Molecular mechanisms of human IRE1 activation through dimerization and ligand binding
2019-11-14
IRE1 autophosphorylation is coupled to RNase activity through formation of a back-to-back dimer, although the conservation of the underlying molecular mechanism is not clear from existing structures.
The researchers have developed a compound that potently inhibits human IRE1 kinase activity while stimulating XBP1 splicing.
A crystal structure of the inhibitor bound to IRE1 shows an increased ordering of the kinase activation loop.
Dr. Richard Bayliss and Dr. Ian Collins said "The endoplasmic reticulum is responsible for folding secretory proteins. At homeostasis the folding capacity of the ER and the amount of secretory protein synthesis are balanced."
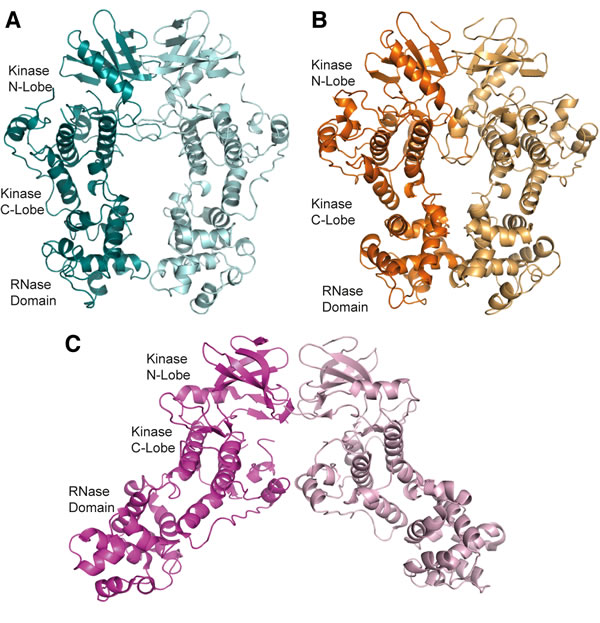
Figure 1: apo-hIRE1 forms a back-to-back dimer. A. The back-to-back dimer conformation of apo-hIRE. Cartoon representation of the two chains of apo-hIRE1 in the crystal structure colored teal and pale blue respectively. B. Cartoon representation of the active phosphorylated yIRE1 dimer in a back-to-back orientation (PDB ID 3FBV) [18], individual chains are shown in light and dark orange. C. Cartoon representation of ADP-hIRE1 (PDB ID 3P23) [22], individual chains are shown in magenta and pink.
RNase activity is coupled to autophosphorylation of the kinase domain activation loop, which is separated from the RNase active site by approximately 50 A.
This orientation of the kinase domains directs the activation loop of one IRE1 molecule towards the active site of the facing IRE1 molecules, and thus provides a rationale for the mechanism of reciprocal autophosphorylation in trans.
However, the kinase active site is lacking several hallmarks of an active kinase, as is often the case in structures of kinases determined without the activating phosphorylation.
The structure of hIRE1 bound to an ATP-competitive inhibitor of kinase and RNase activity forms neither back-to-back dimers nor face-to-face dimers, suggesting perhaps that the compounds might stabilize a monomeric form of IRE1.
These important structures underpin the current model of IRE1 RNase activation, in which trans-autophosphorylation between face-to-face IRE1 dimers generates an active RNase through back-to-back dimerization and higher-order multimerisation.
The Bayliss/Collins Research Team concluded, "Further work will be required to identify which other kinases are regulated through a similar autoinhibitory conformation, and whether formation of a back-to-back dimer provides a general mechanism for release of the tyrosine, leading to kinase activation."
Full text - https://doi.org/10.18632/oncotarget.3864
Correspondence to - Richard Bayliss - [email protected] and Ian Collins - [email protected]
Keywords - UPR, drug discovery, kinase, RNase
Copyright © 2026 Rapamycin Press LLC dba Impact Journals
Oncotarget ® is a registered trademark of Rapamycin Press LLC
Impact Journals ® is a registered trademark of Rapamycin Press LLC
RAPAMYCIN PRESS ® is a registered trademark of Rapamycin Press LLC




