Oncotarget: The Golgi protein TMEM165 controls migration/invasion for carcinoma
2020-07-14
The cover for issue 28 of Oncotarget features Figure 5, "TMEM165 expression levels alters N-linked glycosylation," by Murali, et al., and reported that the TMEM165 protein was not detected in non-malignant matched breast tissues and was detected in invasive ductal breast carcinoma tissues by mass spectrometry.
The hypothesis is that the TMEM165 protein confers a growth advantage to breast cancer.
The authors created a CRISPR/Cas9 knockout of TMEM165 in the human invasive breast cancer cell line MDAMB231.
Furthermore, they find that TMEM165 expression alters the glycosylation of breast cancer cells and these changes promote the invasion and growth of breast cancer by altering the expression levels of key glycoproteins involved in the regulation of the epithelial to mesenchymal transition such as E-cadherin.
These studies illustrate new potential functions for this Golgi membrane protein in the control of breast cancer growth and invasion.
"These studies illustrate new potential functions for this Golgi membrane protein in the control of breast cancer growth and invasion."
Dr. Karen L. Abbott from The University of Oklahoma Health Sciences Center, Department of Biochemistry and Molecular Biology said "Breast cancer is the most commonly diagnosed cancer in women."
The TMEM165 protein was identified by mass spectrometry in invasive breast carcinoma tissue with no detection in patient-matched adjacent normal breast tissues.
The authors analyzed TCGA breast cancer cases to examine TMEM165 expression levels in all molecular types of human breast cancer using UALCAN.
They found that TMEM165 is amplified across all types of breast cancer compared to normal breast tissue with IDC cases having the highest levels of TMEM165 expression.
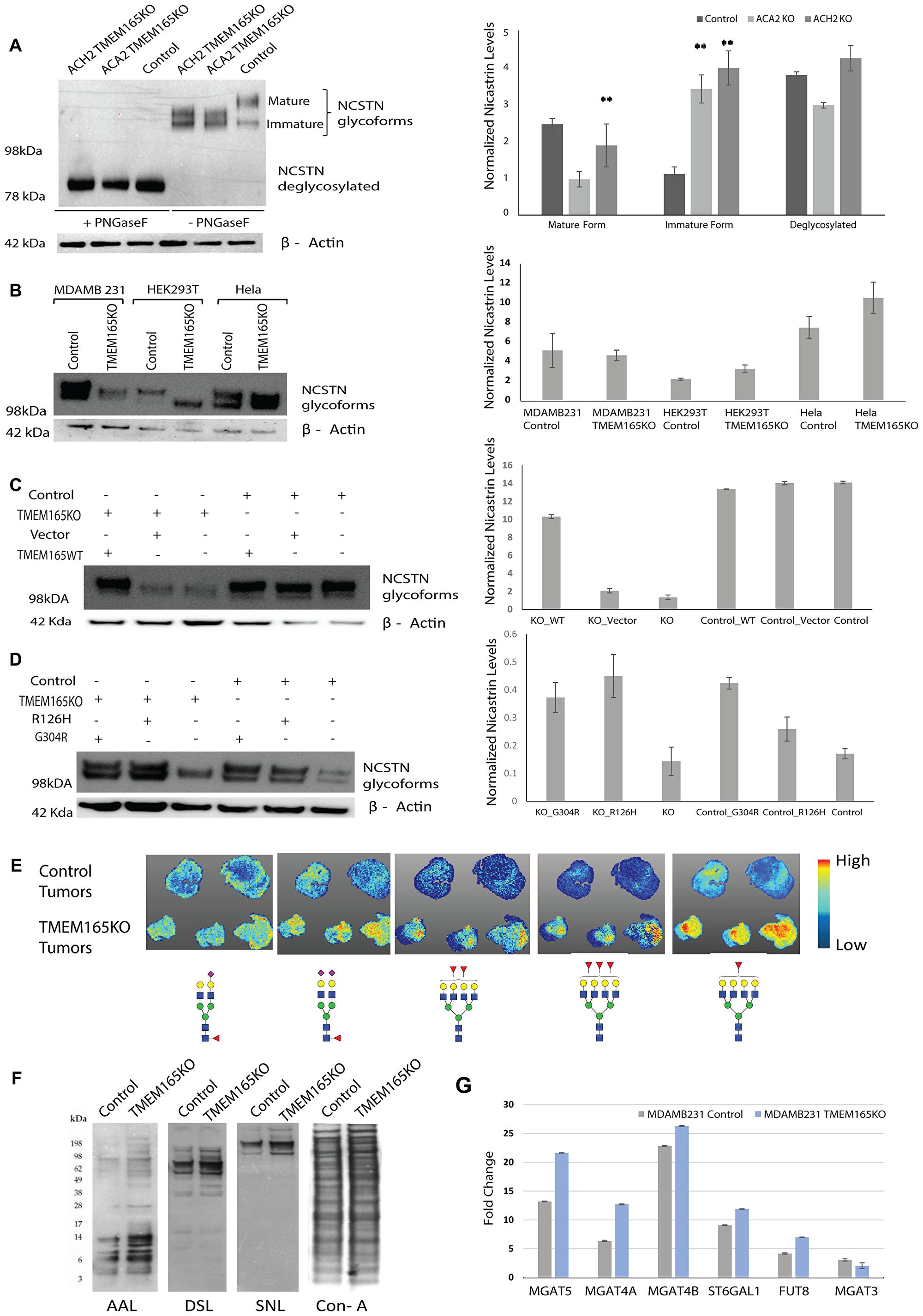
Figure 5: TMEM165 expression levels alters N-linked glycosylation. (A) Western blot analysis of NCSTN from MDAMB231 TMEM165KO (clones ACH2 and ACA2) and MDAMB231 Control (clone ACH11) cell lysates before and after PNGaseF treatment. NCSTN mature, immature, and deglycosylated bands are labeled. The Western blot was imaged on Bio-Rad EZ Imager and has not been manipulated. Any marks on the blot that are not labeled bands are from forceps handling of the membrane. Right panel shows the quantification of mature, immature and deglycosylated forms Nicastrin protein levels after normalization with actin (n = 2, **P < 0.01) (B) Western blot showing different glycoforms of NCSTN in Control and TMEM165KO across different cell lines (MDAMB231, HEK293T and Hela). NCSTN in Control and TMEM165KO across different cell lines (MDAMB231, HEK293T and Hela). Right panel shows the quantification of Nicastrin protein levels after normalization with actin (n = 2). (C, D) Western blot showing rescue of NCSTN glycoforms in TMEM165KO cells after transient transfection with TMEM16WT plasmid and TMEM165CDG mutants (R126H and G304R). Right panel shows the quantification of Nicastrin protein levels after normalization with actin (n = 2,**P < 0.01). (E) Xenograft paraffin embedded tissue sections treated with PNGaseF were subjected to Tissue MALDI-FTICR IMS exhibited differential expression of N-glycan structures among Control and TMEM165KO tumor tissues. Represented glycan structures are in accordance with the guidelines for the Consortium for Functional Glycomics (CFG): blue square, N-acetylglucosamine (GlcNAc); green circle, mannose (Man); yellow circle, galactose (Gal); red triangle, fucose (Fuc); purple diamond, N-acetylneuraminic acid (Neu5Ac). (F) Lectin blot analysis of MDAMB231 control and TMEM165KO cell lysates. (G) Graph representing the relative transcript levels of MGAT5, MGAT4A, MGAT4B, ST6GAL1, FUT8 and MGAT3 in MDAMB231 control or TMEM165KO cells. Normalized data expressed as fold change are plotted for each gene assayed. Error bars represent one standard deviation from the mean of triplicate values from n = 3 experiments.
In the present study, the authors report that TMEM165 is upregulated in human breast cancer cell lines and patient tumor tissues and increased expression of TMEM165 correlates with poor prognosis in breast cancer patients.
Collectively, the data demonstrate that overexpression of TMEM165 promotes EMT in breast cancer suggesting a novel role for TMEM165 as a driver of tumor invasion making it a prognostic marker and potential therapeutic target for breast cancer.
The Abbott Research Team concluded in their Oncotarget Research Paper, "we have expanded on our initial 2012 glycoproteomic study that was the first to identify TMEM165 protein as a potential biomarker for breast carcinoma. In this study, we have provided initial mechanistic studies that indicate that TMEM165 expression drives the growth and invasion of breast cancer. TMEM165 expression levels could be a potential prognostic marker for predicting DCIS cases that may progress to invasive disease. Larger prospective cohorts will need to be analyzed to determine the link between TMEM165 levels and the progression to IDC. We find that IDC patients with higher TMEM165 expression levels have reduced overall survival making this protein a target for the development of new therapeutic strategies to limit the progression of breast cancer."
Sign up for free Altmetric alerts about this article
DOI - https://doi.org/10.18632/oncotarget.27668
Full text - https://www.oncotarget.com/article/27668/text/
Correspondence to - Karen L. Abbott - [email protected]
Keywords - TMEM165, migration, invasion, breast cancer, glycosylation
About Oncotarget
Oncotarget is a biweekly, peer-reviewed, open access biomedical journal covering research on all aspects of oncology.
To learn more about Oncotarget, please visit https://www.oncotarget.com or connect with:
SoundCloud - https://soundcloud.com/oncotarget
Facebook - https://www.facebook.com/Oncotarget/
Twitter - https://twitter.com/oncotarget
LinkedIn - https://www.linkedin.com/company/oncotarget
Pinterest - https://www.pinterest.com/oncotarget/
Reddit - https://www.reddit.com/user/Oncotarget/
Oncotarget is published by Impact Journals, LLC please visit http://www.ImpactJournals.com or connect with @ImpactJrnls
Media Contact
[email protected]
18009220957x105
Copyright © 2026 Rapamycin Press LLC dba Impact Journals
Oncotarget ® is a registered trademark of Rapamycin Press LLC
Impact Journals ® is a registered trademark of Rapamycin Press LLC
RAPAMYCIN PRESS ® is a registered trademark of Rapamycin Press LLC




