Interview with Dr. Kelber from California State University Northridge
Oncotarget published " A novel method for RNA extraction from FFPE samples reveals significant differences in biomarker expression between orthotopic and subcutaneous pancreatic cancer patient-derived xenografts " which reported that performing NGS on FFPE samples is limited by poor RNA purification methods.
To address this hurdle, the researchers developed an improved methodology for extracting high-quality RNA from FFPE samples.
By briefly integrating a newly-designed micro-homogenizing tool with commercially available FFPE RNA extraction protocols, RNA recovery is increased by approximately 3-fold while maintaining standard A260/A280 ratios and RNA quality index values.
Finally, expression analysis of novel biomarkers in KRas mutant PDAC samples revealed that PEAK1 decreases and MST1R increases by over 100-fold in orthotopic versus subcutaneous microenvironments.
Furthermore, this Oncotarget Research Paper offers a new mH-based FFPE RNA extraction method has the potential to enhance and expand future FFPE-RNA-NGS cancer biomarker studies.
Dr. Jonathan A. Kelber from The California State University Northridge said, "Next-Generation Sequencing (NGS), such as RNA-seq or whole-genome analysis, of patient-derived tumor tissue in combination with state-of-the art bioinformatics holds great potential to improve disease outcomes by identifying novel biomarkers of cancer onset, progression and therapy resistance. "
Among the advantages of NGS over other technologies is the ability to perform high-resolution transcriptome monitoring to identify isoform variations, outlier expression patterns and expression signatures of low expressing genes or long noncoding RNAs.
However, understanding the effects of the tumor microenvironment on gene expression and, in particular, cancer-cell biomarker profiles is hindered by the fact that researchers have limited access to fresh tumor tissue, which harbor good-quality and easily-accessible nucleic acid material - these samples are not routinely obtained in hospitals due to the logistical challenges of collecting, processing and banking fresh-frozen tissue .
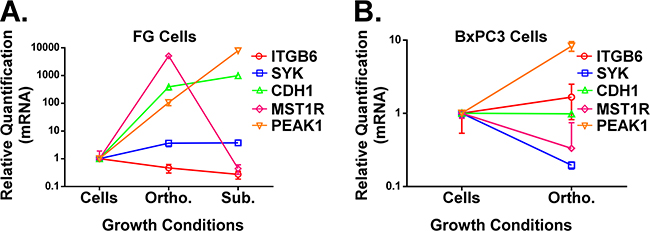
Figure 4: A and B. qPCR analysis of ITGB6, SYK, CDH1, PEAK1 and MST1R in FG (A, KRas mutant G12D line) or BxPC3 (B, KRas wild type line) PDAC cells grown under the indicated microenvironmental conditions. All RNA was extracted using our mH-modified Qiagen FFPE kit protocol and relative quantification (RQ) values for gene expression were normalized to house-keeping genes (GAPDH and/or POLR2A).
Yet, it remains extremely difficult to recover sufficient high-quality RNA from these FFPE samples due to formalin-induced cross-linking and degradation as well as the requirement for large amounts of the FFPE samples and time consuming methods for RNA extraction.
The authors hypothesized that the microHomogenizer ™ device could significantly disrupt FFPE tissue to liberate more high-quality RNA material and increase the efficiency of RNA purification from FFPE samples using standard kits.
In the present study, the combination of their mH technology with the Qiagen FFPE RNA extraction kit has significantly improved RNA quality and yield from pancreatic ductal adenocarcinoma patient and cell line derived xenograft FFPE samples.
The Kelber Research Team concluded in their Oncotarget Research Paper, "Previously developed concepts and strategies of highly selective tumor targeting can now take advantage of these new methods of biomarker measurement using the mH-based RNA extraction method described here and NGS technology to identify differences between the microenvironment of normal and tumor tissue. "
Sign up for free Altmetric alerts about this article
DOI - https://doi.org/10.18632/oncotarget.11809
Full text - https://www.oncotarget.com/article/11809/text/
Correspondence to - Jonathan A. Kelber - [email protected]
Keywords - FFPE RNA extraction, microHomogenizer ™, pancreatic cancer, patient-derived orthotopic xenografts (PDOX) tumor microenvironment, cancer biomarkers

Copyright © 2026 Rapamycin Press LLC dba Impact Journals
Oncotarget ® is a registered trademark of Rapamycin Press LLC
Impact Journals ® is a registered trademark of Rapamycin Press LLC
RAPAMYCIN PRESS ® is a registered trademark of Rapamycin Press LLCß

