Oncotarget: DNA assay predicts treatment response in head and neck carcinoma
2021-06-22
Oncotarget published "Human papilloma virus circulating tumor DNA assay predicts treatment response in recurrent/metastatic head and neck squamous cell carcinoma" which reported that despite the rising incidence of human papillomavirus related oropharyngeal squamous cell carcinoma, treatment of metastatic disease remains palliative.
Even with new treatments such as immunotherapy, response rates are low and can be delayed, while even mild tumor progression in the face of an ineffective therapy can lead to rapid death.
Real-time biomarkers of response to therapy could improve outcomes by guiding early change of therapy in the metastatic setting. Herein, the authors developed and analytically validated a new droplet digital PCR -based assay for HPV16 circulating tumor DNA and evaluated plasma HPV16 ctDNA for predicting treatment response in metastatic HPV OPSCC.
The authors developed and analytically validated a new droplet digital PCR -based assay for HPV16 circulating tumor DNA
The Oncotarget authors found that longitudinal changes of HPV16 ctDNA correlate with treatment response and that ctDNA responses are observed earlier than conventional imaging.
With additional validation in multi-site studies, this assay may enable early identification of treatment failure, allowing patients to be directed promptly toward clinical trials or alternative therapies.
Dr. Paul L. Swiecicki from The University of Michigan said, "Head and neck squamous cell carcinomas (HNSCC) constitute 3–5% of all malignancies worldwide and there are approximately 600,000 newly diagnosed cases annually"
The most studied ctDNA biomarker in HPV OPSCC is HPV ctDNA.
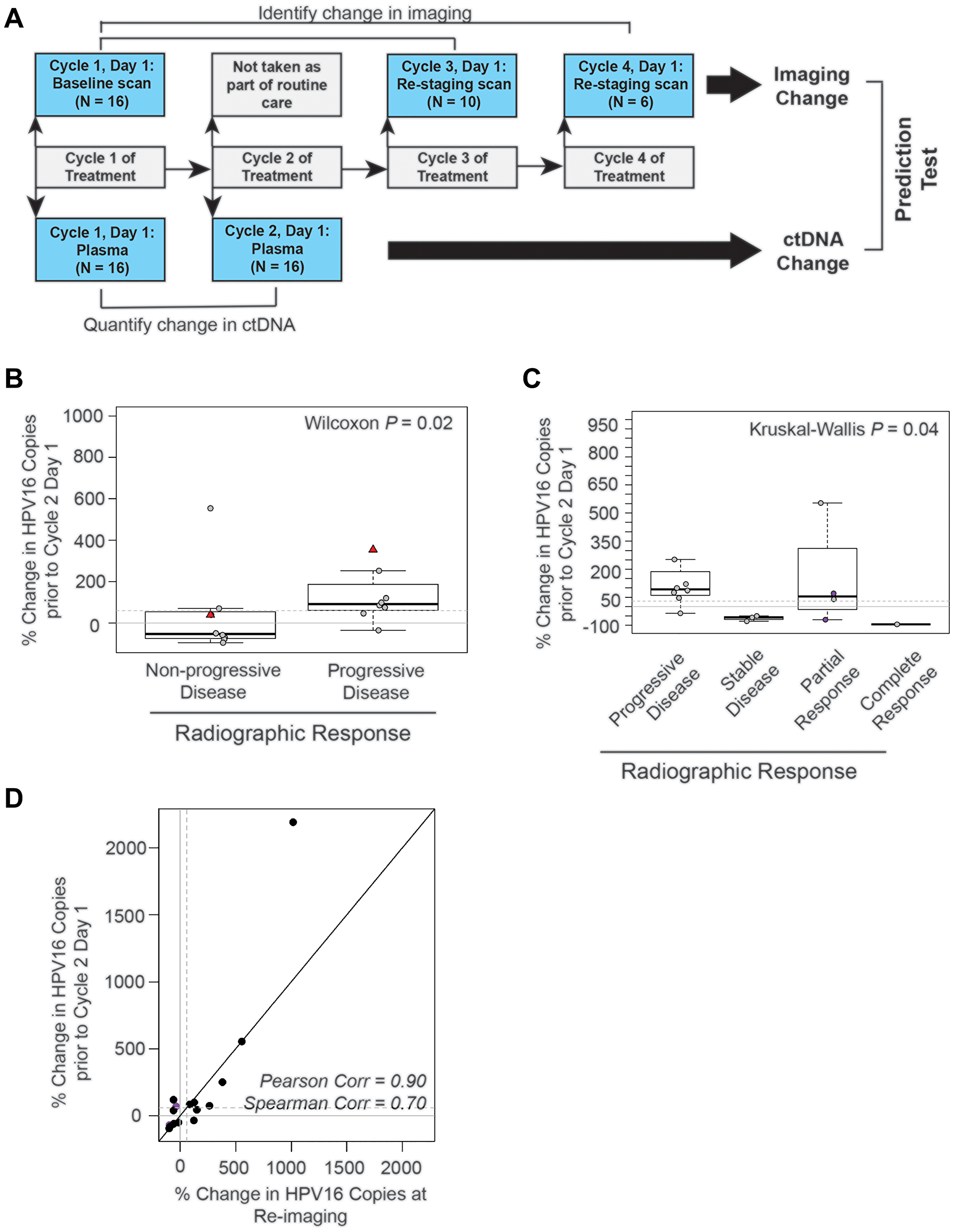
Figure 5: The absolute change in HPV16 ctDNA copies after one cycle of treatment predicts radiographic response in recurrent and metastatic HNSCC patients. (A) Schematic timeline representation of the design of the cohort to evaluate the predictive value of HPV16 ctDNA after one cycle of treatment. Change in radiographic response between each patient's baseline and re-staging CT scans (top bars) were compared to change in HPV16 ctDNA copies between the post-cycle 1 of treatment time point and baseline (bottom bar). For this cohort, we were able to obtain plasma after the first cycle of treatment for 16 of the 18 potential treatment series as two patients missed blood collection; therefore, the total N analyzed is 16. (B) Box-plot analysis. Wilcoxon test was performed to assess the difference between percent change in HPV16 ctDNA after one cycle of treatment in patients with progressive disease (PD) and those deriving benefit (non-PD). The dotted line indicates a 60% change as identified in previous ROC curve analysis. (C) Box-plot analysis. The Kruskal-Wallis analysis test were performed to evaluate for differences in percent change of HPV ctDNA after one cycle of treatment between patients with progressive disease (PD), stable disease (SD), partial response (PR), and complete response (CR) on restaging imaging. Kruskal-Wallis test demonstrated statistically significant changes across response categories (p = 0.04). The dotted line indicates a 60% change. The purple circles highlight patients with pseudoprogression. (D) Scatter plot of percent change in HPV16 ctDNA observed in the draw after one cycle of treatment and the percent change in the blood drawn synchronous with restaging imaging. Spearman and Pearson's rho is reported to assess the magnitude of correlation.
Limited prior data demonstrate that a rapid clearance profile of HPV ctDNA is associated with decreased risk of locoregional recurrence in patients receiving chemoradiation for locally advanced HPV OPSCC, and that levels may increase at the time of recurrence.
Small studies have suggested that HPV ctDNA levels correlate with total disease burden and levels mirror fluctuations in disease status in patients with R/M HPV OPSCC.
However, no study has prospectively examined the potential of HPV ctDNA changes during the course of treatment to predict treatment outcome.
The authors hypothesized that:
- A HPV16 ctDNA test would offer a precise assay for detection of HPV OPSCC and
- The assay would predict progressive disease prior to radiographic imaging in patients undergoing treatment for R/M HPV OPSCC.
The Swiecicki Research Team concluded in their Oncotarget Research Output that this data suggests that changes in HPV16 ctDNA may be predictive of progressive disease in patients with R/M HNSCC receiving systemic therapy.
Changes in HPV16 ctDNA appear to precede radiographic response and thus have the potential to be used as an early predictive biomarker to guide treatment decisions, potentially improving survival and sparing toxicity.
While larger prospective studies are required for validation and clinical utility analyses, this data offers a promising glimpse into the future potential clinical utility of HPV16 ctDNA in HNSCC.
DOI - https://doi.org/10.18632/oncotarget.27992
Full text - https://www.oncotarget.com/article/27992/text/
Correspondence to - Paul L. Swiecicki - [email protected]
Keywords - HPV, head and neck cancer, oropharyngeal cancer, ctDNA, circulating tumor DNA
About Oncotarget
Oncotarget is a bi-weekly, peer-reviewed, open access biomedical journal covering research on all aspects of oncology.
To learn more about Oncotarget, please visit https://www.oncotarget.com or connect with:
SoundCloud - https://soundcloud.com/oncotarget
Facebook - https://www.facebook.com/Oncotarget/
Twitter - https://twitter.com/oncotarget
LinkedIn - https://www.linkedin.com/company/oncotarget
Pinterest - https://www.pinterest.com/oncotarget/
Reddit - https://www.reddit.com/user/Oncotarget/
Oncotarget is published by Impact Journals, LLC please visit https://www.ImpactJournals.com or connect with @ImpactJrnls
Media Contact
[email protected]
18009220957x105
Copyright © 2026 Rapamycin Press LLC dba Impact Journals
Oncotarget ® is a registered trademark of Rapamycin Press LLC
Impact Journals ® is a registered trademark of Rapamycin Press LLC
RAPAMYCIN PRESS ® is a registered trademark of Rapamycin Press LLC




