Elucidating drivers of oral epithelial dysplasia formation and malignant transformation to cancer using RNAseq
2019-11-22
To identify markers of early disease, the research team performed RNA sequencing of 19 matched HPV negative patient trios: normal oral mucosa, dysplasia and associated OSCC.
They highlight novel coding and non-coding candidates for involvement in oral dysplasia development and malignant transformation, and speculate on how their findings may guide further translational research into the treatment of oral dysplasia.
Dr. Lucy F. Stead from Precancer Genomics at Leeds Institute of Cancer and Pathology, Wellcome Trust Brenner Building, St James's University Hospital in Leeds, UK said, "Oral squamous cell carcinoma (OSCC) is the 6th most prevalent cancer worldwide with a 5-year survival rate of just 50%."
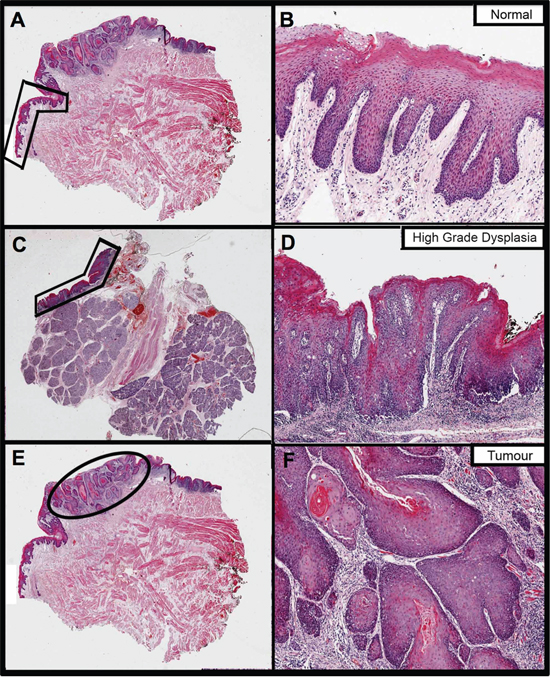
Figure 1: The fixed sections that were H&E stained and annotated to guide RNA extraction from a single patient (ID PG063) in this study. Images on the right are magnifications of the areas annotated on the left, to better show histology. Images A. and B. pertain to the normal oral mucosa sample, images C. and D. to dysplasia and images E. and F. to tumour. Normal and tumour were extracted from the same block whereas dysplasia is from a different block from the same surgery.
OSCC is proposed to occur via a stepwise model whereby genetic abnormalities accumulate, resulting in abnormal lesions called dysplasia, with higher risk of malignant transformation into OSCC than histologically normal oral mucosa.
To further understand the development and malignant transformation of oral dysplasia at the cellular level, they have characterised the transcriptomes of matched normal oral mucosa, oral dysplasia and associated OSCC in 19 patients in unprecedented detail, using high coverage strand-specific RNA sequencing that captures information about both coding and non-coding RNA.
The Stead Research Team concluded, "This includes, but is not limited to, the potential roles of immunoediting, and specifically IL36G, and adherens junction components in malignant transformation, and the role of linc RNA RP11-351J23.1 in de-differentiation of OSCCs."
Full text - https://doi.org/10.18632/oncotarget.5529
Correspondence to - Lucy F. Stead - [email protected]
Keywords - RNAseq, oral squamous cell carcinoma, OSCC, dysplasia, non-coding
Copyright © 2026 Rapamycin Press LLC dba Impact Journals
Oncotarget ® is a registered trademark of Rapamycin Press LLC
Impact Journals ® is a registered trademark of Rapamycin Press LLC
RAPAMYCIN PRESS ® is a registered trademark of Rapamycin Press LLC




