Oncotarget: The comprehensive genomic profiling test, GEM ExTra®
2021-04-13
Oncotarget published "Analytic validation and clinical utilization of the comprehensive genomic profiling test, GEM ExTra®" which reported that the authors developed and analytically validated a comprehensive genomic profiling assay, GEM ExTra, for patients with advanced solid tumors that uses Next Generation Sequencing to characterize whole exomes employing a paired tumor-normal subtraction methodology.
The assay detects single nucleotide variants, indels, focal copy number alterations, TERT promoter region, as well as tumor mutation burden and microsatellite instability status.
Additionally, the assay incorporates whole transcriptome sequencing of the tumor sample that allows for the detection of gene fusions and select special transcripts, including AR-V7, EGFR vIII, EGFRvIV, and MET exon 14 skipping events.
The assay has a mean target coverage of 180X for the normal and 400X for tumor DNA including enhanced probe design to facilitate the sequencing of difficult regions.
Proprietary bioinformatics, paired with comprehensive clinical curation results in reporting that defines clinically actionable, FDA-approved, and clinical trial drug options for the management of the patient's cancer.
Proprietary bioinformatics, paired with comprehensive clinical curation results in reporting that defines clinically actionable, FDA-approved, and clinical trial drug options for the management of the patient's cancer.
Dr. Thomas Royce from Ashion Analytics, LLC said, "Cancer has a high clinical burden and oncology therapies are expensive."
There are a few predominant laboratories that perform CGP tumor tests with the intended use of providing clinical decision support for therapy selection for cancer.
GEM ExTra uses Whole Exome Sequencing for tumor DNA profiling, testing for all protein-coding genes in a sample, indicating that the test will be comprehensive now and in the future.
GEM ExTra provides somatic variant calling based on tumor and matched germline sequencing allowing for improved discrimination of somatic variants from rare, benign germline variants when compared to tumor-only analysis used by other CGP tests.
GEM ExTra also identifies clinically actionable transcript variants and fusion genes through transcriptome sequencing. These are typically undetectable through conventional CGP tests, which only employ DNA analysis.
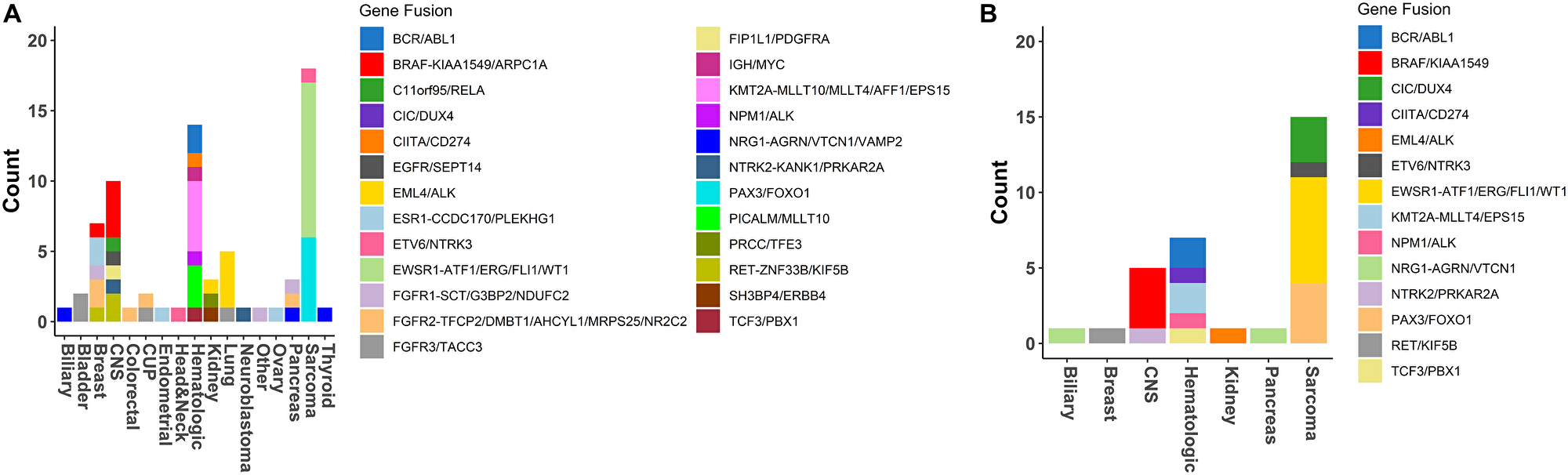
Figure 5: Fusions detection in GEM ExTra. (A) Fusions detected by tumor type. (B) Fusion's detected in tumors types with RNA only findings. For those main fusion genes (e.g., BRAF) that were found to be fused with multiple partner genes (e.g., KIAA1549, or ARPC1A) the partner genes are separated from main fusion gene by a dashed line “-”, and the other partners listed consecutively, separated by a forward slash “/”.
The test is designed to provide healthcare professionals with clinically actionable information to guide patient management decisions based on the genomic profile of a cancer patient's tumor.
The Royce Research Team concluded in their Oncotarget Research Output that the analytic performance characteristics of the assay were validated by comparison of patient samples to reference assays, and actionable variants were identified in tumors to guide oncology patient management decisions.
The test was utilized in over 1400 patient samples during a period of April 2018 and December 2019 across cancer centers to detect multiple actionable alterations in a variety of cancer types.
Reports of these actionable mutations were utilized to inform patient care, including matching patients to available targeted therapies or clinical trials.
The data from the clinical laboratory testing is generally concordant with data in the literature and emphasizes the value of the use of this pan-cancer comprehensive genomic test for the clinical management of patients with advanced cancer.
As of December 2019, Ashion was added to the list of commercial laboratories that are designated to identify and refer eligible patients to the NCI-MATCH trial.
Sign up for free Altmetric alerts about this article
DOI - https://doi.org/10.18632/oncotarget.27945
Full text - https://www.oncotarget.com/article/27945/text/
Correspondence to - Thomas Royce - [email protected]
Keywords - comprehensive genomic profiling, whole exome sequencing, RNA sequencing, precision medicine, pan-cancer
About Oncotarget
Oncotarget is a bi-weekly, peer-reviewed, open access biomedical journal covering research on all aspects of oncology.
To learn more about Oncotarget, please visit https://www.oncotarget.com or connect with:
SoundCloud - https://soundcloud.com/oncotarget
Facebook - https://www.facebook.com/Oncotarget/
Twitter - https://twitter.com/oncotarget
LinkedIn - https://www.linkedin.com/company/oncotarget
Pinterest - https://www.pinterest.com/oncotarget/
Reddit - https://www.reddit.com/user/Oncotarget/
Oncotarget is published by Impact Journals, LLC please visit https://www.ImpactJournals.com or connect with @ImpactJrnls
Media Contact
[email protected]
18009220957x105
Copyright © 2026 Rapamycin Press LLC dba Impact Journals
Oncotarget ® is a registered trademark of Rapamycin Press LLC
Impact Journals ® is a registered trademark of Rapamycin Press LLC
RAPAMYCIN PRESS ® is a registered trademark of Rapamycin Press LLC




