisplatin in human papillomavirus 16/18-infected lung cancer. Clin Cancer Res. 2010; 21: 5200-5210.
31. Power D, Santoso N, Dieringer M, Yu J, Huang H, Simpson S, Seth I, Miao H, Zhu J. IFI44 suppresses HIV-1 LTR promoter activity and facilitates its latency. Virology. 2015: 142-150.
32. Desai TM, Marin M, Chin CR, Savidis G, Brass AL, Melikyan GB. IFITM3 restricts influenza A virus entry by blocking the formation of fusion pores following virus-endosome hemifusion. PLoS Pathog. 2014; 4: e1004048.
33. Mills TC, Rautanen A, Elliott KS, Parks T, Naranbhai V, Ieven MM, Butler CC, Little P, Verheij T, Garrard CS, Hinds C, Goossens H, Chapman S, et al. IFITM3 and susceptibility to respiratory viral infections in the community. J Infect Dis. 2014; 7: 1028-1031.
34. Anafu AA, Bowen CH, Chin CR, Brass AL, Holm GH. Interferon-inducible transmembrane protein 3 (IFITM3) restricts reovirus cell entry. J Biol Chem. 2013; 24: 17261-17271.
35. Everitt AR, Clare S, Pertel T, John SP, Wash RS, Smith SE, Chin CR, Feeley EM, Sims JS, Adams DJ, Wise HM, Kane L, Goulding D, et al. IFITM3 restricts the morbidity and mortality associated with influenza. Nature. 2012; 7395: 519-523.
36. Zhang T, Wang Y, Zhang L, Liu B, Xie J, Wood C, Wang J. Lysine residues of interferon regulatory factor 7 affect the replication and transcription activator-mediated lytic replication of Kaposi's sarcoma-associated herpesvirus/human herpesvirus 8. J Gen Virol. 2011; Pt 1: 181-187.
37. Pichlmair A, Lassnig C, Eberle CA, Gorna MW, Baumann CL, Burkard TR, Burckstummer T, Stefanovic A, Krieger S, Bennett KL, Rulicke T, Weber F, Colinge J, et al. IFIT1 is an antiviral protein that recognizes 5'-triphosphate RNA. Nat Immunol. 2011; 7: 624-630.
38. Li K, Markosyan RM, Zheng YM, Golfetto O, Bungart B, Li M, Ding S, He Y, Liang C, Lee JC, Gratton E, Cohen FS, Liu SL. IFITM proteins restrict viral membrane hemifusion. PLoS Pathog. 2013; 1: e1003124.
39. Rosebeck S, Leaman DW. Mitochondrial localization and pro-apoptotic effects of the interferon-inducible protein ISG12a. Apoptosis. 2008; 4: 562-572.
40. Liu N, Long Y, Liu B, Yang D, Li C, Chen T, Wang X, Liu C, Zhu H. ISG12a mediates cell response to Newcastle disease viral infection. Virology. 2014: 283-294.
41. Zhao C, Hsiang TY, Kuo RL, Krug RM. ISG15 conjugation system targets the viral NS1 protein in influenza A virus-infected cells. Proc Natl Acad Sci USA. 2010; 5: 2253-2258.
42. Hsiao NW, Chen JW, Yang TC, Orloff GM, Wu YY, Lai CH, Lan YC, Lin CW. ISG15 over-expression inhibits replication of the Japanese encephalitis virus in human medulloblastoma cells. Antiviral Res. 2010; 3: 504-511.
43. Eduardo-Correia B, Martinez-Romero C, Garcia-Sastre A, Guerra S. ISG15 is counteracted by vaccinia virus E3 protein and controls the proinflammatory response against viral infection. J Virol. 2014; 4: 2312-2318.
44. Caserta MT, Hall CB, Canfield RL, Davidson P, Lofthus G, Schnabel K, Carnahan J, Shelley L, Wang H. Early developmental outcomes of children with congenital HHV-6 infection. Pediatrics. 2014; 6: 1111-1118.
45. Licastro F, Raschi E, Carbone I, Porcellini E. Variants in Antiviral Genes are Risk Factors for Cognitive Decline and Dementia. J Alzheimers Dis. 2015.
46. Wozniak MA, Shipley SJ, Combrinck M, Wilcock GK, Itzhaki RF. Productive herpes simplex virus in brain of elderly normal subjects and Alzheimer's disease patients. J Med Virol. 2005; 2: 300-306.
47. Itzhaki R, Wozniak M, Appelt D, Balin B. Infiltration of the brain by pathogens causes Alzheimer’s disease. Neurobiol Aging. 2004; 5: 619-627.
48. Schechter I, Ziv E. Cathepsins S, B and L with aminopeptidases display beta-secretase activity associated with the pathogenesis of Alzheimer's disease. Biol Chem. 2011; 6: 555-569.
49. Kiddle SJ, Sattlecker M, Proitsi P, Simmons A, Westman E, Bazenet C, Nelson SK, Williams S, Hodges A, Johnston C, Soininen H, Kloszewska I, Mecocci P, et al. Candidate blood proteome markers of Alzheimer's disease onset and progression: a systematic review and replication study. J Alzheimers Dis. 2014; 3: 515-531.
50. Kim S, Lee HJ. Direct Detection of alpha-1 Antitrypsin in Serum Samples using Surface Plasmon Resonance with a New Aptamer-Antibody Sandwich Assay. Anal Chem. 2015; 14: 7235-7240.
51. Yang MH, Yang YH, Lu CY, Jong SB, Chen LJ, Lin YF, Wu SJ, Chu PY, Chung TW, Tyan YC. Activity-dependent neuroprotector homeobox protein: A candidate protein identified in serum as diagnostic biomarker for Alzheimer's disease. J Proteomics. 2012; 12: 3617-3629.
52. Abad MA, Enguita M, DeGregorio-Rocasolano N, Ferrer I, Trullas R. Neuronal pentraxin 1 contributes to the neuronal damage evoked by amyloid-beta and is overexpressed in dystrophic neurites in Alzheimer's brain. J Neurosci. 2006; 49: 12735-12747.
53. Ko CY, Chang LH, Lee YC, Sterneck E, Cheng CP, Chen SH, Huang AM, Tseng JT, Wang JM. CCAAT/enhancer binding protein delta (CEBPD) elevating PTX3 expression inhibits macrophage-mediated phagocytosis of dying neuron cells. Neurobiol Aging. 2012; 2: 422 e411-425.
54. Lee HW, Choi J, Suk K. Increases of pentraxin 3 plasma levels in patients with Parkinson's disease. Mov Disord. 2011; 13: 2364-2370.
55. Osera C, Pascale A, Amadio M, Venturini L, Govoni S, Ricevuti G. Pentraxins and Alzheimer's disease: at the interface between biomarkers and pharmacological targets. Ageing Res Rev. 2012; 2: 189-198.
56. Antonell A, Mansilla A, Rami L, Llado A, Iranzo A, Olives J, Balasa M, Sanchez-Valle R, Molinuevo JL. Cerebrospinal fluid level of YKL-40 protein in preclinical and prodromal Alzheimer's disease. J Alzheimers Dis. 2014; 3: 901-908.
57. Kester MI, Teunissen CE, Sutphen C, Herries EM, Ladenson JH, Xiong C, Scheltens P, van der Flier WM, Morris JC, Holtzman DM. Cerebrospinal fluid VILIP-1 and YKL-40, candidate biomarkers to diagnose, predict and monitor Alzheimer’s disease in a memory clinic cohort. Alzheimers Res Ther. 2015; 1: 1-9.
58. Wennström M, Surova Y, Hall S, Nilsson C, Minthon L, Hansson O, Nielsen HM. The Inflammatory Marker YKL-40 Is Elevated in Cerebrospinal Fluid from Patients with Alzheimer’s but Not Parkinson’s Disease or Dementia with Lewy Bodies. PLoS One. 2015; 8: e0135458.
59. Gispert JD, Monté GC, Falcon C, Tucholka A, Rojas S, Sánchez-Valle R, Antonell A, Lladó A, Rami L, Molinuevo JL. CSF YKL-40 and pTau181 are related to different cerebral morphometric patterns in early AD. Neurobiol Aging. 2015.
60. Rosén C, Andersson C-H, Andreasson U, Molinuevo JL, Bjerke M, Rami L, Lladó A, Blennow K, Zetterberg H. Increased Levels of Chitotriosidase and YKL-40 in Cerebrospinal Fluid from Patients with Alzheimer's Disease. Dement Geriatr Cogn. 2014; 2: 297-304.
61. Alcolea D, Vilaplana E, Pegueroles J, Montal V, Sanchez-Juan P, Gonzalez-Suarez A, Pozueta A, Rodriguez-Rodriguez E, Bartres-Faz D, Vidal-Pineiro D, Gonzalez-Ortiz S, Medrano S, Carmona-Iragui M, et al. Relationship between cortical thickness and cerebrospinal fluid YKL-40 in predementia stages of Alzheimer's disease. Neurobiol Aging. 2015; 6: 2018-2023.
62. Craig-Schapiro R, Perrin RJ, Roe CM, Xiong C, Carter D, Cairns NJ, Mintun MA, Peskind ER, Li G, Galasko DR, Clark CM, Quinn JF, D'Angelo G, et al. YKL-40: a novel prognostic fluid biomarker for preclinical Alzheimer's disease. Biol Psychiatry. 2010; 10: 903-912.
63. Kamboh MI, Minster RL, Kenney M, Ozturk A, Desai PP, Kammerer CM, DeKosky ST. Alpha-1-antichymotrypsin (ACT or SERPINA3) polymorphism may affect age-at-onset and disease duration of Alzheimer's disease. Neurobiol Aging. 2006; 10: 1435-1439.
64. Yamada T, Horisberger MA, Kawaguchi N, Moroo I, Toyoda T. Immunohistochemistry using antibodies to alpha-interferon and its induced protein, MxA, in Alzheimer's and Parkinson's disease brain tissues. Neurosci Lett. 1994; 1-2: 61-64.
65. Siddon A, Lozovatsky L, Mohamed A, Hudnall SD. Human herpesvirus 6 positive Reed-Sternberg cells in nodular sclerosis Hodgkin lymphoma. Br J Haematol. 2012; 5: 635-643.
66. Sultanova A, Chistjakovs M, Chapenko S, Donina S, Murovska M. Possible interference of human beta-herpesviruses-6 and -7 in gastrointestinal cancer development. Exp Oncol. 2013; 2: 93-96.
67. Flannery T, McQuaid S, McGoohan C, McConnell RS, McGregor G, Mirakhur M, Hamilton P, Diamond J, Cran G, Walker B, Scott C, Martin L, Ellison D, et al. Cathepsin S expression: An independent prognostic factor in glioblastoma tumours--A pilot study. Int J Cancer. 2006; 4: 854-860.
68. Kim JK, Jin X, Ham SW, Lee SY, Seo S, Kim SC, Kim SH, Kim H. IRF7 promotes glioma cell invasion by inhibiting AGO2 expression. Tumour Biol. 2015; 7: 5561-5569.
69. Hattermann K, Held-Feindt J, Ludwig A, Mentlein R. The CXCL16-CXCR6 chemokine axis in glial tumors. J Neuroimmunol. 2013; 1-2: 47-54.
70. Zhao B, Wang H, Zong G, Li P. The role of IFITM3 in the growth and migration of human glioma cells. BMC Neurol. 2013: 210.
71. Locatelli M, Ferrero S, Martinelli Boneschi F, Boiocchi L, Zavanone M, Maria Gaini S, Bello L, Valentino S, Barbati E, Nebuloni M, Mantovani A, Garlanda C. The long pentraxin PTX3 as a correlate of cancer-related inflammation and prognosis of malignancy in gliomas. J Neuroimmunol. 2013; 1-2: 99-106.
72. Colin C, Baeza N, Bartoli C, Fina F, Eudes N, Nanni I, Martin PM, Ouafik L, Figarella-Branger D. Identification of genes differentially expressed in glioblastoma versus pilocytic astrocytoma using Suppression Subtractive Hybridization. Oncogene. 2006; 19: 2818-2826.
73. Pelloski CE, Ballman KV, Furth AF, Zhang L, Lin E, Sulman EP, Bhat K, McDonald JM, Yung WK, Colman H, Woo SY, Heimberger AB, Suki D, et al. Epidermal growth factor receptor variant III status defines clinically distinct subtypes of glioblastoma. J Clin Oncol. 2007; 16: 2288-2294.
74. Bernardi D, Padoan A, Ballin A, Sartori M, Manara R, Scienza R, Plebani M, Della Puppa A. Serum YKL-40 following resection for cerebral glioblastoma. J Neurooncol. 2012; 2: 299-305.
75. Hormigo A, Gu B, Karimi S, Riedel E, Panageas KS, Edgar MA, Tanwar MK, Rao JS, Fleisher M, DeAngelis LM, Holland EC. YKL-40 and matrix metalloproteinase-9 as potential serum biomarkers for patients with high-grade gliomas. Clin Cancer Res. 2006; 19: 5698-5704.
76. Hjelmeland AB, Wu Q, Wickman S, Eyler C, Heddleston J, Shi Q, Lathia JD, Macswords J, Lee J, McLendon RE, Rich JN. Targeting A20 decreases glioma stem cell survival and tumor growth. PLoS Biol. 2010; 2: e1000319.
77. Bellail AC, Olson JJ, Yang X, Chen ZJ, Hao C. A20 ubiquitin ligase-mediated polyubiquitination of RIP1 inhibits caspase-8 cleavage and TRAIL-induced apoptosis in glioblastoma. Cancer Discov. 2012; 2: 140-155.
78. Guo Q, Dong H, Liu X, Wang C, Liu N, Zhang J, Li B, Cao W, Ding T, Yang Z, Zhang X. A20 is overexpressed in glioma cells and may serve as a potential therapeutic target. Expert Opin Ther Targets. 2009; 7: 733-741.
79. Bertrand MJ, Milutinovic S, Dickson KM, Ho WC, Boudreault A, Durkin J, Gillard JW, Jaquith JB, Morris SJ, Barker PA. cIAP1 and cIAP2 facilitate cancer cell survival by functioning as E3 ligases that promote RIP1 ubiquitination. Mol Cell. 2008; 6: 689-700.
80. Gyrd-Hansen M, Darding M, Miasari M, Santoro MM, Zender L, Xue W, Tenev T, da Fonseca PC, Zvelebil M, Bujnicki JM, Lowe S, Silke J, Meier P. IAPs contain an evolutionarily conserved ubiquitin-binding domain that regulates NF-kappaB as well as cell survival and oncogenesis. Nat Cell Biol. 2008; 11: 1309-1317.
81. Yang W, Cooke M, Duckett CS, Yang X, Dorsey JF. Distinctive effects of the cellular inhibitor of apoptosis protein c-IAP2 through stabilization by XIAP in glioblastoma multiforme cells. Cell Cycle. 2014; 6: 992-1005.
82. Wainwright DA, Balyasnikova IV, Han Y, Lesniak MS. The expression of BST2 in human and experimental mouse brain tumors. Exp Mol Pathol. 2011; 1: 440-446.
83. Milutin Gasperov N, Farkas SA, Nilsson TK, Grce M. Epigenetic activation of immune genes in cervical cancer. Immunol Lett. 2014; 2 Pt B: 256-257.
84. Chiang SF, Kan CY, Hsiao YC, Tang R, Hsieh LL, Chiang JM, Tsai WS, Yeh CY, Hsieh PS, Liang Y, Chen JS, Yu JS. Bone Marrow Stromal Antigen 2 Is a Novel Plasma Biomarker and Prognosticator for Colorectal Carcinoma: A Secretome-Based Verification Study. Dis Markers. 2015: 874054.
85. Jones PH, Mahauad-Fernandez WD, Madison MN, Okeoma CM. BST-2/tetherin is overexpressed in mammary gland and tumor tissues in MMTV-induced mammary cancer. Virology. 2013; 1-2: 124-139.
86. Imaizumi T, Murakami K, Ohta K, Seki H, Matsumiya T, Meng P, Hayakari R, Xing F, Aizawa-Yashiro T, Tatsuta T, Yoshida H, Kijima H. MDA5 and ISG56 mediate CXCL10 expression induced by toll-like receptor 4 activation in U373MG human astrocytoma cells. Neurosci Res. 2013; 4: 195-206.
87. Lee J, Goh SH, Song N, Hwang JA, Nam S, Choi IJ, Shin A, Kim IH, Ju MH, Jeong JS, Lee YS. Overexpression of IFITM1 has clinicopathologic effects on gastric cancer and is regulated by an epigenetic mechanism. Am J Pathol. 2012; 1: 43-52.
88. Yu F, Ng SS, Chow BK, Sze J, Lu G, Poon WS, Kung HF, Lin MC. Knockdown of interferon-induced transmembrane protein 1 (IFITM1) inhibits proliferation, migration, and invasion of glioma cells. J Neurooncol. 2011; 2: 187-195.
89. Hatano H, Kudo Y, Ogawa I, Tsunematsu T, Kikuchi A, Abiko Y, Takata T. IFN-induced transmembrane protein 1 promotes invasion at early stage of head and neck cancer progression. Clin Cancer Res. 2008; 19: 6097-6105.
90. Descamps V. Diagnosis of DRESS (drug reaction with eosinophilia and systemic symptoms) in the intensive care unit: essential but challenging. Shock. 2013; 5: 437-438.
91. Ben-Fredj N, Ben-Selma W, Rotola A, Nefzi F, Benedetti S, Frih-Ayed M, Di Luca D, Aouni M, Caselli E. Prevalence of human herpesvirus U94/REP antibodies and DNA in Tunisian multiple sclerosis patients. J Neurovirol. 2013; 1: 42-47.
92. Alenda R, Alvarez-Lafuente R, Costa-Frossard L, Arroyo R, Mirete S, Alvarez-Cermeno JC, Villar LM. Identification of the major HHV-6 antigen recognized by cerebrospinal fluid IgG in multiple sclerosis. Eur J Neurol. 2014; 8: 1096-1101.
93. Pietilainen-Nicklen J, Virtanen JO, Uotila L, Salonen O, Farkkila M, Koskiniemi M. HHV-6-positivity in diseases with demyelination. J Clin Virol. 2014; 2: 216-219.
94. Bonneh-Barkay D, Wang G, Starkey A, Hamilton RL, Wiley CA. In vivo CHI3L1 (YKL-40) expression in astrocytes in acute and chronic neurological diseases. J Neuroinflammation. 2010: 34.
95. Bhardwaj R, Yester JW, Singh SK, Biswas DD, Surace MJ, Waters MR, Hauser KF, Yao Z, Boyce BF, Kordula T. RelB/p50 complexes regulate cytokine-induced YKL-40 expression. J Immunol. 2015; 6: 2862-2870.
96. Malmestrom C, Axelsson M, Lycke J, Zetterberg H, Blennow K, Olsson B. CSF levels of YKL-40 are increased in MS and replaces with immunosuppressive treatment. J Neuroimmunol. 2014; 1-2: 87-89.
97. Martinez MA, Olsson B, Bau L, Matas E, Cobo Calvo A, Andreasson U, Blennow K, Romero-Pinel L, Martinez-Yelamos S, Zetterberg H. Glial and neuronal markers in cerebrospinal fluid predict progression in multiple sclerosis. Mult Scler. 2015; 5: 550-561.
98. Zahn S, Barchet W, Rehkamper C, Hornung T, Bieber T, Tuting T, Wenzel J. Enhanced skin expression of melanoma differentiation-associated gene 5 (MDA5) in dermatomyositis and related autoimmune diseases. J Am Acad Dermatol. 2011; 5: 988-989.
99. Schirmer L, Srivastava R, Kalluri SR, Bottinger S, Herwerth M, Carassiti D, Srivastava B, Gempt J, Schlegel J, Kuhlmann T, Korn T, Reynolds R, Hemmer B. Differential loss of KIR4.1 immunoreactivity in multiple sclerosis lesions. Ann Neurol. 2014; 6: 810-828.
100. Amin B, Maurer A, Voelter W, Melms A, Kalbacher H. New poteintial serum biomarkers in multiple sclerosis identified by proteomic strategies. Curr Med Chem. 2014; 13: 1544-1556.
101. Lewis EC. Expanding the clinical indications for alpha(1)-antitrypsin therapy. Mol Med. 2012: 957-970.
102. Holmoy T, Loken-Amsrud KI, Bakke SJ, Beiske AG, Bjerve KS, Hovdal H, Lilleas F, Midgard R, Pedersen T, Saltyte Benth J, Torkildsen O, Wergeland S, Myhr KM, et al. Inflammation markers in multiple sclerosis: CXCL16 reflects and may also predict disease activity. PLoS One. 2013; 9: e75021.
103. Ibi D, Nagai T, Nakajima A, Mizoguchi H, Kawase T, Tsuboi D, Kano S, Sato Y, Hayakawa M, Lange UC, Adams DJ, Surani MA, Satoh T, et al. Astroglial IFITM3 mediates neuronal impairments following neonatal immune challenge in mice. Glia. 2013; 5: 679-693.
104. Wang H, Wang K, Wang C, Zhong X, Qiu W, Hu X. Increased plasma levels of pentraxin 3 in patients with multiple sclerosis and neuromyelitis optica. Mult Scler. 2013; 7: 926-931.
105. Nexo BA, Hansen B, Nissen KK, Gundestrup L, Terkelsen T, Villesen P, Bahrami S, Petersen T, Pedersen FS, Laska MJ. Restriction genes for retroviruses influence the risk of multiple sclerosis. PLoS One. 2013; 9: e74063.
106. van der Voort LF, Vennegoor A, Visser A, Knol DL, Uitdehaag BM, Barkhof F, Oudejans CB, Polman CH, Killestein J. Spontaneous MxA mRNA level predicts relapses in patients with recently diagnosed MS. Neurology. 2010; 14: 1228-1233.

