Validating and enabling phosphoglycerate dehydrogenase (PHGDH) as a target for fragment-based drug discovery in PHGDH-amplified breast cancer
2019-05-01
The cover for issue 17 of Oncotarget features Figure 4B, "A comparison of the structural elements comprising human PHGDH, sPHGDH and PHGDH-93. Amino acids involved in hydrogen bond formation to the substrate (*) or cofactor NAD+ (*) are highlighted," from Unterlass, et al.
The compounds identified and characterised in this work are structurally different from the reported PHGDH inhibitors CBR-5884 and NCT-503, and thus provide a new starting point for the design of potent and selective PHGDH inhibitors.
CBR-5884 is a non-competitive inhibitor that prevents the oligomerisation of PHGDH, whereas NCT-503, also a non-competitive inhibitor in regards to substrate and cofactor, is supposed to bind close to the active site as mutation of C234 in the active site of the protein reduces the inhibitory effect of PHGDH.
Consistent with the genetic data, knockdown of PHGDH in melanoma and breast cancer cell lines containing a PHGDH amplification, resulted in decreased cell viability.
In addition, PHGDH suppression inhibited the growth of mammary tumours in mice, although the suppressive effect seemed to depend on the tumour stage, as a similar in vivo study in more established mammary tumours reported no significant reduction in tumour growth following PHGDH knockdown.
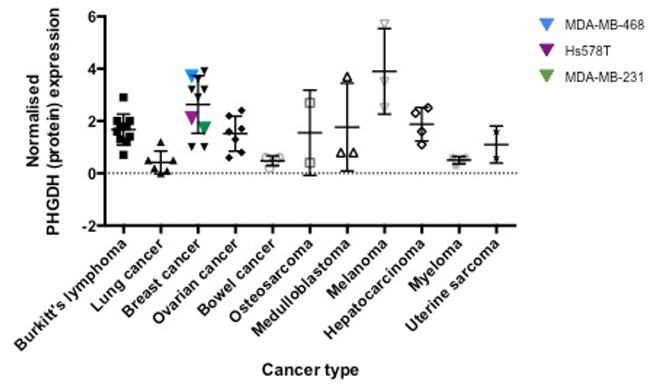
Analysis of PHGDH levels in different cancer cell lines revealed upregulation of PHGDH mRNA rather than changes in enzymatic activity to be the reason for elevated PHGDH activity in human tumour cells.
Subsequent studies showed significantly enhanced PHGDH expression in melanoma and breast cancer, indicating that these cancer types might be susceptible to treatment by PHGDH inhibition.
In an effort to further build on the reported target validation and to provide a chemical probe to interrogate the role of PHGDH activity in cancer, the researchers explored the effects of PHGDH knockdown in a panel of cancer cell lines and developed assays and a crystal system for the identification of fragments that bind to the cofactor binding-site of PHGDH.
Knockdown caused reduced proliferation in the PHGDH-amplified cell line MDA-MB-468, whereas breast cancer cells with low PHGDH expression or with elevated PHGDH expression in the absence of genomic amplification were not affected.
They designed a truncated PHGDH construct that gave crystals which diffracted to high resolution and could be used for fragment soaking.
15 fragments stabilising PHGDH were identified using a thermal shift assay and validated by X-ray crystallography and ITC competition experiments to exhibit 1.5-26.2 mM affinity for PHGDH.
Full text - https://www.oncotarget.com/article/11487/text/
Correspondence to - Martin E.M. Noble - [email protected]
Keywords - cancer metabolism, PHGDH, serine metabolism, drug discovery, fragments
Copyright © 2026 Rapamycin Press LLC dba Impact Journals
Oncotarget ® is a registered trademark of Rapamycin Press LLC
Impact Journals ® is a registered trademark of Rapamycin Press LLC
RAPAMYCIN PRESS ® is a registered trademark of Rapamycin Press LLC




